41:, also known as next-generation sequencing. Such an advantage has critical implications for both genome science and the study of biology in general. However, third generation sequencing data have much higher error rates than previous technologies, which can complicate downstream genome assembly and analysis of the resulting data. These technologies are undergoing active development and it is expected that there will be improvements to the high error rates. For applications that are more tolerant to error rates, such as structural variant calling, third generation sequencing has been found to outperform existing methods, even at a low depth of sequencing coverage.
142:
486:(AS) is the process by which a single gene may give rise to multiple distinct mRNA transcripts and consequently different protein translations. Some evidence suggests that AS is a ubiquitous phenomenon and may play a key role in determining the phenotypes of organisms, especially in complex eukaryotes; all eukaryotes contain genes consisting of introns that may undergo AS. In particular, it has been estimated that AS occurs in 95% of all human multi-exon genes. AS has undeniable potential to influence myriad biological processes. Advancing knowledge in this area has critical implications for the study of biology in general.
168:
448:
multiple overlapping reads are hard to obtain, this further leads to accuracy problems of downstream DNA modification detection. Both the hidden Markov model and statistical methods used with MinION raw data require repeated observations of DNA modifications for detection, meaning that individual modified nucleotides need to be consistently present in multiple copies of the genome, e.g. in multiple cells or plasmids in the sample.
495:
complicated by the highly variable expression levels across transcripts, and consequently variable read coverages across the sequence of the gene. In addition, exons may be shared among individual transcripts, rendering unambiguous inferences essentially impossible. Existing computational methods make inferences based on the accumulation of short reads at various sequence locations often by making simplifying assumptions.
154:. This sequencing machine is roughly the size of a regular USB flash drive and can be used readily by connecting to a laptop. In addition, since the sequencing process is not parallelized across regions of the genome, data could be collected and analyzed in real time. These advantages of third generation sequencing may be well-suited in hospital settings where quick and on-site data collection and analysis is demanded.
230:
PacBio’s single molecular and real time sequencing technology, the DNA polymerase molecule becomes increasingly damaged as the sequencing process occurs. Additionally, since the process happens quickly, the signals given off by individual bases may be blurred by signals from neighbouring bases. This poses a new computational challenge for deciphering the signals and consequently inferring the sequence. Methods such as
361:
513:
longer read lengths. Pacific
Bioscience has introduced the iso-seq platform, proposing to sequence mRNA molecules at their full lengths. It is anticipated that Oxford Nanopore will put forth similar technologies. The trouble with higher error rates may be alleviated by supplementary high quality short reads. This approach has been previously tested and reported to reduce the error rate by more than 3 folds.
294:
547:(EBOV) read was sequenced 44 seconds after data acquisition. There was uniform mapping of reads to genome; at least one read mapped to >88% of the genome. The relatively long reads allowed for sequencing of a near-complete viral genome to high accuracy (97–99% identity) directly from a primary clinical sample.
508:
genes. In comparison, transcript identification sensitivity decreases to 65%. For human, the study reported an exon detection sensitivity averaging to 69% and transcript detection sensitivity had an average of a mere 33%. In other words, for human, existing methods are able to identify less than half
503:
A study published in 2008 surveyed 25 different existing transcript reconstruction protocols. Its evidence suggested that existing methods are generally weak in assembling transcripts, though the ability to detect individual exons are relatively intact. According to the estimates, average sensitivity
499:
takes a parsimonious approach, seeking to explain all the reads with the fewest possible number of transcripts. On the other hand, StringTie attempts to simultaneously estimate transcript abundances while assembling the reads. These methods, while reasonable, may not always identify real transcripts.
494:
The current generation of sequencing technologies produce only short reads, putting tremendous limitation on the ability to detect distinct transcripts; short reads must be reverse engineered into original transcripts that could have given rise to the resulting read observations. This task is further
237:
On average, different individuals of the human population share about 99.9% of their genes. In other words, approximately only one out of every thousand bases would differ between any two person. The high error rates involved with third generation sequencing are inevitably problematic for the purpose
451:
For the PacBio platform, too, depending on what methylation you expect to find, coverage needs can vary. As of March 2017, other epigenetic factors like histone modifications have not been discoverable using third-generation technologies. Longer patterns of methylation are often lost because smaller
280:
Given the short reads produced by the current generation of sequencing technologies, de novo assembly is a major computational problem. It is normally approached by an iterative process of finding and connecting sequence reads with sensible overlaps. Various computational and statistical techniques,
512:
Third generation sequencing technologies have demonstrated promising prospects in solving the problem of transcript detection as well as mRNA abundance estimation at the level of transcripts. While error rates remain high, third generation sequencing technologies have the capability to produce much
311:
Third generation sequencing may also be used in conjunction with second generation sequencing. This approach is often referred to as hybrid sequencing. For example, long reads from third generation sequencing may be used to resolve ambiguities that exist in genomes previously assembled using second
149:
Other important advantages of third generation sequencing technologies include portability and sequencing speed. Since minimal sample preprocessing is required in comparison to second generation sequencing, smaller equipments could be designed. Oxford
Nanopore Technology has recently commercialized
474:
While expression levels can be more or less accurately depicted by second generation sequencing (we can assume that actual abundances of the population of transcripts are randomly sampled), transcript-level information still remains an important challenge. As a consequence, the role of alternative
356:
platform. As a result of short read length, information regarding the longer patterns of methylation are lost. Third generation sequencing technologies offer the capability for single molecule real-time sequencing of longer reads, and detection of DNA modification without the aforementioned assay.
