213:, whereas permanent complexes have a relatively long half-life. Typically, the obligate interactions (protein–protein interactions in an obligate complex) are permanent, whereas non-obligate interactions have been found to be either permanent or transient. Note that there is no clear distinction between obligate and non-obligate interaction, rather there exist a continuum between them which depends on various conditions e.g. pH, protein concentration etc. However, there are important distinctions between the properties of transient and permanent/stable interactions: stable interactions are highly conserved but transient interactions are far less conserved, interacting proteins on the two sides of a stable interaction have more tendency of being co-expressed than those of a transient interaction (in fact, co-expression probability between two transiently interacting proteins is not higher than two random proteins), and transient interactions are much less co-localized than stable interactions. Though, transient by nature, transient interactions are very important for cell biology: the human interactome is enriched in such interactions, these interactions are the dominating players of gene regulation and signal transduction, and proteins with
369:
multimer. Genes that encode multimer-forming polypeptides appear to be common. One interpretation of the data is that polypeptide monomers are often aligned in the multimer in such a way that mutant polypeptides defective at nearby sites in the genetic map tend to form a mixed multimer that functions poorly, whereas mutant polypeptides defective at distant sites tend to form a mixed multimer that functions more effectively. The intermolecular forces likely responsible for self-recognition and multimer formation were discussed by Jehle.
246:
121:
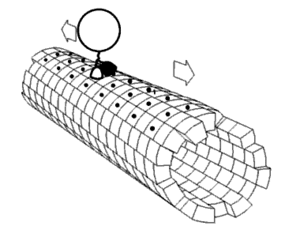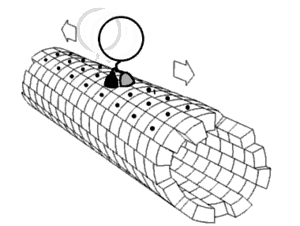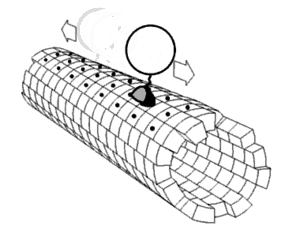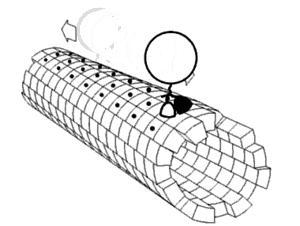
422:, which can identify different intermediate states simultaneously. This has led to the discovery that most complexes follow an ordered assembly pathway. In the cases where disordered assembly is possible, the change from an ordered to a disordered state leads to a transition from function to dysfunction of the complex, since disordered assembly leads to aggregation.
201:, then the complexes formed by such proteins are termed "non-obligate protein complexes". However, some proteins can't be found to create a stable well-folded structure alone, but can be found as a part of a protein complex which stabilizes the constituent proteins. Such protein complexes are called "obligate protein complexes".
267:". These authors also showed that complexes tend to be composed of either essential or non-essential proteins rather than showing a random distribution (see Figure). However, this not an all or nothing phenomenon: only about 26% (105/401) of yeast complexes consist of solely essential or solely nonessential subunits.
228:
have more than one structural form or dynamic structural disorder in the bound state. This means that proteins may not fold completely in either transient or permanent complexes. Consequently, specific complexes can have ambiguous interactions, which vary according to the environmental signals. Hence
262:
belong to protein complexes. This led to the conclusion that essentiality is a property of molecular machines (i.e. complexes) rather than individual components. Wang et al. (2009) noted that larger protein complexes are more likely to be essential, explaining why essential genes are more likely to
441:
proteins assemble in a way that mimics evolution. That is, an intermediate in the assembly process is present in the complex's evolutionary history. The opposite phenomenon is observed in heteromultimeric complexes, where gene fusion occurs in a manner that preserves the original assembly pathway.
368:
of the gene. Separately, the mutants were tested in pairwise combinations to measure complementation. An analysis of the results from such studies led to the conclusion that intragenic complementation, in general, arises from the interaction of differently defective polypeptide monomers to form a
968:
Yu, H; Braun, P; Yildirim, M. A.; Lemmens, I; Venkatesan, K; Sahalie, J; Hirozane-Kishikawa, T; Gebreab, F; Li, N; Simonis, N; Hao, T; Rual, J. F.; Dricot, A; Vazquez, A; Murray, R. R.; Simon, C; Tardivo, L; Tam, S; Svrzikapa, N; Fan, C; De Smet, A. S.; Motyl, A; Hudson, M. E.; Park, J; Xin, X;
425:
The structure of proteins play a role in how the multiprotein complex assembles. The interfaces between proteins can be used to predict assembly pathways. The intrinsic flexibility of proteins also plays a role: more flexible proteins allow for a greater surface area available for interaction.
31:
291:
in the plasma membrane of a neuron are heteromultimeric proteins composed of four of forty known alpha subunits. Subunits must be of the same subfamily to form the multimeric protein channel. The tertiary structure of the channel allows ions to flow through the hydrophobic plasma membrane.
96:
complex and substrates can be vastly improved, leading to higher cellular efficiency. Many of the techniques used to enter cells and isolate proteins are inherently disruptive to such large complexes, complicating the task of determining the components of a complex.
