152:
269:-based ChIP methods, the precision of the ChIP-seq assay is not limited by the spacing of predetermined probes. By integrating a large number of short reads, highly precise binding site localization is obtained. Compared to ChIP-chip, ChIP-seq data can be used to locate the binding site within few tens of base pairs of the actual protein binding site. Tag densities at the binding sites are a good indicator of protein–DNA binding affinity, which makes it easier to quantify and compare binding affinities of a protein to different DNA sites.
383:
369:
341:
to call for broader peaks, spanning over kilobases to megabases in order to search for broader chromatin domains. SICER is more useful for histone marks spanning gene bodies. A mathematical more rigorous method BCP (Bayesian Change Point) can be used for both sharp and broad peaks with faster computational speed, see benchmark comparison of ChIP-seq peak-calling tools by Thomas
171:. However, the widespread use of this method has been limited by the lack of a sufficiently robust method to identify all of the enriched DNA sequences. The ChIP wet lab protocol contains ChIP and hybridization. There are essentially five parts to the ChIP protocol that aid in better understanding the overall process of ChIP. In order to carry out the ChIP, the first step is
327:
analysis pipeline as long as a high-quality genome sequence is available for read mapping and the genome doesn't have repetitive content that confuses the mapping process. ChIP-seq also has the potential to detect mutations in binding-site sequences, which may directly support any observed changes in protein binding and gene regulation.
312:
some of the transcription factors were also identified. Some of the transcription factors regulate genes that control other transcription factors. These genes are not regulated by other factors. Most transcription factors serve as both targets and regulators of other factors, demonstrating a network of regulation.
176:
pieces for ChIP analysis in the end. These fragments should be cut to become under 500 base pairs each to have the best outcome for genome mapping. The third step is called chromatin immunoprecipitation, which is what ChIP is short for. The ChIP process enhances specific crosslinked DNA-protein complexes using an
356:
To reduce spurious sites from ChIP-seq, multiple experimental controls can be used to detect binding sites from an IP experiment. Bay2Ctrls adopts a
Bayesian model to integrate the DNA input control for the IP, the mock IP and its corresponding DNA input control to predict binding sites from the IP.
340:
methods have been developed. One of the most popular methods is MACS which empirically models the shift size of ChIP-Seq tags, and uses it to improve the spatial resolution of predicted binding sites. MACS is optimized for higher resolution peaks, while another popular algorithm, SICER is programmed
180:
against the protein of interest followed by incubation and centrifugation to obtain the immunoprecipitation. The immunoprecipitation step also allows for the removal of non-specific binding sites. The fourth step is DNA recovery and purification, taking place by the reversed effect on the cross-link
311:
to explore genome-wide binding sites of 22 transcription factors. Up to 20% of the annotated candidate genes were assigned to transcription factors. Several transcription factors were assigned to non-coding RNA regions and may be subject to developmental or environmental variables. The functions of
175:
using formaldehyde and large batches of the DNA in order to obtain a useful amount. The cross-links are made between the protein and DNA, but also between RNA and other proteins. The second step is the process of chromatin fragmentation which breaks up the chromatin in order to get high quality DNA
261:
Sensitivity of this technology depends on the depth of the sequencing run (i.e. the number of mapped sequence tags), the size of the genome and the distribution of the target factor. The sequencing depth is directly correlated with cost. If abundant binders in large genomes have to be mapped with
205:
After size selection, all the resulting ChIP-DNA fragments are sequenced simultaneously using a genome sequencer. A single sequencing run can scan for genome-wide associations with high resolution, meaning that features can be located precisely on the chromosomes. ChIP-chip, by contrast, requires
326:
ChIP-seq offers an alternative to ChIP-chip. STAT1 experimental ChIP-seq data have a high degree of similarity to results obtained by ChIP-chip for the same type of experiment, with greater than 64% of peaks in shared genomic regions. Because the data are sequence reads, ChIP-seq offers a rapid
322:
were shown to be more correlated with transcription factor motifs at promoters in comparison to RNA level. Hence author proposed that using histone modification ChIP-seq would provide more reliable inference of gene-regulatory networks in comparison to other methods based on expression.
230:
in parallel using novel fluorescently labelled reversible terminator nucleotides. Templates are sequenced base-by-base during each read. Then, the data collection and analysis software aligns sample sequences to a known genomic sequence to identify the ChIP-DNA fragments.
107:. ChIP produces a library of target DNA sites bound to a protein of interest. Massively parallel sequence analyses are used in conjunction with whole-genome sequence databases to analyze the interaction pattern of any protein with DNA, or the pattern of any epigenetic
357:
This approach is particularly effective for complex samples such as whole model organisms. In addition, the analysis indicates that for complex samples mock IP controls substantially outperform DNA input controls probably due to the active genomes of the samples.
348:
Another relevant computational problem is differential peak calling, which identifies significant differences in two ChIP-seq signals from distinct biological conditions. Differential peak callers segment two ChIP-seq signals and identify differential peaks using
100:. This introduces some bias, as an array is restricted to a fixed number of probes. Sequencing, by contrast, is thought to have less bias, although the sequencing bias of different sequencing technologies is not yet fully understood.
2002:: GeneProf is a freely accessible, easy-to-use analysis environment for ChIP-seq and RNA-seq data and comes with a large database of ready-analysed public experiments, e.g. for transcription factor binding and histone modifications.