307:
Long read lengths offered by third generation sequencing may alleviate many of the challenges currently faced by de novo genome assemblies. For example, if an entire repetitive region can be sequenced unambiguously in a single read, no computation inference would be required. Computational methods
297:
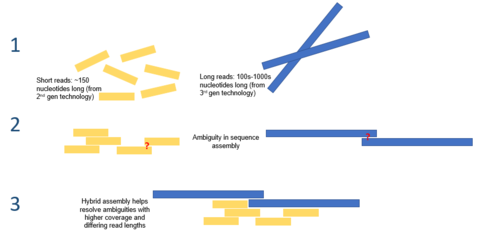
Hybrid assembly – the use of reads from 3rd gen sequencing platforms with shorts reads from 2nd gen platforms – may be used to resolve ambiguities that exist in genomes previously assembled using second generation sequencing. Short second generation reads have also been used to correct errors that
229:
Third generation sequencing, as of 2008, faced important challenges mainly surrounding accurate identification of nucleotide bases; error rates were still much higher compared to second generation sequencing. This is generally due to instability of the molecular machinery involved. For example, in
419:
sequencing has also been used to detect DNA methylation. In this platform, the pulse width – the width of a fluorescent light pulse – corresponds to a specific base. In 2010 it was shown that the interpulse distance in control and methylated samples are different, and there is a "signature" pulse
389:
DNA and the resulting signals measured by the nanopore technology. Then the trained model was used to detect 5mC in MinION genomic reads from a human cell line which already had a reference methylome. The classifier has 82% accuracy in randomly sampled singleton sites, which increases to 95% when
447:
Processing of the raw data – such as normalization to the median signal – was needed on MinION raw data, reducing real-time capability of the technology. Consistency of the electrical signals is still an issue, making it difficult to accurately call a nucleotide. MinION has low throughput; since
257:
When a reference genome is available, as one is in the case of human, newly sequenced reads could simply be aligned to the reference genome in order to characterize its properties. Such reference based assembly is quick and easy but has the disadvantage of “hiding" novel sequences and large copy
127:
are stable and potentially heritable modifications to the DNA molecule that are not in its sequence. An example is DNA methylation at CpG sites, which has been found to influence gene expression. Histone modifications are another example. The current generation of sequencing technologies rely on
110:
It is well known that eukaryotic genomes including primates and humans are complex and have large numbers of long repeated regions. Short reads from second generation sequencing must resort to approximative strategies in order to infer sequences over long ranges for assembly and genetic variant
132:
for the detection of epigenetic markers. These techniques involve tagging the DNA strand, breaking and filtering fragments that contain markers, followed by sequencing. Third generation sequencing may enable direct detection of these markers due to their distinctive signal from the other four
393:
Other methods address different types of DNA modifications using the MinION platform. Stoiber et al. examined 4-methylcytosine (4mC) and 6-methyladenine (6mA), along with 5mC, and also created software to directly visualize the raw MinION data in a human-friendly way. Here they found that in
335:
machinery. DNA modifications and resulting gene expression can vary across cell types, temporal development, with genetic ancestry, can change due to environmental stimuli and are heritable. After the discovery of DNAm, researchers have also found its correlation to diseases like cancer and
106:
In comparison to the current generation of sequencing technologies, third generation sequencing has the obvious advantage of producing much longer reads. It is expected that these longer read lengths will alleviate numerous computational challenges surrounding genome assembly, transcript
272:
assembly is the alternative genome assembly approach to reference alignment. It refers to the reconstruction of whole genome sequences entirely from raw sequence reads. This method would be chosen when there is no reference genome, when the species of the given organism is unknown as in
115:
have been leveraged by second generation sequencing to combat these limitations. However, exact fragment lengths of pair ends are often unknown and must also be approximated as well. By making long reads lengths possible, third generation sequencing technologies have clear advantages.
471:, genetic information flows from double stranded DNA molecules to single stranded mRNA molecules where they can be readily translated into functional protein molecules. By studying the transcriptome, one can gain valuable insight into the regulation of gene expression.
77:
involves passing a DNA molecule through a nanoscale pore structure and then measuring changes in electrical field surrounding the pore; while
Quantapore has a different proprietary nanopore approach. Stratos Genomics spaces out the DNA bases with polymeric inserts,
578:
pathogens were not identified. Ease of carryover contamination when re-using the same flow cell (standard wash protocols don’t work) is also a concern. Unique barcodes may allow for more multiplexing. Furthermore, performing accurate species identification for
1182:
Stoiber, Marcus H.; Quick, Joshua; Egan, Rob; Lee, Ji Eun; Celniker, Susan E.; Neely, Robert; Loman, Nicholas; Pennacchio, Len; Brown, James B. (2016-12-15). "De novo
Identification of DNA Modifications Enabled by Genome-Guided Nanopore Signal Processing".
312:
generation sequencing. On the other hand, short second generation reads have been used to correct errors in that exist in the long third generation reads. In general, this hybrid approach has been shown to improve de novo genome assemblies significantly.
175:
Parts of this article (those related to long-read sequencing technologies producing low-accuracy reads. While true 5 years ago, circular consensus reads with the PacBio Sequel II long-read sequencer can easily achieve an even higher read accuracy than
957:
Chin, Chen-Shan; Alexander, David H.; Marks, Patrick; Klammer, Aaron A.; Drake, James; Heiner, Cheryl; Clum, Alicia; Copeland, Alex; Huddleston, John (2013-06-01). "Nonhybrid, finished microbial genome assemblies from long-read SMRT sequencing data".