278:
The subunits of a multimeric protein may be identical as in a homomultimeric (homooligomeric) protein or different as in a heteromultimeric protein. Many soluble and membrane proteins form homomultimeric complexes in a cell, majority of proteins in the
417:
Proper assembly of multiprotein complexes is important, since misassembly can lead to disastrous consequences. In order to study pathway assembly, researchers look at intermediate steps in the pathway. One such technique that allows one to do that is
253:
Although some early studies suggested a strong correlation between essentiality and protein interaction degree (the "centrality-lethality" rule) subsequent analyses have shown that this correlation is weak for binary or transient interactions (e.g.,
1466:
Rodríguez-Pombo P, Pérez-Cerdá C, Pérez B, Desviat LR, Sánchez-Pulido L, Ugarte M. Towards a model to explain the intragenic complementation in the heteromultimeric protein propionyl-CoA carboxylase. Biochim
Biophys Acta. 2005;1740(3):489-498.
148:. Individual proteins can participate in a variety of protein complexes. Different complexes perform different functions, and the same complex can perform multiple functions depending on various factors. Factors include:
1517:
Raicu V, Stoneman MR, Fung R, Melnichuk M, Jansma DB, Pisterzi LF, Rath S, Fox, M, Wells, JW, Saldin DK (2008). "Determination of supramolecular structure and spatial distribution of protein complexes in living cells".
283:
are homomultimeric. Homooligomers are responsible for the diversity and specificity of many pathways, may mediate and regulate gene expression, activity of enzymes, ion channels, receptors, and cell adhesion processes.
89:. These complexes are a cornerstone of many (if not most) biological processes. The cell is seen to be composed of modular supramolecular complexes, each of which performs an independent, discrete biological function.
624:
Amoutzias G, Van de Peer Y (2010). "Single-Gene and Whole-Genome
Duplications and the Evolution of Protein–Protein Interaction Networks. Evolutionary genomics and systems biology". In Caetano-Anolles G (ed.).
1664:
Sudha, Govindarajan; Nussinov, Ruth; Srinivasan, Narayanaswamy (2014). "An overview of recent advances in structural bioinformatics of protein–protein interactions and a guide to their principles".
323:
of a particular gene, the mixed multimer may exhibit greater functional activity than the unmixed multimers formed by each of the mutants alone. In such a case, the phenomenon is referred to as
429:
While assembly is a different process from disassembly, the two are reversible in both homomeric and heteromeric complexes. Thus, the overall process can be referred to as (dis)assembly.
401:. Recently, Raicu and coworkers developed a method to determine the quaternary structure of protein complexes in living cells. This method is based on the determination of pixel-level
229:
different ensembles of structures result in different (even opposite) biological functions. Post-translational modifications, protein interactions or alternative splicing modulate the
217:(IDR: regions in protein that show dynamic inter-converting structures in the native state) are found to be enriched in transient regulatory and signaling interactions.
1249:
Lage, K; Karlberg, E. O.; Størling, Z. M.; Olason, P. I.; Pedersen, A. G.; Rigina, O; Hinsby, A. M.; Tümer, Z; Pociot, F; Tommerup, N; Moreau, Y; Brunak, S (2007).
327:(also called inter-allelic complementation). Intragenic complementation has been demonstrated in many different genes in a variety of organisms including the fungi
1450:
Smallwood S, Cevik B, Moyer SA. Intragenic complementation and oligomerization of the L subunit of the sendai virus RNA polymerase. Virology. 2002;304(2):235-245.
65:
2136:
1036:"Why do hubs in the yeast protein interaction network tend to be essential: Reexamining the connection between the network topology and essentiality"
1545:
144:
Protein complex formation can activate or inhibit one or more of the complex members and in this way, protein complex formation can be similar to
2028:
595:
316:
form a complex, this protein structure is referred to as a multimer. When a multimer is formed from polypeptides produced by two different
1147:
Wang, H; Kakaradov, B; Collins, S. R.; Karotki, L; Fiedler, D; Shales, M; Shokat, K. M.; Walther, T. C.; Krogan, N. J.; Koller, D (2009).
1516:
409:. The distribution of FRET efficiencies are simulated against different models to get the geometry and stoichiometry of the complexes.
233:
of fuzzy complexes, to fine-tune affinity or specificity of interactions. These mechanisms are often used for regulation within the
1841:
263:
have high co-complex interaction degree. Ryan et al. (2013) referred to the observation that entire complexes appear essential as "
324:
748:
Tompa P, Fuxreiter M (January 2008). "Fuzzy complexes: polymorphism and structural disorder in protein–protein interactions".
270:
In humans, genes whose protein products belong to the same complex are more likely to result in the same disease phenotype.
258:). However, the correlation is robust for networks of stable co-complex interactions. In fact, a disproportionate number of
17:
402:
108:. In stable complexes, large hydrophobic interfaces between proteins typically bury surface areas larger than 2500 square
2221:
2021:
42:
2114:
249:
Essential proteins in yeast complexes occur much less randomly than expected by chance. Modified after Ryan et al. 2013
489:
288:
2179:
1869:
86:
2184:
2156:
2129:
2107:
2092:
2057:
2014:
907:
Jeong, H; Mason, S. P.; Barabási, A. L.; Oltvai, Z. N. (2001). "Lethality and centrality in protein networks".