225:
substrate to create clusters of approximately 1000 clonal copies each. The resulting high density array of template clusters on the flow cell surface is sequenced by a genome analyzing program. Each template cluster undergoes
335:
As with many high-throughput sequencing approaches, ChIP-seq generates extremely large data sets, for which appropriate computational analysis methods are required. To predict DNA-binding sites from ChIP-seq read count data,
1050:
Jung, Youngsook L.; Luquette, Lovelace J.; Ho, Joshua W. K.; Ferrari, Francesco; Tolstorukov, Michael; Minoda, Aki; Issner, Robbyn; Epstein, Charles B.; Karpen, Gary H.; Kuroda, Mitzi I.; Park, Peter J. (May 2014).
284:
S3 cells which are clones of the HeLa line that are used for analysis of cell populations. The performance of ChIP-seq was then compared to the alternative protein–DNA interaction methods of ChIP-PCR and ChIP-chip.
1762:
Xu, Jinrui; Kudron, Michelle M; Victorsen, Alec; Gao, Jiahao; Ammouri, Haneen N; Navarro, Fabio C P; Gevirtzman, Louis; Waterston, Robert H; White, Kevin P; Reinke, Valerie; Gerstein, Mark (21 December 2020).
1292:
Robertson G, Hirst M, Bainbridge M, Bilenky M, Zhao Y, Zeng T, et al. (August 2007). "Genome-wide profiles of STAT1 DNA association using chromatin immunoprecipitation and massively parallel sequencing".
993:
Chen, Yiwen; Negre, Nicolas; Li, Qunhua; Mieczkowska, Joanna O.; Slattery, Matthew; Liu, Tao; Zhang, Yong; Kim, Tae-Kyung; He, Housheng Hansen; Zieba, Jennifer; Ruan, Yijun (June 2012).
262:
high sensitivity, costs are high as an enormously high number of sequence tags will be required. This is in contrast to ChIP-chip in which the costs are not correlated with sensitivity.
291:
Using ChIP-seq, it was determined that Yeast genes seem to have a minimal nucleosome-free promoter region of 150bp in which RNA polymerase can initiate transcription.
297:
ChIP-seq was used to compare conservation of TFs in the forebrain and heart tissue in embryonic mice. The authors identified and validated the heart functionality of
181:
between DNA and protein to separate them and cleaning DNA with an extraction. The fifth and final step is the analyzation step of the ChIP protocol by the process of
2030:
463:
218:
128:
301:, and determined that transcription enhancers for the heart are less conserved than those for the forebrain during the same developmental stage.
426:, antibody-targeted controlled cleavage by micrococcal nuclease instead of ChIP, allowing for enhanced signal-to-noise ratio during sequencing.
487:
432:, antibody-targeted controlled cleavage by transposase Tn5 instead of ChIP, allowing for enhanced signal-to-noise ratio during sequencing.
1724:
Allhoff M, Seré K, Chauvistré H, Lin Q, Zenke M, Costa IG (December 2014). "Detecting differential peaks in ChIP-seq signals with ODIN".
222:
2008:
1986:
analysis of regulatory elements from +2800 ChIP-seq datasets, giving a catalogue of 80 million peaks from 485 transcription regulators.
401:
239:
ChIP-seq offers us a fast analysis, however, a quality control must be performed to make sure that the results obtained are reliable:
197:. Through the analysis, the sequences can then be identified and interpreted by the gene or region to where the protein was bound.
774:
Kumar, Vibhor; Muratani, Masafumi; Rayan, Nirmala Arul; Kraus, Petra; Lufkin, Thomas; Ng, Huck Hui; Prabhakar, Shyam (July 2013).
50:
131:. As an alternative to the dependence on specific antibodies, different methods have been developed to find the superset of all
1577:"Genome-wide localization of protein-DNA binding and histone modification by a Bayesian change-point method with ChIP-seq data"
246:: low-complexity regions should be removed as they are not informative and may interfere with mapping in the reference genome.
662:"DNase-seq: a high-resolution technique for mapping active gene regulatory elements across the genome from mammalian cells"
60:
of DNA-associated proteins. It can be used to map global binding sites precisely for any protein of interest. Previously,
1108:
Ho, Joshua W. K.; Bishop, Eric; Karchenko, Peter V.; Nègre, Nicolas; White, Kevin P.; Park, Peter J. (28 February 2011).
719:"FAIRE (Formaldehyde-Assisted Isolation of Regulatory Elements) isolates active regulatory elements from human chromatin"
182:
111:
modifications. This can be applied to the set of ChIP-able proteins and modifications, such as transcription factors,
2063:"ReMap 2018: an updated atlas of regulatory regions from an integrative analysis of DNA-binding ChIP-seq experiments"
643:
103:
Specific DNA sites in direct physical interaction with transcription factors and other proteins can be isolated by
2018:
528:"RNA-seq and ChIP-seq as Complementary Approaches for Comprehension of Plant Transcriptional Regulatory Mechanism"
17:
2189:
1685:"An HMM approach to genome-wide identification of differential histone modification sites from ChIP-seq data"
460:, same goal and first steps, but does not use cross linking methods and uses microarray instead of sequencing
104:
46:
1979:
2184:
193:
adaptors are then added to the small stretches of DNA that were bound to the protein of interest to enable
526:
Muhammad, Isiaka
Ibrahim; Kong, Sze Ling; Akmar Abdullah, Siti Nor; Munusamy, Umaiyal (25 December 2019).