285:
and overlap layout consensus graphs, have been leveraged to solve this problem. Nonetheless, due to the highly repetitive nature of eukaryotic genomes, accurate and complete reconstruction of genome sequences in de novo assembly remains challenging.
308:
have been proposed to alleviate the issue of high error rates. For example, in one study, it was demonstrated that de novo assembly of a microbial genome using PacBio sequencing alone performed superior to that of second generation sequencing.
594:
The per base sequencing cost is still significantly more than that of MiSeq. However, the prospect of supplementing reference databases with full-length sequences from organisms below the limit of detection from the
86:
475:
splicing in molecular biology remains largely elusive. Third generation sequencing technologies hold promising prospects in resolving this issue by enabling sequencing of mRNA molecules at their full lengths.
829:
Simpson, Jared T.; Workman, Rachael; Zuzarte, Philip C.; David, Matei; Dursi, Lewis
Jonathan; Timp, Winston (2016-04-04). "Detecting DNA Methylation using the Oxford Nanopore Technologies MinION sequencer".
1479:
Pan, Qun; Shai, Ofer; Lee, Leo J.; Frey, Brendan J.; Blencowe, Benjamin J. (2008-12-01). "Deep surveying of alternative splicing complexity in the human transcriptome by high-throughput sequencing".
375:
has been used to detect DNAm. As each DNA strand passes through a pore, it produces electrical signals which have been found to be sensitive to epigenetic changes in the nucleotides, and a
439:
Other forms of DNA modifications – from heavy metals, oxidation, or UV damage – are also possible avenues of research using Oxford
Nanopore and PacBio third generation sequencing.
1384:
Steijger, Tamara; Abril, Josep F.; Engström, Pär G.; Kokocinski, Felix; The RGASP Consortium; Hubbard, Tim J.; Guigó, Roderic; Harrow, Jennifer; Bertone, Paul (2013-12-01).
1594:
Trapnell, Cole; Williams, Brian A.; Pertea, Geo; Mortazavi, Ali; Kwan, Gordon; van Baren, Marijke J.; Salzberg, Steven L.; Wold, Barbara J.; Pachter, Lior (2010-05-01).
558:
gene. Both MinION and PacBio's SMRT platform have been used to sequence this gene. In this context the PacBio error rate was comparable to that of shorter reads from
1128:
Flusberg, Benjamin A.; Webster, Dale R.; Lee, Jessica H.; Travers, Kevin J.; Olivares, Eric C.; Clark, Tyson A.; Korlach, Jonas; Turner, Stephen W. (2010-06-01).
1261:
Greer, Eric
Lieberman; Blanco, Mario Andres; Gu, Lei; Sendinc, Erdem; Liu, Jianzhao; Aristizábal-Corrales, David; Hsu, Chih-Hung; Aravind, L.; He, Chuan (2015).
1654:
Abdel-Ghany, Salah E.; Hamilton, Michael; Jacobi, Jennifer L.; Ngam, Peter; Devitt, Nicholas; Schilkey, Faye; Ben-Hur, Asa; Reddy, Anireddy S. N. (2016-06-24).
1310:
Wu, Tao P.; Wang, Tao; Seetin, Matthew G.; Lai, Yongquan; Zhu, Shijia; Lin, Kaixuan; Liu, Yifei; Byrum, Stephanie D.; Mackintosh, Samuel G. (2016-04-21).
536:
is their speed of sequencing in comparison to second generation techniques. Speed of sequencing is important for example in the clinical setting (i.e.
543:
Oxford
Nanopore's MinION was used in 2015 for real-time metagenomic detection of pathogens in complex, high-background clinical samples. The first
71:. Signals are in the form of fluorescent light emission from each nucleotide incorporated by a DNA polymerase bound to the bottom of the zL well.
1204:
Clark, T. A.; Murray, I. A.; Morgan, R. D.; Kislyuk, A. O.; Spittle, K. E.; Boitano, M.; Fomenkov, A.; Roberts, R. J.; Korlach, J. (2012-02-01).
1794:; Naccache, Samia N.; Federman, Scot; Yu, Guixia; Mbala, Placide; Bres, Vanessa; Stryke, Doug; Bouquet, Jerome; Somasekar, Sneha (2015-01-01).
60:, Quantapore (CA-USA), and Stratos (WA-USA). These companies are taking fundamentally different approaches to sequencing single DNA molecules.
637:
Bleidorn, Christoph (2016-01-02). "Third generation sequencing: technology and its potential impact on evolutionary biodiversity research".
49:
Sequencing technologies with a different approach than second-generation platforms were first described as "third-generation" in 2008–2009.
402:, event windows of 5 base pairs long can be used to divide and statistically analyze the raw MinION electrical signals. A straightforward
900:
Li, Ruiqiang; Zhu, Hongmei; Ruan, Jue; Qian, Wubin; Fang, Xiaodong; Shi, Zhongbin; Li, Yingrui; Li, Shengting; Shan, Gao (2010-02-01).
1596:"Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation"
1009:
Goodwin, Sara; Gurtowski, James; Ethe-Sayers, Scott; Deshpande, Panchajanya; Schatz, Michael C.; McCombie, W. Richard (2015-11-01).
1537:
Pertea, Mihaela; Pertea, Geo M.; Antonescu, Corina M.; Chang, Tsung-Cheng; Mendell, Joshua T.; Salzberg, Steven L. (2015-03-01).