419:
2102:
1834:
76:
1500:
Jehle H. Intermolecular forces and biological specificity. Proc Natl Acad Sci U S A. 1963;50(3):516-524.
818:
Fuxreiter M, Simon I, Bondos S (August 2011). "Dynamic protein–DNA recognition: beyond what can be seen".
394:
2274:
390:
197:
If a protein can form a stable well-folded structure on its own (without any other associated protein)
184:
complexes. They compared 6000 yeast proteins to those from 2026 other fungi and 4325 other eukaryotes.
126:
1098:"A high-accuracy consensus map of yeast protein complexes reveals modular nature of gene essentiality"
341:
83:
2206:
1813:
1347:"Caught in self-interaction: evolutionary and functional mechanisms of protein homooligomerization"
386:
335:
234:
230:
164:
1809:
2238:
2216:
2196:
1827:
1483:
Crick FH, Orgel LE. The theory of inter-allelic complementation. J Mol Biol. 1964 Jan;8:161-5.
300:. A cluster of connexons forms the gap-junction in two neurons that transmit signals through an
172:
wide and the elucidation of most of its protein complexes is ongoing. In 2021, researchers used
2201:
2097:
397:
is also becoming available. One method that is commonly used for identifying the meomplexes is
1973:
1750:
Levy, Emmanuel D; Boeri Erba, Elisabetta; Robinson, Carol V; Teichmann, Sarah A (July 2008).
1539:
406:
382:
361:
360:
defective in the same gene were often isolated and mapped in a linear order on the basis of
1864:
1763:
1568:
1358:
1047:
1027:
982:
926:
456:
353:
1615:
Marsh JA, Hernández H, Hall Z, Ahnert SE, Perica T, Robinson CV, Teichmann SA (Apr 2013).
1251:"A human phenome-interactome network of protein complexes implicated in genetic disorders"
860:"All or nothing: Protein complexes flip essentiality between distantly related eukaryotes"
699:"Unequal evolutionary conservation of human protein interactions in interologous networks"
8:
2253:
2243:
2211:
398:
378:
1767:
1572:
1362:
1051:
986:
930:
245:
2248:
1784:
1751:
1727:
1700:
1641:
1616:
1592:
1428:
1403:
1379:
1370:
1346:
1327:
1278:
1226:
1199:
1175:
1148:
1124:
1097:
1070:
1035:
1003:
970:
950:
916:
884:
859:
725:
698:
572:
547:
301:
38:
1488:
674:
649:
1905:
1879:
1789:
1732:
1681:
1677:
1646:
1617:"Protein complexes are under evolutionary selection to assemble via ordered pathways"
1584:
1433:
1404:"Intragenic Complementation among Temperature Sensitive Mutants of Bacteriophage T4D"
1384:
1319:
1314:
1297:
1270:
1231:
1180:
1129:
1075:
1008:
942:
889:
835:
800:
765:
730:
679:
577:
528:
485:
329:
280:
61:
1331:
1250:
168:(yeast). For this relatively simple organism, the study of protein complexes is now
1983:
1941:
1936:
1931:
1779:
1771:
1722:
1712:
1673:
1636:
1628:
1596:
1576:
1527:
1501:
1484:
1468:
1451:
1423:
1415:
1374:
1366:
1309:
1282:
1262:
1221:
1211:
1170:
1160:
1119:
1109:
1065:
1055:
1031:
998:
990:
934:
879:
871:
827:
792:
757:
720:
710:
669:
661:
630:
596:"AI cracks the code of protein complexes—providing a road map for new drug targets"
567:
559:
518:
255:
137:
46:
954:
2006:
1889:
1874:
1717:
1472:
1060:
783:
Fuxreiter M (January 2012). "Fuzziness: linking regulation to protein dynamics".
548:"The origins and evolution of functional modules: lessons from protein complexes"
461:
259:
145:
1419:
482:
Fundamentals of
Enzymology: The Cell and Molecular Biology of Catalytic Proteins
2189:
2072:
2049:
1910:
1632:
1165:
831:
761:
634:
451:
162:
Many protein complexes are well understood, particularly in the model organism
105:
1216:
971:"High-quality binary protein interaction map of the yeast interactome network"
2268:
2084:
2067:
1968:
1963:
1701:"Protein flexibility facilitates quaternary structure assembly and evolution"
1531:
432:
225:
173:
109:
92:
Through proximity, the speed and selectivity of binding interactions between
1149:"A complex-based reconstruction of the Saccharomyces cerevisiae interactome"
1114:
994:
715:
1793:
1736:
1685:
1650:
1588:
1455:
1437:
1388:
1323:
1274:
1235:
1184:
1133:
1079:
1012:
946:
893:
839:
804:
769:
734:
683:
665:
581:
563:
532:
365:
130:
1505:
381:
of protein complexes can be determined by experimental techniques such as
2062:
1988:
1978:
1559:
Dobson, Christopher M (December 2003). "Protein folding and misfolding".