411:
194:
1996:
data. It provides the most comprehensive ChIP-Seq data set for various cell/tissue types and conditions.
586:
1218:
Bernstein BE, Kamal M, Lindblad-Toh K, Bekiranov S, Bailey DK, Huebert DJ, et al. (January 2005).
2040:
1267:
214:
717:
Giresi, Paul G.; Kim, Jonghwan; McDaniell, Ryan M.; Iyer, Vishwanath R.; Lieb, Jason D. (June 2007).
167:
is a powerful method to selectively enrich for DNA sequences bound by a particular protein in living
838:
116:
2005:
1930:"Calling Cards enable multiplexed identification of the genomic targets of DNA-binding proteins"
2174:
2027:: Uncovering correlated variability in epigenomic datasets using the Karhunen-Loeve transform.
1436:"Diverse transcription factor binding features revealed by genome-wide ChIP-seq in C. elegans"
252:
ratio of reads that are located in peaks over reads that are located where there isn't a peak.
92:
and expression analysis. ChIP-seq technology is currently seen primarily as an alternative to
429:
423:
73:
2123:
1835:
1588:
601:
454:, another method for identifying the binding sites of cellular RNA-binding proteins (RBPs).
319:
151:
124:
509:
Calling Cards, uses a transposase to mark the sequence where a transcription factor binds.
8:
350:
298:
172:
2127:
1839:
1592:
605:
84:
is essential for fully understanding many biological processes and disease states. This
2146:
2111:
2087:
2062:
1954:
1929:
1905:
1880:
1856:
1823:
1799:
1764:
1660:
1635:
1611:
1576:
1552:
1525:
1460:
1435:
1411:
1386:
1385:
Blow MJ, McCulley DJ, Li Z, Zhang T, Akiyama JA, Holt A, et al. (September 2010).
1367:
1318:
1249:
1195:
1168:
1144:
1109:
1085:
1052:
1027:
994:
967:
934:
813:
751:
718:
694:
661:
625:
562:
527:
97:
1822:
Licatalosi DD, Mele A, Fak JJ, Ule J, Kayikci M, Chi SW, et al. (November 2008).
2151:
2092:
1959:
1910:
1861:
1804:
1786:
1741:
1706:
1665:
1616:
1557:
1524:
Zhang Y, Liu T, Meyer CA, Eeckhoute J, Johnson DS, Bernstein BE, et al. (2008).
1506:
1465:
1416:
1359:
1310:
1241:
1200:
1149:
1131:
1090:
1072:
1032:
1014:
972:
954:
915:
880:
805:
797:
756:
738:
699:
681:
617:
567:
549:
388:
120:
1737:
1701:
1684:
1371:
1322:
1169:"Genome-wide identification of in vivo protein–DNA binding sites from ChIP-seq data"
817:
2141:
2131:
2082:
2074:
1949:
1941:
1900:
1892:
1851:
1843:
1794:
1776:
1733:
1696:
1655:
1647:
1606:
1596:
1547:
1537:
1496:
1455:
1447:
1434:
Niu W, Lu ZJ, Zhong M, Sarov M, Murray JI, Brdlik CM, et al. (February 2011).
1406:
1398:
1349:
1302:
1253:
1231:
1220:"Genomic maps and comparative analysis of histone modifications in human and mouse"
1190:
1180:
1139:
1121:
1080:
1064:
1022:
1006:
962:
946:
907:
872:
787:
746:
730:
689:
673:
609:
557:
539:
469:
217:
used in this sequencing step. Some technologies that analyze the sequences can use
1765:"To mock or not: a comprehensive comparison of mock IP and DNA input for ChIP-seq"
1483:
Kumar V, Muratani M, Rayan NA, Kraus P, Lufkin T, Ng HH, Prabhakar S (July 2013).
629:
2136:
2012:
1601:
898:
Kim TH, Dekker J (April 2018). "Preparation of Cross-Linked
Chromatin for ChIP".
190:
81:
1815:
500:
uses S9.6 antibody to precipitate three-stranded DND:RNA hybrids called R-loops.
135:-depleted or nucleosome-disrupted active regulatory regions in the genome, like
2110:
Bailey T, Krajewski P, Ladunga I, Lefebvre C, Li Q, Liu T, et al. (2013).
1354:
1337:
1236:
1219:
406:
374:
227:
168:
53:
2024:
1110:"ChIP-chip versus ChIP-seq: lessons for experimental design and data analysis"
80:-affecting mechanisms. Determining how proteins interact with DNA to regulate
2168:
1824:"HITS-CLIP yields genome-wide insights into brain alternative RNA processing"
1790:
1542:
1135:
1126:
1076:
1018:
958:
801:
742:
685:
553:
64:
was the most common technique utilized to study these protein–DNA relations.
613:
2179:
2155:
2096:
2061:
Chèneby J, Gheorghe M, Artufel M, Mathelier A, Ballester B (January 2018).
