375:. In wild-type lambda, lysis occurs at about 50 min, releasing approximately 100 completed virions. The timing of lysis is determined by the holin and antiholin proteins, with the latter inhibiting the former. In overview, the holin protein accumulates in the cytoplasmic membrane until suddenly forming micron-scale holes, which triggers lysis. The endolysin R is released to the periplasm, where it attacks the peptidoglycan. The spanin proteins Rz and Rz1 accumulate in the cytoplasmic and outer membranes, respectively, and form a complex spanning the periplasm through the meshwork of the peptidoglycan. When the endolysin degrades the peptidoglycan, the spanin complexes are liberated and cause disruption of the outer membrane. Destruction of the peptidoglycan by the endolysin and disruption of the outer membrane by the spanin complex are both required for lysis in lambda infections.
682:
930:
918:
954:
942:
38:
307:
There are three classes of genes in the phage genome that regulate whether the lytic or lysogenic cycles will emerge. The first class is the immediate early genes, the second is the delayed early genes and the third is the late genes. The following refers to the well-studied temperate phage lambda of
116:
In the lytic cycle, the viral DNA exists as a separate free floating molecule within the bacterial cell, and replicates separately from the host bacterial DNA, whereas in the lysogenic cycle, the viral DNA is integrated into the host genome. This is the key difference between the lytic and lysogenic
175:
and (if present) the cell wall. The virus does so by either attaching to a receptor on the cell's surface or by simple mechanical force. The binding is due to electrostatic interactions and is influenced by pH and the presence of ions. The virus then releases its genetic material (either single- or
358:
Q-mediated turn-on of late transcription begins about 6–8 min after infection if the lytic pathway is chosen. More than 25 genes are expressed from the single late promoter, resulting in four parallel biosynthetic pathways. Three of the pathways are for production of the three components of the
359:
virion: the DNA-filled head, the tail, and the side tail fibers. The virions self-assemble from these components, with the first virion appearing at about 20 min after infection. The fourth pathway is for lysis. In lambda 5 proteins are involved in lysis: the holin and antiholin from gene
335:
expression. The lysis-lysogeny decision is mainly influenced by the competition between Cro and CII, resulting in the determination of whether or not sufficient CI repressor is made. If so, CI represses the early promoters and the infection is shunted into the lysogenic pathway. N is an
299:(water pressure) that can no longer be constrained by the cell wall. This releases progeny virions into the surrounding environment, where they can go on to infect other cells and another lytic cycle begins. The phage that causes lysis of the host is called a lytic or virulent phage.
386:, that inhibit the T4 holin, if the infected cell undergoes super-infection by another T4 (or closely related) virion. Repeated super-infection can cause the T4 infection to continue without lysis for hours, leading to accumulation of virions to levels 10-fold higher than normal.
204:
During the transcription and biosynthesis stages, the virus hijacks the cell's replication and translation mechanisms, using them to make more viruses. The virus's nucleic acid uses the host cell's metabolic machinery to make large amounts of viral components.
290:
About 25 minutes after initial infection, approximately 200 new virions (viral bodies) are formed. Once enough virions have matured and accumulated, specialized viral proteins are used to dissolve the cells' walls. The cell bursts (i.e. it undergoes
224:
transcribes the viral RNA into DNA, which is then transcribed again into RNA. Once the viral DNA has taken control it induces the host cell's machinery to synthesize viral DNA, protein and starts multiplying.
188:. At this stage the cell becomes infected and can also be targeted by the immune system. It is mostly aided by receptors on the surface of the cell. The sequence of events that occur during initiation of
125:
The lytic cycle is often broken into six-stages. The six stages are: attachment, penetration, transcription, biosynthesis, maturation, and lysis.
216:(mRNA) molecules that are then used to direct the cell's ribosomes. One of the first polypeptides to be translated destroys the host's DNA. In
848:
402:
Molineux, Ian J. (January 2006). "Fifty-three years since
Hershey and Chase; much ado about pressure but which pressure is it?".