921:
875:
69:
1775:
1580:
1200:"Using protein complexes to predict phenotypic effects of gene mutation"
2174:
2151:
1915:
796:
347:
120:
101:
1993:
938:
523:
506:
438:
296:
are an example of a homomultimeric protein composed of six identical
181:
177:
1266:
1946:
1884:
357:
297:
293:
969:
Cusick, M. E.; Moore, T; Boone, C; Snyder, M; Roth, F. P. (2008).
30:
1956:
1951:
1850:
1025:
133:
79:
34:
505:
Hartwell LH, Hopfield JJ, Leibler S, Murray AW (December 1999).
204:
2146:
2141:
2037:
1749:
320:
317:
273:
169:
93:
2231:
2226:
1344:
858:
Ryan, C. J.; Krogan, N. J.; Cunningham, P; Cagney, G (2013).
504:
209:
Transient protein complexes form and break down transiently
2124:
2119:
1819:
1146:
313:
192:
1345:
Hashimoto K, Nishi H, Bryant S, Panchenko AR (June 2011).
1248:
857:
433:
Evolutionary significance of multiprotein complex assembly
405:(FRET) efficiency in conjunction with spectrally resolved
240:
545:
1614:
906:
1663:
1401:
967:
546:
Pereira-Leal JB, Levy ED, Teichmann SA (March 2006).
356:, an RNA virus and humans. In such studies, numerous
2137:
Branched-chain alpha-keto acid dehydrogenase complex
817:
623:
2239:
Phosphoenolpyruvate sugar phosphotransferase system
1402:Bernstein, H; Edgar, RS; Denhardt, GH (June 1965).
312:When multiple copies of a polypeptide encoded by a
2036:
1752:"Assembly reflects evolution of protein complexes"
484:(3rd ed.). Oxford: Oxford University Press.
2266:
619:
617:
615:
2058:Photosynthetic reaction center complex proteins
1610:
1608:
1606:
1095:
1699:Marsh, Joseph; Teichmann, Sarah A (May 2014).
1698:
747:
647:
2022:
1835:
1197:
1096:Hart, G. T.; Lee, I; Marcotte, E. R. (2007).
696:
612:
307:
205:Transient vs permanent/stable protein complex
187:
1666:Progress in Biophysics and Molecular Biology
1603:
1544:: CS1 maint: multiple names: authors list (
1338:
811:
479:
274:Homomultimeric and heteromultimeric proteins
1091:
1089:
1019:
853:
851:
849:
782:
776:
741:
690:
641:
539:
498:
2029:
2015:
1842:
1828:
1295:
1191:
372:
100:Examples of protein complexes include the
1812:at the U.S. National Library of Medicine
1783:
1726:
1716:
1640:
1427:
1378:
1313:
1225:
1215:
1174:
1164:
1123:
1113:
1069:
1059:
1002:
920:
900:
883:
724:
714:
673:
571:
522:
393:. Increasingly the theoretical option of
72:are found in a single polypeptide chain.
1298:"The modular nature of genetic diseases"
1242:
1140:
1086:
846:
552:Philos. Trans. R. Soc. Lond. B Biol. Sci
507:"From molecular to modular cell biology"
244:
193:Obligate vs non-obligate protein complex
119:
37:is a protein functioning as a molecular
29:
1289:
241:Essential proteins in protein complexes
14:
2267:
1558:
1198:Fraser, H. B.; Plotkin, J. B. (2007).
961:
64:. Protein complexes are distinct from
2010:
1823:
60:is a group of two or more associated
27:Type of stable macromolecular complex
648:Nooren IM, Thornton JM (July 2003).
2222:Mitochondrial trifunctional protein
1153:Molecular & Cellular Proteomics
650:"Diversity of protein interactions"
104:for molecular degradation and most
82:in a protein complex are linked by
24:
25:
2286:
1803:
437:In homomultimeric complexes, the
403:Förster resonance energy transfer
1678:10.1016/j.pbiomolbio.2014.07.004
1315:10.1111/j.1399-0004.2006.00708.x
289:voltage-gated potassium channels
220:
215:intrinsically disordered regions
176:software RoseTTAFold along with
75:Protein complexes are a form of
2180:Carbamoyl phosphate synthase II
1870:Post-translational modification
1743:
1692:
1657:
1552:
1510:
1494:
1477:
1460:
1444:
1395:
1296:Oti, M; Brunner, H. G. (2007).
180:to solve the structures of 712
2185:Aspartate carbamoyltransferase
2093:Pyruvate dehydrogenase complex
588:
473:
420:electrospray mass spectrometry
13:
1:
2217:Glycine decarboxylase complex
2212:Fatty acid synthetase complex
1489:10.1016/s0022-2836(64)80156-x
697:Brown KR, Jurisica I (2007).
467:
1849:
1718:10.1371/journal.pbio.1001870
1473:10.1016/j.bbadis.2004.10.009
1371:10.1088/1478-3975/8/3/035007
1061:10.1371/journal.pcbi.1000140
864:Genome Biology and Evolution
480:Price NC, Stevens L (1999).