1963:
1914:
1865:
1808:
1781:
1745:
1710:
1669:
1620:
1561:
1510:
1469:
1420:
1363:
1314:
1245:
1204:
1153:
1094:
1036:
976:
919:
911:
884:
876:
809:
760:
703:
621:
571:
396:
337:
207:
186:
61:
57:
2078:
1945:
1451:
353:. Examples for two-stage differential peak callers are ChIPDiff and ODIN.
1651:
1185:
1068:
677:
544:
1847:
1010:
734:
484:
uses exonuclease treatment to achieve up to single base-pair resolution
266:
132:
112:
85:
2112:"Practical guidelines for the comprehensive analysis of ChIP-seq data"
1881:"HITS-CLIP: panoramic views of protein-RNA regulation in living cells"
1306:
1501:
1484:
792:
775:
475:
441:
140:
136:
108:
93:
77:
2033:: a tool for probabilistic pattern discovery on multiple normalized
1896:
950:
2044:
2043:: a tool for regression and peak prediction on multiple normalized
2034:
1993:
1989:
1983:
1485:"Uniform, optimal signal processing of mapped deep-sequencing data"
1402:
1217:
776:"Uniform, optimal signal processing of mapped deep-sequencing data"
497:
491:
481:
457:
451:
445:
435:
368:
177:
89:
1992:: a database for exploring transcription factor binding maps from
525:
506:, principally similar method to measure mRNA translation dynamics.
438:, identical to ChIP-Seq but skipping the immunoprecipitation step.
382:
1338:"ChIP-Seq data reveal nucleosome architecture of human promoters"
503:
38:
1636:"Features that define the best ChIP-seq peak calling algorithms"
995:"Systematic evaluation of factors influencing ChIP-seq fidelity"
2060:
1291:
2109:
935:"ChIP-seq: advantages and challenges of a maturing technology"
1387:"ChIP-Seq identification of weakly conserved heart enhancers"
863:
Kim TH, Dekker J (April 2018). "Formaldehyde Cross-Linking".
1928:
Wang H, Mayhew D, Chen X, Johnston M, Mitra RD (May 2011).
1633:
281:
164:
1999:
587:"Genome-wide mapping of in vivo protein-DNA interactions"
584:
42:
1723:
1482:
773:
585:
Johnson DS, Mortazavi A, Myers RM, Wold B (June 2007).
1927:
1523:
1107:
992:
716:
1049:
660:
Song, Lingyun; Crawford, Gregory E. (February 2010).
448:), for finding interactions with RNA rather than DNA.
1821:
1761:
1634:
Thomas R, Thomas S, Holloway AK, Pollard KS (2017).
1053:"Impact of sequencing depth in ChIP-seq experiments"
364:
27:
Method used to analyze protein interactions with DNA
2015:: Tutorial for differential peak calling with ODIN.
1384:
466:, a method for finding a consensus binding sequence
472:, to measure relative replacement dynamics on DNA.
76:and other chromatin-associated proteins influence
644:"Whole-Genome Chromatin IP Sequencing (ChIP-Seq)"
221:of adapter-ligated ChIP DNA fragments on a solid
2166:
1574:
1433:
839:"ChIP guide: epigenetics applications | Abcam"
146:
1683:Xu H, Wei CL, Lin F, Sung WK (October 2008).
494:to achieve up to single base-pair resolution.
1682:
1335:
659:
280:ChIP-seq was used to study STAT1 targets in
72:ChIP-seq is primarily used to determine how
532:International Journal of Molecular Sciences
2021:: Comprehensive analysis of ChIP-seq data.
1476:
307:ChIP-sequencing was completed on the worm
2145:
2135:
2086:
1953:
1904:
1855:
1798:
1780:
1700:
1659:
1610:
1600:
1551:
1541:
1526:"Model-based analysis of ChIP-Seq (MACS)"
1500:
1459:
1410:
1353:
1235:
1194:
1184:
1143:
1125:
1084:
1026:
966:
897:
862:
791:
750:
693:
561:
543:
330:
150:
2019:Bioinformatic analysis of ChIP-seq data
1878:
1575:Xing H, Mo Y, Liao W, Zhang MQ (2012).
14:
2167:
478:to measure RNA-bound DNA and proteins.
1757:
1755:
1336:Schmid CD, Bucher P (November 2007).
1166:
988:
986:
289:Nucleosome Architecture of Promoters:
1885:Wiley Interdisciplinary Reviews. RNA
1268:"HeLa S3 from ATCC | Biocompare.com"
932:
833:
831:
829:
827:
2000:GeneProf database and analysis tool
272:
189:(hybrid array) or ChIP sequencing.
24:
1752:
983:
649:. Illumina, Inc. 26 November 2007.
417:
295:Transcription factor conservation:
234:
25:
2201:
1973:
824:
1348:(5): 831–2, author reply 832–3.
381:
367:
88:information is complementary to
2103:
2054:
1921:
1872:
1717:
1676:
1627:
1568:
1517:
1427:
1378:
1329:
1285:
1260:
1211:
1160:
1101:
1043:
933:Park, Peter J. (October 2009).