2026:
467:, usually by characterizing the relative abundances of messenger RNA molecules in the tissue under study. According to the
249:
is the reconstruction of whole genome DNA sequences. This is generally done with two fundamentally different approaches.
1856:
Schloss, Patrick D.; Jenior, Matthew L.; Koumpouras, Charles C.; Westcott, Sarah L.; Highlander, Sarah K. (2016-01-01).
468:
1796:"Rapid metagenomic identification of viral pathogens in clinical samples by real-time nanopore sequencing analysis"
413:
It seems likely that in the future, MinION raw data will be used to detect many different epigenetic marks in DNA.
290:
have been posed as a possible solution, though exact fragment lengths are often unknown and must be approximated.
52:
There are several companies currently at the heart of third generation sequencing technology development, namely,
2036:
730:"NanoVar: accurate characterization of patients' genomic structural variants using low-depth nanopore sequencing"
1909:"Species-level resolution of 16S rRNA gene amplicons sequenced through the MinION™ portable nanopore sequencer"
693:
Gupta, Pushpendra K. (2008-11-01). "Single-molecule DNA sequencing technologies for future genomics research".
613:
368:
277:, or when there exist genetic variants of interest that may not be detected by reference genome alignment.
181:
57:
1206:"Characterization of DNA methyltransferase specificities using single-molecule, real-time DNA sequencing"
608:
38:
141:
37:
Third generation sequencing technologies have the capability to produce substantially longer reads than
2021:
571:
1444:
Graveley, Brenton R. (2001). "Alternative splicing: increasing diversity in the proteomic world".
1011:"Oxford Nanopore sequencing, hybrid error correction, and de novo assembly of a eukaryotic genome"
591:
is very difficult, as they share a larger portion of the genome, and some only differ by <5%.
403:
2031:
1791:
425:
177:
599:
approach; this could possibly greatly help the identification of organisms in metagenomics.
107:
reconstruction, and metagenomics among other important areas of modern biology and medicine.
420:
width for each methylation type. In 2012 using the PacBio platform the binding sites of DNA
1732:
1667:
1323:
1184:
831:
646:
483:
349:
8:
433:
416:
376:
238:
of characterizing individual differences that exist between members of the same species.
231:
90:
74:
68:
53:
1736:
1671:
1327:
650:
212:
Please help update this article to reflect recent events or newly available information.
1943:
1908:
1884:
1857:
1830:
1795:
1763:
1720:
1696:
1655:
1628:
1595:
1571:
1538:
1512:
1418:
1385:
1352:
1311:
1287:
1262:
1238:
1205:
1154:
1129:
1102:
1067:
1043:
1010:
991:
934:
901:
756:
729:
670:
364:
PacBio SMRT technology and Oxford
Nanopore can use unaltered DNA to detect methylation.
353:
1457:
2000:
1992:
1948:
1930:
1889:
1835:
1817:
1768:
1750:
1701:
1683:
1633:
1615:
1576:
1558:
1504:
1496:
1461:
1423:
1405:
1357:
1339:
1292:
1243:
1225:
1159:
1107:
1089:
1066:
Fraser, Hunter B.; Lam, Lucia L.; Neumann, Sarah M.; Kobor, Michael S. (2012-02-09).
1048:
1030:
995:
983:
975:
939:
921:
882:
874:
802:
761:
710:
662:
596:
559:
421:
258:
number variants. In addition, reference genomes do not yet exist for most organisms.
246:
183:
674:
340:. In this disease etiology context DNAm is an important avenue of further research.
89:'s single molecule fluorescence approach, but the company entered bankruptcy in the
1982:
1938:
1920:
1879:
1869:
1825:
1807:
1758:
1740:
1691:
1675:
1623:
1607:
1566:
1550:
1516:
1488:
1453:
1413:
1397:
1347:
1331:
1282:
1274:
1233:
1217:
1149:
1141:
1097:
1079:
1038:
1022:
967:
929:
913:
864:
792:
751:
741:
702:
654:
574:
markers, for which single nucleotide resolution is necessary. For the same reason,
524:
is the analysis of genetic material recovered directly from environmental samples.
410:
sequence, as well as further split the modifications into 4mC, 6mA or 5mC regions.
384:
380:
1130:"Direct detection of DNA methylation during single-molecule, real-time sequencing"
706:
658:
1745:
1539:"StringTie enables improved reconstruction of a transcriptome from RNA-seq reads"
902:"De novo assembly of human genomes with massively parallel short read sequencing"
496:
460:
320:
282:
129:
64:
1858:"Sequencing 16S rRNA gene fragments using the PacBio SMRT DNA sequencing system"
1987:
1970:
1719:
Au, Kin Fai; Underwood, Jason G.; Lee, Lawrence; Wong, Wing Hung (2012-10-04).
1278:
746:
540:
identification), to allow for efficient diagnosis and timely clinical actions.
287:
112:
31:
27:
1925:
1812:
2015:
1996:
1934:
1821:
1754:
1687:
1619:
1562:
1500:
1409:
1343:
1229:
1093:
1034:
979:
925:
878:
666:
555:
551:
464:
383:(5mC) DNA modification. The model was trained using synthetically methylated
1084:
2004:
1952:
1893:
1839:
1772:
1705:
1637:
1580:
1508:
1465:
1427:
1361:
1296:
1247:
1163:
1111:
1052:
987:
943:
886:
806:
765:
714:
533:
521:
328:
274:
203:
199:
195:
191:
187:
151:
1221:
1026:
917:
82:", to circumvent the signal to noise challenge of nanopore ssDNA reading.