87:protein–protein interactions
7:
445:
412:
115:
10:
2291:
2249:Sucrase-isomaltase complex
2115:Oxoglutarate dehydrogenase
1911:Protein structural domains
1633:10.1016/j.cell.2013.02.044
1166:10.1074/mcp.M800490-MCP200
1040:PLOS Computational Biology
832:10.1016/j.tibs.2011.04.006
762:10.1016/j.tibs.2007.10.003
635:10.1002/9780470570418.ch19
391:nuclear magnetic resonance
325:intragenic complementation
308:Intragenic complementation
188:Types of protein complexes
127:Bacillus amyloliquefaciens
2167:
2083:
2048:
1924:
1898:
1857:
1420:10.1093/genetics/51.6.987
1217:10.1186/gb-2007-8-11-r252
342:Schizosaccharomyces pombe
152:Cell compartment location
2207:Electron transport chain
1814:Medical Subject Headings
1532:10.1038/nphoton.2008.291
387:Single particle analysis
336:Saccharomyces cerevisiae
235:eukaryotic transcription
231:conformational ensembles
165:Saccharomyces cerevisiae
2197:P450-containing systems
1115:10.1186/1471-2105-8-236
1026:Zotenko, E; Mestre, J;
995:10.1126/science.1158684
716:10.1186/gb-2007-8-5-r95
395:protein–protein docking
373:Structure determination
226:Fuzzy protein complexes
158:Cell nutritional status
43:protein domain dynamics
2202:Cytochrome b6f complex
1810:Multiprotein+Complexes
1456:10.1006/viro.2002.1720
564:10.1098/rstb.2005.1807
517:(6761 Suppl): C47–52.
364:frequencies to form a
250:
141:
49:
2042:multienzyme complexes
1974:Photoreceptor protein
1506:10.1073/pnas.50.3.516
627:Evolutionary Genomics
407:two-photon microscope
383:X-ray crystallography
248:
123:
77:quaternary structure.
33:
1865:Protein biosynthesis
1255:Nature Biotechnology
666:10.1093/emboj/cdg359
629:. pp. 413–429.
457:Biomolecular complex
265:modular essentiality
68:, in which multiple
58:multiprotein complex
18:Multiprotein complex
2254:Tryptophan synthase
2244:Polyketide synthase
1776:10.1038/nature06942
1768:2008Natur.453.1262L
1581:10.1038/nature02261
1573:2003Natur.426..884D
1363:2011PhBio...8c5007H
1052:2008PLSCB...4E0140Z
987:2008Sci...322..104Y
931:2001Natur.411...41J
820:Trends Biochem. Sci
750:Trends Biochem. Sci
399:immunoprecipitation
379:molecular structure
140:(blue) in a complex
66:multidomain enzymes
1102:BMC Bioinformatics
876:10.1093/gbe/evt074
797:10.1039/c1mb05234a
302:electrical synapse
251:
142:
136:(colored) and its
62:polypeptide chains
50:
39:biological machine
2275:Protein complexes
2262:
2261:
2004:
2003:
1906:Protein structure
1880:Protein targeting
1302:Clinical Genetics
330:Neurospora crassa
281:Protein Data Bank
70:catalytic domains
16:(Redirected from
2282:
2031:
2024:
2017:
2008:
2007:
1984:Phycobiliprotein
1942:Globular protein
1937:Membrane protein
1932:List of proteins
1844:
1837:
1830:
1821:
1820:
1798:
1797:
1787:
1762:(7199): 1262–5.
1747:
1741:
1740:
1730:
1720:
1696:
1690:
1689:
1661:
1655:
1654:
1644:
1612:
1601:
1600:
1567:(6968): 884–90.
1556:
1550:
1549:
1543:
1535:
1520:Nature Photonics
1514:
1508:
1498:
1492:
1481:
1475:
1464:
1458:
1448:
1442:
1441:
1431:
1399:
1393:
1392:
1382:
1342:
1336:
1335:
1317:
1293:
1287:
1286:
1246:
1240:
1239:
1229:
1219:
1195:
1189:
1188:
1178:
1168:
1144:
1138:
1137:
1127:
1117:
1093:
1084:
1083:
1073:
1063:
1032:Przytycka, T. M.
1023:
1017:
1016:
1006:
981:(5898): 104–10.
965:
959:
958:
939:10.1038/35075138
924:
922:cond-mat/0105306
904:
898:
897:
887:
855:
844:
843:
815:
809:
808:
780:
774:
773:
745:
739:
738:
728:
718:
694:
688:
687:
677:
645:
639:
638:
621:
610:
609:
607:
606:
592:
586:
585:
575:
558:(1467): 507–17.
543:
537:
536:
526:
524:10.1038/35011540
502:
496:
495:
477:
354:bacteriophage T4
345:; the bacterium
256:yeast two-hybrid
155:Cell cycle stage
21:
2290:
2289:
2285:
2284:
2283:
2281:
2280:
2279:
2265:
2264:
2263:
2258:
2163:
2079:
2044:
2035:
2005:
2000:
1964:Fibrous protein
1920:
1894:
1890:Protein methods
1875:Protein folding
1853:
1848:
1806:
1801:
1748:
1744:
1711:(5): e1001870.