926:
891:
856:
767:
710:
653:
636:
578:
519:
256:
37:, is a method used to analyze
13:
1:
1982:: An integrative and uniform
1738:10.1093/bioinformatics/btu722
1702:10.1093/bioinformatics/btn402
513:
316:Inferring regulatory network:
200:
195:massively parallel sequencing
105:chromatin immunoprecipitation
47:chromatin immunoprecipitation
2137:10.1371/journal.pcbi.1003326
1602:10.1371/journal.pcbi.1002613
900:Cold Spring Harbor Protocols
865:Cold Spring Harbor Protocols
666:Cold Spring Harbor Protocols
7:
1167:Jothi, et al. (2008).
412:Mammalian promoter database
360:
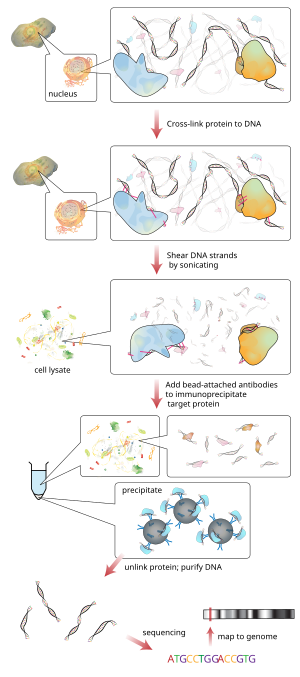
147:Workflow of ChIP-sequencing
10:
2206:
2116:PLOS Computational Biology
1355:10.1016/j.cell.2007.11.017
1237:10.1016/j.cell.2005.01.001
2006:Differential Peak Calling
117:transcriptional machinery
1543:10.1186/gb-2008-9-9-r137
1127:10.1186/1471-2164-12-134
939:Nature Reviews. Genetics
155:ChIP-sequencing workflow
614:10.1126/science.1141319
299:transcription enhancers
228:sequencing-by-synthesis
159:
67:
2067:Nucleic Acids Research
1769:Nucleic Acids Research
1057:Nucleic Acids Research
912:10.1101/pdb.prot082602
877:10.1101/pdb.prot082594
430:CUT&Tag sequencing
424:CUT&RUN sequencing
331:Computational analysis
278:STAT1 DNA association:
244:Non-redundant fraction
215:new sequencing methods
210:for lower resolution.
156:
1946:10.1101/gr.114850.110
1452:10.1101/gr.114587.110
906:(4): pdb.prot082602.
871:(4): pdb.prot082594.
305:Genome-wide ChIP-seq:
219:cluster amplification
154:
125:protein modifications
74:transcription factors
2190:Proteomic sequencing
1782:10.1093/nar/gkaa1155
1489:Nature Biotechnology
780:Nature Biotechnology
678:10.1101/pdb.prot5384
545:10.3390/ijms21010167
490:improved version of
351:Hidden Markov Models
320:Histone modification
45:. ChIP-seq combines
2185:Genomics techniques
2128:2013PLSCB...9E3326B
2079:10.1093/nar/gkx1092
2011:15 May 2021 at the
1879:Darnell RB (2010).
1848:10.1038/nature07488
1840:2008Natur.456..464L
1593:2012PLSCB...8E2613X
672:(2): pdb.prot5384.
606:2007Sci...316.1497J
318:ChIP-seq signal of
250:Fragments in peaks:
121:structural proteins
98:hybridization array
1652:10.1093/bib/bbw035
1272:www.biocompare.com
1186:10.1093/nar/gkn488
1069:10.1093/nar/gku178
1011:10.1038/nmeth.1985
735:10.1101/gr.5533506
600:(5830): 1497–502.
157:
51:massively parallel
41:interactions with
2073:(D1): D267–D275.
1990:ChIPBase database
1307:10.1038/nmeth1068
1179:(16): 5221–5231.
1173:Nucleic Acids Res
389:Technology portal
129:DNA modifications
96:which requires a
16:(Redirected from
2197:
2160:
2159:
2149:
2139:
2122:(11): e1003326.
2107:
2101:
2100:
2090:
2058:
2041:FullSignalRanker
1968:
1967:
1957:
1925:
1919:
1918:
1908:
1876:
1870:
1869:
1859:
1819:
1813:
1812:
1802:
1784:
1759:
1750:
1749:
1721:
1715:
1714:
1704:
1680:
1674:
1673:
1663:
1631:
1625:
1624:
1614:
1604:
1581:PLOS Comput Biol
1572:
1566:
1565:
1555:
1545:
1521:
1515:
1514:
1504:
1502:10.1038/nbt.2596
1480:
1474:
1473:
1463:
1431:
1425:
1424:
1414:
1382:
1376:
1375:
1357:
1333:
1327:
1326:
1289:
1283:
1282:
1280:
1278:
1264:
1258:
1257:
1239:
1215:
1209:
1208:
1198:
1188:
1164:
1158:
1157:
1147:
1129:
1105:
1099:
1098:
1088:
1047:
1041:
1040:
1030:
990:
981:
980:
970:
930:
924:
923:
895:
889:
888:
860:
854:
853:
851:
849:
835:
822:
821:
795:
793:10.1038/nbt.2596
771:
765:
764:
754:
714:
708:
707:
697:
657:
651:
650:
648:
640:
634:
633:
591:
582:
576:
575:
565:
547:
523:
470:Competition-ChIP
391:
386:
385:
377:
372:
371:
273:Current research
56:to identify the
33:, also known as
21:
2205:
2204:
2200:
2199:
2198:
2196:
2195:
2194:
2165:
2164:
2163:
2108:
2104:
2059:
2055:
2047:signal profiles
2037:signal profiles
2013:Wayback Machine
1980:ReMap catalogue
1976:
1971:
1934:Genome Research
1926:
1922:
1897:10.1002/wrna.31
1877:
1873:
1834:(7221): 464–9.