869:
852:
797:
780:
544:
332:
124:
1679:
1656:"A survey of the sorghum transcriptome using single-molecule long reads"
1335:
1874:
1401:
1145:
971:
588:
352:
that fragments DNA before standard second generation sequencing on the
234:, for example, have been leveraged for this purpose with some success.
1907:
Benítez-Páez, Alfonso; Portune, Kevin J.; Sanz, Yolanda (2016-01-01).
360:
1611:
1554:
575:
399:
1008:
1492:
1189:
836:
580:
537:
532:
The main advantage for third-generation sequencing technologies in
1312:"DNA methylation on N6-adenine in mammalian embryonic stem cells"
293:
348:
The current most common methods for examining methylation state
1855:
1593:
1383:
584:
570:
MinION's high error rate (~10-40%) prevented identification of
372:
337:
1721:"Improving PacBio Long Read Accuracy by Short Read Alignment"
1653:
1386:"Assessment of transcript reconstruction methods for RNA-seq"
554:
marker for microbial community diversity studies is the 16S
145:
MinION Portable Gene
Sequencer, Oxford Nanopore Technologies
1790:
1536:
828:
1127:
956:
324:
1203:
1906:
851:
Schadt, E. E.; Turner, S.; Kasarskis, A. (2010-10-15).
424:
were characterized. The detection of N6-methylation in
1065:
850:
1260:
1181:
1123:
1121:
504:to detect exons across the 25 protocols is 80% for
1718:
1068:"Population-specificity of human DNA methylation"
2013:
1118:
379:(HMM) was used to analyze MinION data to detect
1309:
1478:
1263:"DNA Methylation on N6-Adenine in C. elegans"
899:
778:
489:
432:-adenine using the PacBio platform in mouse
63:PacBio developed the sequencing platform of
853:"A window into third-generation sequencing"
562:and Illumina's MiSeq sequencing platforms.
65:single molecule real time sequencing (SMRT)
1971:"Method of the year: long-read sequencing"
1986:
1942:
1924:
1883:
1873:
1829:
1811:
1762:
1744:
1695:
1627:
1570:
1417:
1351:
1286:
1237:
1188:
1153:
1101:
1083:
1042:
933:
868:
835:
796:
781:"Genome sequencing: the third generation"
755:
745:
298:exist in the long third generation reads.
180:with a combination of other sequencers.
1443:
636:
359:
292:
140:
136:
478:
390:more stringent thresholds are applied.
44:
34:, under active development since 2008.
2014:
1649:
1647:
428:was shown in 2015. DNA methylation on
331:– is the best understood component of
323:(DNAm) – the covalent modification of
252:
30:methods which produce longer sequence
1851:
1849:
1786:
1784:
1782:
1532:
1530:
1528:
1526:
1439:
1437:
1379:
1377:
1375:
1373:
1371:
1177:
1175:
1173:
692:
315:
1968:
824:
822:
820:
818:
816:
727:
688:
686:
684:
632:
630:
628:
452:contigs still need to be assembled.
406:can detect modified portions of the
161:
1644:
327:at CpG sites resulting in attached
261:
13:
1846:
1779:
1523:
1434:
1368:
1170:
779:Check Hayden, Erika (2009-02-06).
469:central dogma of molecular biology
455:
302:
241:
14:
2048:
1962:
813:
681:
625:
1969:Marx, Vivien (12 January 2023).
166:
1900:
1712:
1587:
1472:
1303:
1254:
1197:
1059:
728:Tham, Cheng Yong (2020-03-03).
527:
516:
343:
101:
1002:
950:
893:
844:
772:
721:
565:
128:laboratory techniques such as
119:
1:
1458:10.1016/s0168-9525(00)02176-4
707:10.1016/j.tibtech.2008.07.003
659:10.1080/14772000.2015.1099575
619:
157:
96:
67:, based on the properties of
2027:Molecular biology techniques
1746:10.1371/journal.pone.0046679
639:Systematics and Biodiversity
614:Second-generation sequencing
509:of all existing transcript.
442:
369:Oxford Nanopore Technologies
75:Oxford Nanopore’s technology
39:second generation sequencing
7:
609:First-generation sequencing
602:
20:Third-generation sequencing
10:
2053:
1988:10.1038/s41592-022-01730-w
1279:10.1016/j.cell.2015.04.005
747:10.1186/s13059-020-01968-7
740:(Article number: 56): 56.
58:Oxford Nanopore Technology
1926:10.1186/s13742-016-0111-z
1813:10.1186/s13073-015-0220-9
490:Transcript reconstruction
857:Human Molecular Genetics
572:antimicrobial resistance
1792:Greninger, Alexander L.
1085:10.1186/gb-2012-13-2-r8
695:Trends in Biotechnology
2037:DNA sequencing methods
1210:Nucleic Acids Research
506:Caenorhabditis elegans
365:
299:
178:hybrid genome assembly
146:
1660:Nature Communications
1027:10.1101/gr.191395.115
918:10.1101/gr.097261.109
363:
296:
144:
137:Portability and speed
1600:Nature Biotechnology
1543:Nature Biotechnology
798:10.1038/news.2009.86
484:Alternative splicing
479:Alternative splicing
463:is the study of the
434:embryonic stem cells
398:, which has a known
232:Hidden Markov Models
69:zero-mode waveguides
45:Current technologies
24:long-read sequencing
1737:2012PLoSO...746679A
1680:10.1038/ncomms11706
1672:2016NatCo...711706A
1336:10.1038/nature17640
1328:2016Natur.532..329W
1222:10.1093/nar/gkr1146
651:2016SyBio..14....1B
436:was shown in 2016.