1697:
1693:
1672:(2–3): 141–50.
1662:
1658:
1613:
1604:
1557:
1553:
1537:
1536:
1515:
1511:
1499:
1495:
1491:. PMID 14149958
1482:
1478:
1465:
1461:
1449:
1445:
1414:(6): 987–1002.
1400:
1396:
1343:
1339:
1294:
1290:
1267:10.1038/nbt1295
1247:
1243:
1196:
1192:
1145:
1141:
1094:
1087:
1046:(8): e1000140.
1024:
1020:
966:
962:
905:
901:
856:
847:
816:
812:
781:
777:
746:
742:
695:
691:
660:(14): 3486–92.
646:
642:
622:
613:
604:
602:
600:www.science.org
594:
593:
589:
544:
540:
503:
499:
492:
478:
474:
470:
462:Protein subunit
448:
435:
415:
375:
310:
276:
260:essential genes
243:
223:
207:
195:
190:
146:phosphorylation
118:
106:RNA polymerases
54:protein complex
28:
23:
22:
15:
12:
11:
5:
2288:
2278:
2277:
2260:
2259:
2257:
2256:
2251:
2246:
2241:
2236:
2235:
2234:
2229:
2219:
2214:
2209:
2204:
2199:
2194:
2193:
2192:
2190:Dihydroorotase
2187:
2182:
2171:
2169:
2165:
2164:
2162:
2161:
2160:
2159:
2154:
2149:
2144:
2134:
2133:
2132:
2127:
2122:
2112:
2111:
2110:
2105:
2100:
2089:
2087:
2081:
2080:
2078:
2077:
2076:
2075:
2070:
2060:
2054:
2052:
2050:Photosynthesis
2046:
2045:
2034:
2033:
2026:
2019:
2011:
2002:
2001:
1999:
1998:
1997:
1996:
1991:
1986:
1976:
1971:
1966:
1961:
1960:
1959:
1954:
1949:
1939:
1934:
1928:
1926:
1922:
1921:
1919:
1918:
1913:
1908:
1902:
1900:
1896:
1895:
1893:
1892:
1887:
1882:
1877:
1872:
1867:
1861:
1859:
1855:
1854:
1847:
1846:
1839:
1832:
1824:
1818:
1817:
1805:
1804:External links
1802:
1800:
1799:
1742:
1691:
1656:
1627:(2): 461–470.
1602:
1551:
1526:(2): 107–113.
1509:
1493:
1476:
1459:
1443:
1394:
1337:
1288:
1241:
1204:Genome Biology
1190:
1159:(6): 1361–81.
1139:
1085:
1028:O'Leary, D. P.
1018:
960:
915:(6833): 41–2.
899:
870:(6): 1049–59.
845:
810:
775:
740:
689:
640:
611:
587:
538:
497:
490:
471:
469:
466:
465:
464:
459:
454:
452:Heterotetramer
447:
444:
434:
431:
414:
411:
374:
371:
309:
306:
275:
272:
242:
239:
222:
219:
206:
203:
194:
191:
189:
186:
160:
159:
156:
153:
117:
114:
26:
9:
6:
4:
3:
2:
2287:
2276:
2273:
2272:
2270:
2255:
2252:
2250:
2247:
2245:
2242:
2240:
2237:
2233:
2230:
2228:
2225:
2224:
2223:
2220:
2218:
2215:
2213:
2210:
2208:
2205:
2203:
2200:
2198:
2195:
2191:
2188:
2186:
2183:
2181:
2178:
2177:
2176:
2173:
2172:
2170:
2166:
2158:
2155:
2153:
2150:
2148:
2145:
2143:
2140:
2139:
2138:
2135:
2131:
2128:
2126:
2123:
2121:
2118:
2117:
2116:
2113:
2109:
2106:
2104:
2101:
2099:
2096:
2095:
2094:
2091:
2090:
2088:
2086:
2085:Dehydrogenase
2082:
2074:
2071:
2069:
2066:
2065:
2064:
2061:
2059:
2056:
2055:
2053:
2051:
2047:
2043:
2039:
2032:
2027:
2025:
2020:
2018:
2013:
2012:
2009:
1995:
1992:
1990:
1987:
1985:
1982:
1981:
1980:
1977:
1975:
1972:
1970:
1969:Chromoprotein
1967:
1965:
1962:
1958:
1955:
1953:
1950:
1948:
1945:
1944:
1943:
1940:
1938:
1935:
1933:
1930:
1929:
1927:
1923:
1917:
1914:
1912:
1909:
1907:
1904:
1903:
1901:
1897:
1891:
1888:
1886:
1883:
1881:
1878:
1876:
1873:
1871:
1868:
1866:
1863:
1862:
1860:
1856:
1852:
1845:
1840:
1838:
1833:
1831:
1826:
1825:
1822:
1815:
1811:
1808:
1807:
1795:
1791:
1786:
1781:
1777:
1773:
1769:
1765:
1761:
1757:
1753:
1746:
1738:
1734:
1729:
1724:
1719:
1714:
1710:
1706:
1702:
1695:
1687:
1683:
1679:
1675:
1671:
1667:
1660:
1652:
1648:
1643:
1638:
1634:
1630:
1626:
1622:
1618:
1611:
1609:
1607:
1598:
1594:
1590:
1586:
1582:
1578:
1574:
1570:
1566:
1562:
1555:
1547:
1541:
1533:
1529:
1525:
1521:
1513:
1507:
1503:
1497:
1490:
1486:
1480:
1474:
1470:
1463:
1457:
1453:
1447:
1439:
1435:
1430:
1425:
1421:
1417:
1413:
1409:
1405:
1398:
1390:
1386:
1381:
1376:
1372:
1368:
1364:
1360:
1357:(3): 035007.