1820:
1816:
1775:(3): gkaa1155.
1760:
1753:
1732:(24): 3467–75.
1722:
1718:
1681:
1677:
1640:Brief Bioinform
1632:
1628:
1587:(7): e1002613.
1573:
1569:
1522:
1518:
1481:
1477:
1440:Genome Research
1432:
1428:
1391:Nature Genetics
1383:
1379:
1334:
1330:
1290:
1286:
1276:
1274:
1266:
1265:
1261:
1216:
1212:
1165:
1161:
1106:
1102:
1048:
1044:
991:
984:
951:10.1038/nrg2641
945:(10): 669–680.
931:
927:
896:
892:
861:
857:
847:
845:
837:
836:
825:
772:
768:
723:Genome Research
715:
711:
658:
654:
646:
642:
641:
637:
589:
583:
579:
524:
520:
516:
420:
418:Similar methods
387:
380:
373:
366:
363:
333:
275:
259:
237:
235:Quality control
213:There are many
203:
191:Oligonucleotide
162:
149:
82:gene expression
70:
31:ChIP-sequencing
28:
23:
22:
18:Chip-sequencing
15:
12:
11:
5:
2203:
2193:
2192:
2187:
2182:
2177:
2162:
2161:
2102:
2052:
2049:
2048:
2038:
2028:
2022:
2016:
2003:
1997:
1987:
1975:
1974:External links
1972:
1970:
1969:
1920:
1891:(2): 266–286.
1871:
1814:
1751:
1726:Bioinformatics
1716:
1695:(20): 2344–9.
1689:Bioinformatics
1675:
1646:(3): 441–450.
1626:
1567:
1530:Genome Biology
1516:
1475:
1426:
1403:10.1038/ng.650
1377:
1328:
1295:Nature Methods
1284:
1259:
1210:
1159:
1100:
1042:
1005:(6): 609–614.
999:Nature Methods
982:
925:
890:
855:
823:
786:(7): 615–622.
766:
729:(6): 877–885.
709:
652:
635:
577:
517:
515:
512:
511:
510:
507:
501:
495:
485:
479:
473:
467:
461:
455:
449:
439:
433:
427:
419:
416:
415:
414:
409:
404:
399:
393:
392:
378:
375:Biology portal
362:
359:
332:
329:
274:
271:
258:
255:
254:
253:
247:
236:
233:
206:large sets of
202:
199:
161:
158:
148:
145:
69:
66:
54:DNA sequencing
26:
9:
6:
4:
3:
2:
2202:
2191:
2188:
2186:
2183:
2181:
2178:
2176:
2175:Biotechnology
2173:
2172:
2170:
2157:
2153:
2148:
2143:
2138:
2133:
2129:
2125:
2121:
2117:
2113:
2106:
2098:
2094:
2089:
2084:
2080:
2076:
2072:
2068:
2064:
2057:
2053:
2051:
2046:
2042:
2039:
2036:
2032:
2029:
2026:
2023:
2020:
2017:
2014:
2010:
2007:
2004:
2001:
1998:
1995:
1991:
1988:
1985:
1981:
1978:
1977:
1965:
1961:
1956:
1951:
1947:
1943:
1940:(5): 748–55.
1939:
1935:
1931:
1924:
1916:
1912:
1907:
1902:
1898:
1894:
1890:
1886:
1882:
1875:
1867:
1863:
1858:
1853:
1849:
1845:
1841:
1837:
1833:
1829:
1825:
1818:
1810:
1806:
1801:
1796:
1792:
1788:
1783:
1778:
1774:
1770:
1766:
1758:
1756:
1747:
1743:
1739:
1735:
1731:
1727:
1720:
1712:
1708:
1703:
1698:
1694:
1690:
1686:
1679:
1671:
1667:
1662:
1657:
1653:
1649:
1645:
1641:
1637:
1630:
1622:
1618:
1613:
1608:
1603:
1598:
1594:
1590:
1586:
1582:
1578:
1571:
1563:
1559:
1554:
1549:
1544:
1539:
1535:
1531:
1527:
1520:
1512:
1508:
1503:
1498:
1495:(7): 615–22.
1494:
1490:
1486:
1479:
1471:
1467:
1462:
1457:
1453:
1449:
1446:(2): 245–54.
1445:
1441:
1437:
1430:
1422:
1418:
1413:
1408:
1404:
1400:
1397:(9): 806–10.
1396:
1392:
1388:
1381:
1373:
1369:
1365:
1361:
1356:
1351:
1347:
1343:
1339:
1332:
1324:
1320:
1316:
1312:
1308:
1304:
1300:
1296:
1288:
1273:
1269:
1263:
1255:
1251:
1247:
1243:
1238:
1233:
1230:(2): 169–81.