606:
462:
78:
315:
Immediate early genes: These genes are expressed from promoters recognized by the host RNA polymerase, and include
156:– the replicated material assembles into fully formed viral phages (each made up of a head, a tail and tail fibers)
958:
599:
144:– the host cell's DNA is degraded and the cell's metabolism is directed to initiate phage biosynthesis
640:
132:– the phage attaches itself to the surface of the host cell in order to inject its DNA into the cell
946:
209:
110:
171:
To infect a host cell, the virus must first inject its own nucleic acid into the cell through the
985:
635:
327:. CII is a transcription factor that stimulates expression of the main lysogenic repressor gene,
229:
564:
841:
478:
Malys, N (2012). "Shine-Dalgarno sequence of bacteriophage T4: GAGG prevails in early genes".
592:
221:
17:
893:
888:
232:) regulated in three phases of mRNA production followed by a phase of protein production.
184:) into the cell. In some viruses this genetic material is circular and mimics a bacterial
162:- cell wall or membrane ruptures, disintegrating it and releasing the virus in the process
8:
883:
868:
630:
429:
549:
522:
351:, which encodes the anti-terminator responsible for transcription of all the late genes.
336:
anti-termination factor that is needed for the transcription of the delayed early genes.
257:
are not recognized any more but now recognize T4 middle proteins. For protein synthesis
117:
cycles. However, in both cases the virus/phage replicates using the host DNA machinery.
521:
Gummalla, Vimathi S.; Zhang, Yujie; Liao, Yen-Te; Wu, Vivian C. H. (21 February 2023).
503:
254:
138:– the phage injects its DNA into the host cell by penetrating through the cell membrane
980:
917:
858:
703:
495:
458:
250:
196:
ejection from the virion into the host cell (penetration), was reviewed by
Molineux.
105:
and its membrane. Bacteriophages that can only go through the lytic cycle are called
507:
934:
754:
690:
544:
539:
534:
487:
411:
296:
150:– the phage DNA replicates inside the cell, synthesizing new phage DNA and proteins
52:
764:
749:
744:
728:
172:
98:
31:
415:
922:
853:
821:
708:
666:
258:
242:
102:
491:
974:
873:
785:
718:
671:
213:
189:
94:
87:
836:
831:
795:
780:
759:
681:
615:
499:
246:
90:
826:
805:
698:
903:
878:
217:
584:
863:
800:
790:
713:
645:
106:
185:
898:
661:
281:
Structural proteins including those for the head and the tail.
619:
292:
101:. The lytic cycle results in the destruction of the infected
339:
Delayed early genes: These include the replication genes
523:"The Role of Temperate Phages in Bacterial Pathogenicity"
193:
181:
177:
67:
61:
37:
271:
Virus nucleic acid (DNA or RNA depending on virus type).
261:
subsequence GAGG dominates an early genes translation.
453:
Madigan, Michael T.; Martinko, John M., eds. (2006).
241:
Enzymes modify the host's transcriptional process by
220:(which inject an RNA strand), a unique enzyme called
79:
70:
245:. Amongst other modifications, virus T4 changes the
64:
58:
520:
55:
378:Lysis inhibition: T4-like phages have two genes,
199:
972:
192:infection, from adsorption (attachment) through
452:
302:
600:
565:"The Lytic Cycle of the T-Even Bacteriophage"
166:
607:
593:
548:
538:
849:Laboratory diagnosis of viral infections
401:
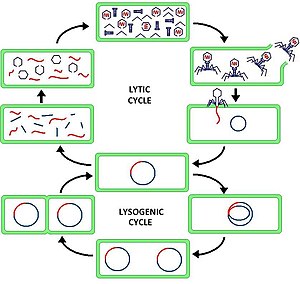
41:Lytic cycle, compared to lysogenic cycle
36:
614:
285:
14:
973:
588:
477:
941:
446:
208:In the case of DNA viruses, the DNA
953:
367:and the spanin proteins from genes
93:(referring to bacterial viruses or
24:
25:
997:
331:, whereas Cro is a repressor for
952:
940:
929:
928:
916:
680:
51:
455:Brock biology of microorganisms
557:
540:10.3390/microorganisms11030541
514:
471:
457:(11 ed.). Prentice Hall.