404:Mann-Whitney U test
377:hidden Markov model
253:Reference alignment
54:Pacific Biosciences
1875:10.7717/peerj.1869
1446:Trends in Genetics
1402:10.1038/nmeth.2714
1146:10.1038/nmeth.1459
972:10.1038/nmeth.2474
870:10.1093/hmg/ddq416
422:methyltransferases
366:
316:Epigenetic markers
300:
147:
133:nucleotide bases.
125:Epigenetic markers
2022:Molecular biology
1487:(12): 1413–1415.
1396:(12): 1177–1184.
1322:(7599): 329–333.
1021:(11): 1750–1756.
863:(R2): R227–R240.
791:(7231): 768–769.
227:
226:
2044:
2008:
1990:
1957:
1956:
1946:
1928:
1904:
1898:
1897:
1887:
1877:
1853:
1844:
1843:
1833:
1815:
1788:
1777:
1776:
1766:
1748:
1716:
1710:
1709:
1699:
1651:
1642:
1641:
1631:
1612:10.1038/nbt.1621
1591:
1585:
1584:
1574:
1555:10.1038/nbt.3122
1534:
1521:
1520:
1476:
1470:
1469:
1441:
1432:
1431:
1421:
1381:
1366:
1365:
1355:
1307:
1301:
1300:
1290:
1258:
1252:
1251:
1241:
1201:
1195:
1194:
1192:
1179:
1168:
1167:
1157:
1125:
1116:
1115:
1105:
1087:
1063:
1057:
1056:
1046:
1006:
1000:
999:
954:
948:
947:
937:
897:
891:
890:
872:
848:
842:
841:
839:
826:
811:
810:
800:
776:
770:
769:
759:
749:
725:
719:
718:
690:
679:
678:
634:
381:5-methylcytosine
350:require an assay
283:de bruijn graphs
222:
219:
213:
170:
169:
162:
152:MinION sequencer
85:Also notable is
26:) is a class of
2052:
2051:
2047:
2046:
2045:
2043:
2042:
2041:
2012:
2011:
1965:
1960:
1905:
1901:
1854:
1847:
1800:Genome Medicine
1789:
1780:
1717:
1713:
1652:
1645:
1592:
1588:
1535:
1524:
1481:Nature Genetics
1477:
1473:
1442:
1435:
1382:
1369:
1308:
1304:
1259:
1255:
1202:
1198:
1180:
1171:
1126:
1119:
1064:
1060:
1015:Genome Research
1007:
1003:
955:
951:
906:Genome Research
898:
894:
849:
845:
827:
814:
777:
773:
726:
722:
701:(11): 602–611.
691:
682:
635:
626:
622:
605:
568:
530:
519:
492:
481:
461:Transcriptomics
458:
456:Transcriptomics
445:
346:
321:DNA methylation
318:
305:
303:Hybrid assembly
267:
255:
247:Genome assembly
244:
242:Genome assembly
223:
217:
214:
211:
171:
167:
160:
139:
130:ChIP-sequencing
122:
104:
99:
47:
22:(also known as
17:
12:
11:
5:
2050:
2040:
2039:
2034:
2029:
2024:
2010:
2009:
1975:Nature Methods
1964:
1963:External links
1961:
1959:
1958:
1899:
1845:
1778:
1731:(10): e46679.
1711:
1643:
1606:(5): 511–515.
1586:
1549:(3): 290–295.
1522:
1493:10.1038/ng.259
1471:
1452:(2): 100–107.
1433:
1390:Nature Methods
1367:
1302:
1273:(4): 868–878.
1253:
1196:
1190:10.1101/094672
1169:
1140:(6): 461–465.
1134:Nature Methods
1117:
1072:Genome Biology
1058:
1001:
966:(6): 563–569.
960:Nature Methods
949:
912:(2): 265–272.