1229:
1225:
1221:
1214:
1206:
1202:
1197:
1192:
1187:
1182:
1178:
1174:
1170:
1163:
1155:
1151:
1146:
1141:
1137:
1133:
1128:
1123:
1119:
1115:
1111:
1104:
1096:
1092:
1087:
1082:
1078:
1074:
1070:
1066:
1062:
1058:
1054:
1046:
1038:
1034:
1029:
1024:
1020:
1016:
1012:
1008:
1004:
1000:
996:
989:
987:
978:
974:
969:
964:
960:
956:
952:
948:
944:
940:
936:
929:
921:
917:
913:
909:
905:
901:
894:
886:
882:
878:
874:
870:
866:
859:
844:
843:www.abcam.com
840:
834:
832:
830:
828:
819:
815:
811:
807:
803:
799:
794:
789:
785:
781:
777:
770:
762:
758:
753:
748:
744:
740:
736:
732:
728:
724:
720:
713:
705:
701:
696:
691:
687:
683:
679:
675:
671:
667:
663:
656:
645:
639:
631:
627:
623:
619:
615:
611:
607:
603:
599:
595:
588:
581:
573:
569:
564:
559:
555:
551:
546:
541:
537:
533:
529:
522:
518:
508:
505:
502:
499:
496:
493:
489:
486:
483:
480:
477:
474:
471:
468:
465:
462:
459:
456:
453:
450:
447:
444:(also called
443:
440:
437:
434:
431:
428:
425:
422:
421:
413:
410:
408:
405:
403:
400:
398:
395:
394:
390:
384:
379:
376:
370:
365:
358:
354:
352:
346:
344:
339:
328:
324:
321:
317:
313:
310:
306:
302:
300:
296:
292:
290:
286:
283:
279:
270:
268:
263:
251:
248:
245:
242:
241:
240:
232:
229:
224:
220:
216:
211:
209:
208:tiling arrays
198:
196:
192:
188:
184:
179:
174:
173:cross-linking
170:
166:
153:
144:
142:
138:
134:
130:
126:
122:
118:
114:
110:
106:
101:
99:
95:
91:
87:
83:
79:
75:
65:
63:
59:
58:binding sites
55:
52:
48:
44:
40:
36:
32:
19:
2119:
2115:
2105:
2070:
2066:
2056:
2050:
2031:SignalSpider
2025:KLTepigenome
1937:
1933:
1923:
1888:
1884:
1874:
1831:
1827:
1817:
1772:
1768:
1729:
1725:
1719:
1692:
1688:
1678:
1643:
1639:
1629:
1584:
1580:
1570:
1533:
1529:
1519:
1492:
1488:
1478:
1443:
1439:
1429:
1394:
1390:
1380:
1345:
1341:
1331:
1301:(8): 651–7.
1298:
1294:
1287:
1275:. Retrieved
1271:
1262:
1227:
1223:
1213:
1176:
1172:
1162:
1117:
1114:BMC Genomics
1113:
1103:
1060:
1056:
1045:
1002:
998:
942:
938:
928:
903:
899:
893:
868:
864:
858:
846:. Retrieved
842:
783:
779:
769:
726:
722:
712:
669:
665:
655:
638:
597:
593:
580:
535:
531:
521:
397:ChIP-on-chip
355:
347:
342:
338:peak calling
334:
325:
315:
314:
308:
304:
303:
294:
293:
288:
287:
277:
276:
264:
260:
249:
243:
238:
212:
204:
187:ChIP-on-chip
163:
102:
71:
62:ChIP-on-chip
49:(ChIP) with
34:
30:
29:
1536:(9): R137.
257:Sensitivity
113:polymerases
2169:Categories
1063:(9): e74.
538:(1): 167.
514:References
488:ChIP-nexus
309:C. elegans
267:microarray
201:Sequencing
133:nucleosome
86:epigenetic
1791:0305-1048
1136:1471-2164
1077:1362-4962
1019:1548-7105
959:1471-0064
802:1087-0156
743:1088-9051
686:1559-6095
554:1422-0067
476:ChiRP-Seq
442:HITS-CLIP
223:flow cell
141:FAIRE-Seq
137:DNase-Seq
109:chromatin
94:ChIP-chip
78:phenotype
2156:24244136
2097:29126285
2045:ChIP-Seq
2035:ChIP-Seq
2009:Archived
1994:ChIP-Seq
1984:ChIP-Seq
1964:21471402
1915:21935890
1866:18978773
1809:33347581
1746:25371479
1711:18667444
1670:27169896
1621:22844240
1562:18798982
1511:23770639
1470:21177963
1421:20729851
1372:29234049
1364:18045524
1323:28531263
1315:17558387
1277:21 March
1246:15680324
1205:18684996
1154:21356108
1095:24598259
1037:22522655
977:19736561
920:29610358
885:29610357
818:32510475
810:23770639
761:17179217
704:20150147
622:17540862
572:31881735
498:DRIP-seq
492:ChIP-exo
482:ChIP-exo
458:RIP-Chip
452:PAR-CLIP
446:CLIP-Seq
436:Sono-Seq
407:ChIP-PET
402:ChIP-PCR
361:See also
345:(2017).
178:antibody
90:genotype
35:ChIP-seq
2147:3828144
2124:Bibcode
2088:5753247
1955:3083092
1906:3222227
1857:2597294
1836:Bibcode
1800:7897498
1661:5429005
1612:5429005
1589:Bibcode
1553:2592715
1461:3032928
1412:3138496
1254:7193829
1196:2532738
1145:3053263
1120:: 134.