1356:
1352:
1348:
1341:
1333:
1329:
1325:
1321:
1316:
1311:
1307:
1303:
1299:
1292:
1284:
1280:
1276:
1272:
1268:
1264:
1261:(3): 309–16.
1260:
1256:
1252:
1245:
1237:
1233:
1228:
1223:
1218:
1213:
1209:
1205:
1201:
1194:
1186:
1182:
1177:
1172:
1167:
1162:
1158:
1154:
1150:
1143:
1135:
1131:
1126:
1121:
1116:
1111:
1107:
1103:
1099:
1092:
1090:
1081:
1077:
1072:
1067:
1062:
1057:
1053:
1049:
1045:
1041:
1037:
1033:
1029:
1022:
1014:
1010:
1005:
1000:
996:
992:
988:
984:
980:
976:
972:
964:
956:
952:
948:
944:
940:
936:
932:
928:
923:
918:
914:
910:
903:
895:
891:
886:
881:
877:
873:
869:
865:
861:
854:
852:
850:
841:
837:
833:
829:
826:(8): 415–23.
825:
821:
814:
806:
802:
798:
794:
791:(1): 168–77.
790:
786:
779:
771:
767:
763:
759:
755:
751:
744:
736:
732:
727:
722:
717:
712:
708:
704:
700:
693:
685:
681:
676:
671:
667:
663:
659:
655:
651:
644:
636:
632:
628:
620:
618:
616:
601:
597:
591:
583:
579:
574:
569:
565:
561:
557:
553:
549:
542:
534:
530:
525:
520:
516:
512:
508:
501:
493:
491:0-19-850229-X
487:
483:
476:
472:
463:
460:
458:
455:
453:
450:
449:
443:
440:
430:
427:
423:
421:
410:
408:
404:
400:
396:
392:
388:
384:
380:
370:
367:
363:
362:recombination
359:
355:
351:
349:
344:
343:
338:
337:
332:
331:
326:
322:
319:
315:
305:
303:
299:
295:
290:
285:
282:
271:
268:
266:
261:
257:
247:
238:
236:
232:
227:
221:Fuzzy complex
218:
216:
212:
202:
200:
185:
183:
179:
175:
174:deep learning
171:
167:
166:
157:
154:
151:
150:
149:
147:
139:
135:
132:
129:
128:
122:
113:
111:
107:
103:
98:
95:
90:
88:
85:
81:
78:
73:
71:
67:
63:
59:
55:
48:
44:
40:
36:
32:
19:
2041:
1759:
1755:
1745:
1708:
1705:PLOS Biology
1704:
1694:
1669:
1665:
1659:
1624:
1620:
1564:
1560:
1554:
1540:cite journal
1523:
1519:
1512:
1496:
1479:
1462:
1446:
1411:
1407:
1397:
1354:
1350:
1340:
1305:
1301:
1291:
1258:
1254:
1244:
1210:(11): R252.
1207:
1203:
1193:
1156:
1152:
1142:
1105:
1101:
1043:
1039:
1021:
978:
974:
963:
912:
908:
902:
867:
863:
823:
819:
813:
788:
784:
778:
753:
749:
743:
706:
702:
692:
657:
653:
643:
626:
603:. Retrieved
599:
590:
555:
551:
541:
514:
510:
500:
481:
475:
436:
428:
424:
416:
376:
352:; the virus
346:
340:
334:
328:
311:
286:
277:
269:
264:
252:
224:
214:
210:
208:
198:
196:
163:
161:
143:
131:ribonuclease
125:
99:
91:
84:non-covalent
74:
57:
53:
51:
2063:Photosystem
1989:Phytochrome
1979:Biliprotein
1308:(1): 1–11.
785:Mol Biosyst
703:Genome Biol
366:genetic map
350:typhimurium
237:machinery.
1916:Proteasome
1899:Structures
756:(1): 2–8.
709:(5): R95.
605:2021-11-14
468:References
348:Salmonella
102:proteasome
47:nanoscales
41:. It uses
1994:Lipocalin
1858:Processes
1351:Phys Biol
439:homomeric
358:mutations
298:connexins
294:Connexons
182:eukaryote
178:AlphaFold
138:inhibitor
94:enzymatic
2269:Category
1947:Globulin
1885:Proteome
1851:Proteins
1794:18563089
1737:24866000
1686:25077409
1651:23582331
1589:14685248
1438:14337770
1408:Genetics
1389:21572178
1332:24615025
1324:17204041
1275:17344885
1236:18042286
1185:19176519
1134:17605818
1080:18670624
1034:(2008).