422:
395:
200:Transcription and biosynthesis
120:
86:) is one of the two cycles of
13:
1:
389:
430:"Ebola - Lytic or Lysogenic"
303:Gene regulation biochemistry
249:of the host by producing an
7:
416:10.1016/j.virol.2005.09.014
27:Cycle of viral reproduction
10:
1002:
363:, the endolysin from gene
228:The biosynthesis is (e.g.
167:Attachment and penetration
29:
912:
814:
773:
737:
689:
678:
654:
641:Social history of viruses
626:
492:10.1007/s11033-011-0707-4
480:Molecular Biology Reports
30:Not to be confused with
295:) due to high internal
109:phages (in contrast to
97:), the other being the
842:Helper dependent virus
42:
222:reverse transcriptase
40:
894:Virus quantification
889:Virus classification
286:Maturation and lysis
884:Virus-like particle
569:nemetoadreviews.com
43:
968:
967:
859:Neurotropic virus
704:Viral replication
464:978-0-13-144329-7
253:so that the host
251:anti-sigma factor
16:(Redirected from
993:
956:
955:
944:
943:
932:
931:
920:
755:Phenotype mixing
691:Viral life cycle
684:
609:
602:
595:
586:
585:
580:
579:
577:
575:
561:
555:
554:
552:
542:
518:
512:
511:
475:
469:
468:
450:
444:
443:
441:
440:
426:
420:
419:
399:
297:osmotic pressure
176:double-stranded
82:
77:
76:
73:
72:
69:
66:
63:
60:
57:
21:
1001:
1000:
996:
995:
994:
992:
991:
990:
971:
970:
969:
964:
908:
810:
769:
765:Viral evolution
750:Antigenic shift
745:Antigenic drift
733:
729:Lysogenic cycle
685:
676:
650:
622:
613:
583:
573:
571:
563:
562:
558:
519:
515:
476:
472:
465:
451:
447:
438:
436:
434:Ebola-Cases.com
428:
427:
423:
400:
396:
392:
305:
288:
202:
173:plasma membrane
169:
123:
99:lysogenic cycle
80:
54:
50:
35:
32:Lysogenic cycle
28:
23:
22:
15:
12:
11:
5:
999:
989:
988:
986:Bacteriophages
983:
966:
965:
963:
962:
950:
938:
926:
913:
910:
909:
907:
906:
901:
896:
891:
886:
881:
876:
871:
866:
861:
856:
854:Marine viruses
851:
846:
845:
844:
834:
829:
824:
822:Antiviral drug
818:
816:
812:
811:
809:
808:
803:
798:
793:
788:
783:
777:
775:
771:
770:
768:
767:
762:
757:
752:
747:
741:
739:
735:
734:
732:
731:
726:
721:
716:
711:
709:Viral shedding
706:
701:
695:
693:
687:
686:
679:
677:
675:
674:
669:
667:Viral envelope
664:
658:
656:
652:
651:
649:
648:
643:
638:
633:
627:
624:
623:
612:
611:
604:
597:
589:
582:
581:
556:
527:Microorganisms
513:
470:
463:
445:
421:
410:(1): 221–229.