892:
843:
837:10.1101/047142
812:
771:
734:Genome Biology
720:
680:
623:
621:
618:
617:
616:
611:
604:
601:
567:
564:
529:
526:
518:
515:
491:
488:
480:
477:
457:
454:
444:
441:
345:
342:
317:
314:
304:
301:
288:Pair end reads
266:
260:
254:
251:
243:
240:
225:
224:
174:
172:
165:
159:
156:
138:
135:
121:
118:
113:Pair end reads
103:
100:
98:
95:
46:
43:
28:DNA sequencing
16:DNA sequencing
15:
9:
6:
4:
3:
2:
2049:
2038:
2035:
2033:
2032:Biotechnology
2030:
2028:
2025:
2023:
2020:
2019:
2017:
2006:
2002:
1998:
1994:
1989:
1984:
1980:
1976:
1972:
1967:
1966:
1954:
1950:
1945:
1940:
1936:
1932:
1927:
1922:
1918:
1914:
1910:
1903:
1895:
1891:
1886:
1881:
1876:
1871:
1867:
1863:
1859:
1852:
1850:
1841:
1837:
1832:
1827:
1823:
1819:
1814:
1809:
1805:
1801:
1797:
1793:
1787:
1785:
1783:
1774:
1770:
1765:
1760:
1756:
1752:
1747:
1742:
1738:
1734:
1730:
1726:
1722:
1715:
1707:
1703:
1698:
1693:
1689:
1685:
1681:
1677:
1673:
1669:
1665:
1661:
1657:
1650:
1648:
1639:
1635:
1630:
1625:
1621:
1617:
1613:
1609:
1605:
1601:
1597:
1590:
1582:
1578:
1573:
1568:
1564:
1560:
1556:
1552:
1548:
1544:
1540:
1533:
1531:
1529:
1527:
1518:
1514:
1510:
1506:
1502:
1498:
1494:
1490:
1486:
1482:
1475:
1467:
1463:
1459:
1455:
1451:
1447:
1440:
1438:
1429:
1425:
1420:
1415:
1411:
1407:
1403:
1399:
1395:
1391:
1387:
1380:
1378:
1376:
1374:
1372:
1363:
1359:
1354:
1349:
1345:
1341:
1337:
1333:
1329:
1325:
1321:
1317:
1313:
1306:
1298:
1294:
1289:
1284:
1280:
1276:
1272:
1268:
1264:
1257:
1249:
1245:
1240:
1235:
1231:
1227:
1223:
1219:
1215:
1211:
1207:
1200:
1191:
1186:
1178:
1176:
1174:
1165:
1161:
1156:
1151:
1147:
1143:
1139:
1135:
1131:
1124:
1122:
1113:
1109:
1104:
1099:
1095:
1091:
1086:
1081:
1077:
1073:
1069:
1062:
1054:
1050:
1045:
1040:
1036:
1032:
1028:
1024:
1020:
1016:
1012:
1005:
997:
993:
989:
985:
981:
977:
973:
969:
965:
961:
953:
945:
941:
936:
931:
927:
923:
919:
915:
911:
907:
903:
896:
888:
884:
880:
876:
871:
866:
862:
858:
854:
847:
838:
833:
825:
823:
821:
819:
817:
808:
804:
799:
794:
790:
786:
782:
775:
767:
763:
758:
753:
748:
743:
739:
735:
731:
724:
716:
712:
708:
704:
700:
696:
689:
687:
685:
676:
672:
668:
664:
660:
656:
652:
648:
644:
640:
633:
631:
629:
624:
615:
612:
610:
607:
606:
600:
598:
592:
590:
586:
582:
577:
573:
563:
561:
557:
556:ribosomal RNA
553:
548:
546:
541:
539:
535:
525:
523:
514:
510:
507:
501:
498:
487:
485:
476:
472:
470:
466:
465:transcriptome
462:
453:
449:
440:
437:
435:
431:
427:
423:
418:
414:
411:
409:
405:
401:
397:
391:
388:
387:
382:
378:
374:
370:
362:
358:
355:
351:
341:
339:
334:
330:
329:methyl groups
326:
322:
313:
309:
295:
291:
289:
284:
278:
276:
271:
264:
259:
250:
248:
239:
235:
233:
221:
209:
206:) need to be
205:
201:
197:
193:
189:
185:
182:
179:
173:
164:
163:
155:
153:
143:
134:
131:
126:
117:
114:
108:
94:
92:
88:
83:
81:
76:
72:
70:
66:
61:
59:
55:
50:
42:
40:
35:
33:
29:
25:
21:
1978:
1974:
1916:
1912:
1902:
1865:
1861:
1803:
1799:
1728:
1724:
1714:
1663:
1659:
1603:
1599:
1589:
1546:
1542:
1484:
1480:
1474:
1449:
1445:
1393:
1389:
1319:
1315:
1305:
1270:
1266:
1256:
1213:
1209:
1199:
1137:
1133:
1075:
1071:
1061:
1018:
1014:
1004:
963:
959:
952:
909:
905:
895:
860:
856:
846:
788:
784:
774:
737:
733:
723:
698:
694:
642:
638:
593:
569:
552:phylogenetic
549:
542:
534:metagenomics
531:
522:Metagenomics
520:
517:Metagenomics
511:
505:
502:
493:
482:
473:
459:
450:
446:
438:
429:
415:
412:
407:
395:
392:
385:
367:
347:
319:
310:
306:
279:
275:metagenomics
269:
268:
262:
256:
245:
236:
228:
218:January 2020
215:
207:
148:
123:
109:
105:
102:Longer reads
91:fall of 2015
84:
79:
73:
62:
51:
48:
36:
23:
19:
18:
1981:(1): 6–11.
1913:GigaScience
785:Nature News
545:Ebola virus
120:Epigenetics
2016:Categories
1216:(4): e29.
645:(1): 1–8.
620:References
576:eukaryotic
528:Advantages
344:Advantages
333:epigenetic
158:Challenges
97:Advantages
80:Xpandomers
1997:1548-7105
1935:2047-217X
1868:: e1869.
1822:1756-994X
1755:1932-6203
1688:2041-1723
1666:: 11706.
1620:1087-0156
1563:1087-0156
1501:1061-4036
1410:1548-7091
1344:0028-0836
1230:0305-1048
1094:1474-760X
1078:(2): R8.
1035:1088-9051
996:205421576
980:1548-7091
926:1088-9051
879:0964-6906
667:1477-2000
589:parasites
566:Drawbacks
550:A common
497:Cufflinks
443:Drawbacks
426:C Elegans
400:methylome
111:calling.