1013:18719252
947:11333967
894:23661563
840:21620710
805:21927770
770:18054235
735:17535438
684:12853464
582:16524839
533:10591225
446:See also
413:Assembly
116:Function
80:Proteins
2038:Enzymes
1957:Albumin
1952:Edestin
1785:2658002
1764:Bibcode
1728:4035275
1642:4009401
1597:1036192
1569:Bibcode
1429:1210828
1380:3148176
1359:Bibcode
1283:5691546
1227:2258176
1176:2690481
1125:1940025
1108:: 236.
1071:2467474
1048:Bibcode
1004:2746753
983:Bibcode
975:Science
927:Bibcode
885:3698920
726:1929159
573:1609335
321:alleles
211:in vivo
199:in vivo
134:barnase
35:Kinesin
2147:BCKDHB
2142:BCKDHA
1816:(MeSH)
1792:
1782:
1756:Nature
1735:
1725:
1684:
1649:
1639:
1595:
1587:
1561:Nature
1436:
1426:
1387:
1377:
1330:
1322:
1281:
1273:
1234:
1224:
1183:
1173:
1132:
1122:
1078:
1068:
1011:
1001:
955:258942
953:
945:
909:Nature
892:
882:
838:
803:
768:
733:
723:
682:
675:165629
672:
654:EMBO J
580:
570:
531:
511:Nature
488:
318:mutant
170:genome
2232:HADHB
2227:HADHA
2168:Other
1925:Types
1593:S2CID
1328:S2CID
1279:S2CID
951:S2CID
917:arXiv
2125:DLST
2120:OGDH
1790:PMID
1733:PMID
1682:PMID
1647:PMID
1621:Cell
1585:PMID
1546:link
1434:PMID
1385:PMID
1320:PMID
1271:PMID
1232:PMID
1181:PMID
1130:PMID
1076:PMID
1009:PMID
943:PMID
890:PMID
836:PMID
801:PMID
766:PMID
731:PMID
680:PMID
578:PMID
529:PMID
486:ISBN
377:The
339:and
314:gene
287:The
124:The
2175:CAD
2157:DLD
2152:DBT
2130:DLD
1780:PMC
1772:doi
1760:453
1723:PMC
1713:doi
1674:doi
1670:116
1637:PMC
1629:doi
1625:153
1577:doi
1565:426
1528:doi
1502:doi
1485:doi
1469:doi
1452:doi
1424:PMC
1416:doi
1375:PMC
1367:doi
1310:doi
1263:doi
1222:PMC
1212:doi
1171:PMC
1161:doi
1120:PMC
1110:doi
1066:PMC
1056:doi
999:PMC
991:doi
979:322
935:doi
913:411
880:PMC
872:doi
828:doi
793:doi
758:doi
721:PMC
711:doi
670:PMC
662:doi
631:doi
568:PMC
560:doi
556:361
519:doi
515:402
389:or
56:or
45:on
2271::
2108:E3
2103:E2
2098:E1
2073:II
2040::
1788:.
1778:.
1770:.
1758:.
1754:.
1731:.
1721:.
1709:12
1707:.
1703:.
1680:.
1668:.
1645:.
1635:.
1623:.
1619:.
1605:^
1591:.
1583:.
1575:.
1563:.
1542:}}
1538:{{
1522:.
1432:.
1422:.
1412:51
1410:.
1406:.
1383:.
1373:.
1365:.
1353:.
1349:.
1326:.
1318:.
1306:71
1304:.
1300:.
1277:.
1269:.
1259:25
1257:.
1253:.
1230:.
1220:.
1206:.
1202:.
1179:.
1169:.
1155:.
1151:.
1128:.
1118:.
1104:.
1100:.
1088:^
1074:.
1064:.
1054:.
1042:.
1038:.
1030:;
1007:.
997:.
989:.
977:.
973:.
949:.
941:.
933:.
925:.
911:.
888:.
878:.
866:.
862:.
848:^
834:.
824:36
822:.
799:.
787:.
764:.
754:33
752:.
729:.
719:.
705:.
701:.
678:.
668:.
658:22
656:.
652:.
614:^
598:.
576:.
566:.
554:.
550:.
527:.
513:.
509:.
385:,
333:,
304:.
112:.
110:Ås
52:A
2068:I
2030:e
2023:t
2016:v
1843:e
1836:t
1829:v
1796:.
1774::
1766::
1739:.
1715::
1688:.
1676::
1653:.
1631::
1599:.
1579::
1571::
1548:)
1534:.
1530::
1524:3
1504::
1487::
1471::
1454::
1440:.
1418::
1391:.
1369::
1361::
1355:8
1334:.
1312::
1285:.
1265::
1238:.
1214::
1208:8
1187:.
1163::
1157:8
1136:.
1112::
1106:8
1082:.
1058::
1050::
1044:4
1015:.
993::
985::
957:.
937::
929::
919::
896:.
874::
868:5
842:.
830::
807:.
795::
789:8
772:.
760::
737:.
713::
707:8
686:.
664::
637:.
633::
608:.
584:.
562::
535:.
521::
494:.
20:)
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.