393:
391:
388:
356:
355:
352:
337:
304:
301:
287:
284:
283:
282:
279:
277:
273:
272:
269:
267:
263:
262:
259:Shine-Dalgarno
243:RNA polymerase
239:
237:
201:
198:
168:
165:
164:
163:
157:
151:
145:
139:
133:
122:
119:
95:bacteriophages
26:
9:
6:
4:
3:
2:
998:
987:
984:
982:
979:
978:
976:
961:
960:
951:
949:
948:
939:
937:
936:
927:
925:
924:
919:
915:
914:
911:
905:
902:
900:
897:
895:
892:
890:
887:
885:
882:
880:
877:
875:
874:Viral disease
872:
870:
867:
865:
862:
860:
857:
855:
852:
850:
847:
843:
840:
839:
838:
835:
833:
830:
828:
825:
823:
820:
819:
817:
813:
807:
804:
802:
799:
797:
794:
792:
789:
787:
786:Bacteriophage
784:
782:
779:
778:
776:
772:
766:
763:
761:
758:
756:
753:
751:
748:
746:
743:
742:
740:
736:
730:
727:
725:
722:
720:
719:Virus latency
717:
715:
712:
710:
707:
705:
702:
700:
697:
696:
694:
692:
688:
683:
673:
672:Viral protein
670:
668:
665:
663:
660:
659:
657:
653:
647:
644:
642:
639:
637:
634:
632:
629:
628:
625:
621:
617:
610:
605:
603:
598:
596:
591:
590:
587:
570:
566:
560:
551:
546:
541:
536:
532:
528:
524:
517:
509:
505:
501:
497:
493:
489:
485:
481:
474:
466:
460:
456:
449:
435:
431:
425:
417:
413:
409:
405:
398:
394:
387:
385:
381:
376:
374:
370:
366:
362:
353:
350:
346:
342:
338:
334:
330:
326:
322:
318:
314:
313:
312:
311:
300:
298:
294:
280:
278:
275:
274:
270:
268:
265:
264:
260:
256:
252:
248:
244:
240:
238:
235:
234:
233:
231:
226:
223:
219:
215:
214:messenger RNA
211:
206:
197:
195:
191:
190:bacteriophage
187:
183:
179:
174:
161:
158:
155:
152:
149:
146:
143:
142:Transcription
140:
137:
134:
131:
128:
127:
126:
118:
114:
112:
108:
104:
100:
96:
92:
89:
85:
84:
75:
48:
39:
33:
19:
957:
945:
933:
921:
837:Viral vector
832:Helper virus
796:Human virome
781:Animal virus
760:Reassortment
723:
636:Introduction
616:Microbiology
572:. Retrieved
568:
559:
530:
526:
516:
483:
479:
473:
454:
448:
437:. Retrieved
433:
424:
407:
403:
397:
383:
379:
377:
372:
368:
364:
360:
357:
348:
344:
340:
332:
328:
324:
320:
316:
309:
306:
289:
266:Middle phase
247:sigma factor
227:
218:retroviruses
212:itself into
207:
203:
170:
159:
153:
148:Biosynthesis
147:
141:
135:
129:
124:
115:
91:reproduction
46:
44:
959:WikiProject
827:Giant virus
806:Plant virus
724:Lytic cycle
699:Viral entry
486:(1): 33–9.
354:Late genes:
236:Early phase
210:transcribes
136:Penetration
121:Description
47:lytic cycle
975:Categories
904:Virosphere
879:Viral load
869:Satellites
655:Components
574:January 9,
533:(3): 541.
439:2023-01-26
390:References
276:Late phase
154:Maturation
130:Attachment
864:Oncovirus
801:Mycovirus
791:Virophage
714:Viroplasm
347:and also
255:promotors
113:phages).
111:temperate
981:Virology
935:Category
738:Genetics
646:Virology
550:10052878
508:17854788
500:21533668
404:Virology
310:E. coli.
107:virulent
947:Commons
774:By host
631:History
186:plasmid
923:Portal
899:Virome
662:Capsid
547:
506:
498:
461:
323:, and
815:Other
620:Virus
504:S2CID
293:lysis
160:Lysis
88:viral
18:Lytic
576:2018
496:PMID
459:ISBN
384:rIII
382:and
371:and
343:and
103:cell
45:The
545:PMC
535:doi
488:doi
412:doi
408:344
373:Rz1
321:cII
317:Cro
194:DNA
182:DNA
180:or
178:RNA
83:-ik
81:LIT
977::
618::
567:.
543:.
531:11
529:.
525:.
502:.
494:.
484:39
482:.
432:.
406:.
380:rI
369:Rz
333:cI
329:cI
319:,
230:T4
608:e
601:t
594:v
578:.
553:.
537::
510:.
490::
467:.
442:.
418:.
414::
365:R
361:S
349:Q
345:P
341:O
325:N
74:/
71:k
68:ɪ
65:t
62:ɪ
59:l
56:ˈ
53:/
49:(
34:.
20:)
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.