2005:36635542
1953:26823973
1894:27069806
1840:26416663
1773:23056399
1725:PLOS ONE
1706:27339290
1638:20436464
1581:25690850
1509:18978789
1466:11173120
1428:24185837
1362:27027282
1297:25936839
1248:22156058
1164:20453866
1112:22322129
1053:26447147
988:23644548
944:20019144
887:20858600
807:19212365
766:32127024
715:18722683
675:85991118
603:See also
581:bacteria
538:pathogen
354:Illumina
281:such as
265:assembly
204:31483244
200:31897449
196:31406327
192:28364362
188:31885515
1944:4730766
1885:4824876
1831:4587849
1764:3464235
1733:Bibcode
1697:4931028
1668:Bibcode
1629:3146043
1572:4643835
1517:9228930
1419:3851240
1353:4977844
1324:Bibcode
1288:4427530
1239:3287169
1185:bioRxiv
1155:2879396
1103:3334571
1044:4617970
935:2813482
832:bioRxiv
757:7055087
647:Bibcode
408:E. coli
396:E. coli
386:E. coli
270:De novo
263:De novo
208:updated
87:Helicos
2003:
1995:
1951:
1941:
1933:
1892:
1882:
1838:
1828:
1820:
1806:: 99.
1771:
1761:
1753:
1704:
1694:
1686:
1636:
1626:
1618:
1579:
1569:
1561:
1515:
1507:
1499:
1464:
1426:
1416:
1408:
1360:
1350:
1342:
1316:Nature
1295:
1285:
1246:
1236:
1228:
1187:
1162:
1152:
1110:
1100:
1092:
1051:
1041:
1033:
994:
986:
978:
942:
932:
924:
885:
877:
834:
805:
764:
754:
713:
673:
665:
597:Sanger
417:PacBio
373:MinION
338:autism
186:
1919:: 4.
1862:PeerJ
1513:S2CID
992:S2CID
671:S2CID
585:fungi
32:reads
2001:PMID
1993:ISSN
1949:PMID
1931:ISSN
1890:PMID
1836:PMID
1818:ISSN
1769:PMID
1751:ISSN
1702:PMID
1684:ISSN
1634:PMID
1616:ISSN
1577:PMID
1559:ISSN
1505:PMID
1497:ISSN
1462:PMID
1424:PMID
1406:ISSN
1358:PMID
1340:ISSN
1293:PMID
1267:Cell
1244:PMID
1226:ISSN
1160:PMID
1108:PMID
1090:ISSN
1049:PMID
1031:ISSN
984:PMID
976:ISSN
940:PMID
922:ISSN
883:PMID
875:ISSN
803:PMID
762:PMID
711:PMID
663:ISSN
587:and
184:PMID
150:the
1983:doi
1939:PMC
1921:doi
1880:PMC
1870:doi
1826:PMC
1808:doi
1759:PMC
1741:doi
1692:PMC
1676:doi
1624:PMC
1608:doi
1567:PMC
1551:doi
1489:doi
1454:doi
1414:PMC
1398:doi
1348:PMC
1332:doi
1320:532
1283:PMC
1275:doi
1271:161
1234:PMC
1218:doi
1150:PMC
1142:doi
1098:PMC
1080:doi
1039:PMC
1023:doi
968:doi
930:PMC
914:doi
865:doi
793:doi
789:457
752:PMC
742:doi
703:doi
655:doi
560:454
325:DNA
2018::
1999:.
1991:.
1979:20
1977:.
1973:.
1947:.
1937:.
1929:.
1915:.
1911:.
1888:.
1878:.
1864:.
1860:.
1848:^
1834:.
1824:.
1816:.
1802:.
1798:.
1781:^
1767:.
1757:.
1749:.
1739:.
1727:.
1723:.
1700:.
1690:.
1682:.
1674:.
1662:.
1658:.
1646:^
1632:.
1622:.
1614:.
1604:28
1602:.
1598:.
1575:.
1565:.
1557:.
1547:33
1545:.
1541:.
1525:^
1511:.
1503:.
1495:.
1485:40
1483:.
1460:.
1450:17
1448:.
1436:^
1422:.
1412:.
1404:.
1394:10
1392:.
1388:.
1370:^
1356:.
1346:.
1338:.
1330:.
1318:.
1314:.
1291:.
1281:.
1269:.
1265:.
1242:.
1232:.
1224:.
1214:40
1212:.
1208:.
1172:^
1158:.
1148:.
1136:.
1132:.
1120:^
1106:.
1096:.
1088:.
1076:13
1074:.
1070:.
1047:.
1037:.
1029:.
1019:25
1017:.
1013:.
990:.
982:.
974:.
964:10
962:.
938:.
928:.
920:.
910:20
908:.
904:.
881:.
873:.
861:19
859:.
855:.
815:^
801:.
787:.
783:.
760:.
750:.
738:21
736:.
732:.
709:.
699:26
697:.
683:^
669:.
661:.
653:.
643:14
641:.
627:^
583:,
371:’
202:,
198:,
194:,
190:,
93:.
56:,
2007:.
1985::
1955:.
1923::
1917:5
1896:.
1872::
1866:4
1842:.
1810::
1804:7
1775:.
1743::
1735::
1729:7
1708:.
1678::
1670::
1664:7
1640:.
1610::
1583:.
1553::
1519:.
1491::
1468:.
1456::
1430:.
1400::
1364:.
1334::
1326::
1299:.
1277::
1250:.
1220::
1193:.
1166:.
1144::
1138:7
1114:.
1082::
1055:.
1025::
998:.
970::
946:.
916::
889:.
867::
840:.
809:.
795::
768:.
744::
717:.
705::
677:.
657::
649::
430:N
220:)
216:(
210:.
78:"
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.