1086:4027199
1028:3477507
968:3191340
848:2 March
752:1891346
695:3627383
602:Bibcode
594:Science
563:6981605
504:TCP-seq
265:Unlike
39:protein
2154:
2144:
2095:
2085:
1962:
1952:
1913:
1903:
1864:
1854:
1828:Nature
1807:
1797:
1789:
1744:
1709:
1668:
1658:
1619:
1609:
1560:
1550:
1509:
1468:
1458:
1419:
1409:
1370:
1362:
1321:
1313:
1252:
1244:
1203:
1193:
1152:
1142:
1134:
1093:
1083:
1075:
1035:
1025:
1017:
975:
965:
957:
918:
883:
816:
808:
800:
759:
749:
741:
702:
692:
684:
630:519841
628:
620:
570:
560:
552:
343:et al.
127:, and
1368:S2CID
1319:S2CID
1250:S2CID
814:S2CID
647:(PDF)
626:S2CID
590:(PDF)
464:SELEX
169:cells
2152:PMID
2093:PMID
1960:PMID
1911:PMID
1862:PMID
1805:PMID
1787:ISSN
1742:PMID
1707:PMID
1666:PMID
1617:PMID
1558:PMID
1507:PMID
1466:PMID
1417:PMID
1360:PMID
1342:Cell
1311:PMID
1279:2020
1242:PMID
1224:Cell
1201:PMID
1150:PMID
1132:ISSN
1091:PMID
1073:ISSN
1033:PMID
1015:ISSN
973:PMID
955:ISSN
916:PMID
904:2018
881:PMID
869:2018
850:2020
806:PMID
798:ISSN
757:PMID
739:ISSN
700:PMID
682:ISSN
670:2010
618:PMID
568:PMID
550:ISSN
282:HeLa
183:qPCR
165:ChIP
160:ChIP
139:and
115:and
68:Uses
2180:DNA
2142:PMC
2132:doi
2083:PMC
2075:doi
1950:PMC
1942:doi
1901:PMC
1893:doi
1852:PMC
1844:doi
1832:456
1795:PMC
1777:doi
1734:doi
1697:doi
1656:PMC
1648:doi
1607:PMC
1597:doi
1548:PMC
1538:doi
1497:doi
1456:PMC
1448:doi
1407:PMC
1399:doi
1350:doi
1346:131
1303:doi
1232:doi
1228:120
1191:PMC
1181:doi
1140:PMC
1122:doi
1081:PMC
1065:doi
1023:PMC
1007:doi
963:PMC
947:doi
908:doi
873:doi
788:doi
747:PMC
731:doi
690:PMC
674:doi
610:doi
598:316
558:PMC
540:doi
43:DNA
2171::
2150:.
2140:.
2130:.
2118:.
2114:.
2091:.
2081:.
2071:46
2069:.
2065:.
1958:.
1948:.
1938:21
1936:.
1932:.
1909:.
1899:.
1887:.
1883:.
1860:.
1850:.
1842:.
1830:.
1826:.
1803:.
1793:.
1785:.
1773:49
1771:.
1767:.
1754:^
1740:.
1730:30
1728:.
1705:.
1693:24
1691:.
1687:.
1664:.
1654:.
1644:18
1642:.
1638:.
1615:.
1605:.
1595:.
1583:.
1579:.
1556:.
1546:.
1532:.
1528:.
1505:.
1493:31
1491:.
1487:.
1464:.
1454:.
1444:21
1442:.
1438:.
1415:.
1405:.
1395:42
1393:.
1389:.
1366:.
1358:.
1344:.
1340:.
1317:.
1309:.
1297:.
1270:.
1248:.
1240:.
1226:.
1222:.
1199:.
1189:.
1177:36
1175:.
1171:.
1148:.
1138:.
1130:.
1118:12
1116:.
1112:.
1089:.
1079:.
1071:.
1061:42
1059:.
1055:.
1031:.
1021:.
1013:.
1001:.
997:.
985:^
971:.
961:.
953:.
943:10
941:.
937:.
914:.
902:.
879:.
867:.
841:.
826:^
812:.
804:.
796:.
784:31
782:.
778:.
755:.
745:.
737:.
727:17
725:.
721:.
698:.
688:.
680:.
668:.
664:.
624:.
616:.
608:.
596:.
592:.
566:.
556:.
548:.
536:21
534:.
530:.
185:,
143:.
123:,
119:,
2158:.
2134::
2126::
2120:9
2099:.
2077::
1966:.
1944::
1917:.
1895::
1889:1
1868:.
1846::
1838::
1811:.
1779::
1748:.
1736::
1713:.
1699::
1672:.
1650::
1623:.
1599::
1591::
1585:8
1564:.
1540::
1534:9
1513:.
1499::
1472:.
1450::
1423:.
1401::
1374:.
1352::
1325:.
1305::
1299:4
1281:.
1256:.
1234::
1207:.
1183::
1156:.
1124::
1097:.
1067::
1039:.
1009::
1003:9
979:.
949::
922:.
910::
887:.
875::
852:.
820:.
790::
763:.
733::
706:.
676::
632:.
612::
604::
574:.
542::
20:)
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.