183:
1264:
31:
246:
3828:
199:
238:
1272:
386:, sometimes also NMR-spectroscopy. Structural details, in particular distances between donor and acceptor which are smaller than the sum of the van der Waals radii can be taken as indication of the hydrogen bond strength. One scheme gives the following somewhat arbitrary classification: those that are 15 to 40 kcal/mol, 5 to 15 kcal/mol, and >0 to 5 kcal/mol are considered strong, moderate, and weak, respectively.
3822:
1044:
3834:
744:
736:
267:
that there are many examples of weaker hydrogen bonding involving donor other than N, O, or F and/or acceptor Ac with electronegativity approaching that of hydrogen (rather than being much more electronegative). Although weak (≈1 kcal/mol), "non-traditional" hydrogen bonding interactions are ubiquitous and influence structures of many kinds of materials.
795:. Owing to the difficulty of breaking these bonds, water has a very high boiling point, melting point, and viscosity compared to otherwise similar liquids not conjoined by hydrogen bonds. Water is unique because its oxygen atom has two lone pairs and two hydrogen atoms, meaning that the total number of bonds of a water molecule is up to four.
599:
prior to melting, variable-temperature infrared spectroscopy can reveal the temperature dependence of hydrogen bonds and the dynamics of both the anions and the cations. The sudden weakening of hydrogen bonds during the solid-solid phase transition seems to be coupled with the onset of orientational
225:
In a hydrogen bond, the electronegative atom not covalently attached to the hydrogen is named the proton acceptor, whereas the one covalently bound to the hydrogen is named the proton donor. This nomenclature is recommended by the IUPAC. The hydrogen of the donor is protic and therefore can act as a
772:
and is often used as a model system. When more molecules are present, as is the case with liquid water, more bonds are possible because the oxygen of one water molecule has two lone pairs of electrons, each of which can form a hydrogen bond with a hydrogen on another water molecule. This can repeat
266:
In weaker hydrogen bonds, hydrogen atoms tend to bond to elements such as sulfur (S) or chlorine (Cl); even carbon (C) can serve as a donor, particularly when the carbon or one of its neighbors is electronegative (e.g., in chloroform, aldehydes and terminal acetylenes). Gradually, it was recognized
1213:
Several studies have shown that hydrogen bonds play an important role for the stability between subunits in multimeric proteins. For example, a study of sorbitol dehydrogenase displayed an important hydrogen bonding network which stabilizes the tetrameric quaternary structure within the mammalian
594:
stretching frequency and a decrease in the bond length. H-bonds can also be measured by IR vibrational mode shifts of the acceptor. The amide I mode of backbone carbonyls in α-helices shifts to lower frequencies when they form H-bonds with side-chain hydroxyl groups. The dynamics of hydrogen bond
802:
liquid water simulations at 25 °C, it was estimated that each water molecule participates in an average of 3.59 hydrogen bonds. At 100 °C, this number decreases to 3.24 due to the increased molecular motion and decreased density, while at 0 °C, the average number of hydrogen bonds
1023:
bonds that comprise most polymers, hydrogen bonds are far weaker, perhaps 5%. Thus, hydrogen bonds can be broken by chemical or mechanical means while retaining the basic structure of the polymer backbone. This hierarchy of bond strengths (covalent bonds being stronger than hydrogen-bonds being
815:
A single hydrogen atom can participate in two hydrogen bonds. This type of bonding is called "bifurcated" (split in two or "two-forked"). It can exist, for instance, in complex organic molecules. It has been suggested that a bifurcated hydrogen atom is an essential step in water reorientation.
433:
distance is ≈160 to 200 pm. The typical length of a hydrogen bond in water is 197 pm. The ideal bond angle depends on the nature of the hydrogen bond donor. The following hydrogen bond angles between a hydrofluoric acid donor and various acceptors have been determined experimentally:
381:
The strength of intermolecular hydrogen bonds is most often evaluated by measurements of equilibria between molecules containing donor and/or acceptor units, most often in solution. The strength of intramolecular hydrogen bonds can be studied with equilibria between conformers with and without
806:
Because water may form hydrogen bonds with solute proton donors and acceptors, it may competitively inhibit the formation of solute intermolecular or intramolecular hydrogen bonds. Consequently, hydrogen bonds between or within solute molecules dissolved in water are almost always unfavorable
2529:
Luo, Jiangshui; Jensen, Annemette H.; Brooks, Neil R.; Sniekers, Jeroen; Knipper, Martin; Aili, David; Li, Qingfeng; Vanroy, Bram; Wübbenhorst, Michael; Yan, Feng; Van
Meervelt, Luc; Shao, Zhigang; Fang, Jianhua; Luo, Zheng-Hong; De Vos, Dirk E.; Binnemans, Koen; Fransaer, Jan (2015).
3413:
Ozeryanskii, Valery A.; Pozharskii, Alexander F.; Bieńko, Agnieszka J.; Sawka-Dobrowolska, Wanda; Sobczyk, Lucjan (2005-03-01). "+ Hydrogen
Bonding in Protonated 1,8-Bis(dimethylamino)-2,7-dimethoxynaphthalene. X-ray Diffraction, Infrared, and Theoretical ab Initio and DFT Studies".
42:
2022:
Arunan, Elangannan; Desiraju, Gautam R.; Klein, Roger A.; Sadlej, Joanna; Scheiner, Steve; Alkorta, Ibon; Clary, David C.; Crabtree, Robert H.; Dannenberg, Joseph J.; Hobza, Pavel; Kjaergaard, Henrik G.; Legon, Anthony C.; Mennucci, Benedetta; Nesbitt, David J. (2011).
149:
The hydrogen bond is also responsible for many of the physical and chemical properties of compounds of N, O, and F that seem unusual compared with other similar structures. In particular, intermolecular hydrogen bonding is responsible for the high boiling point of
122:(occurring among parts of the same molecule). The energy of a hydrogen bond depends on the geometry, the environment, and the nature of the specific donor and acceptor atoms and can vary between 1 and 40 kcal/mol. This makes them somewhat stronger than a
1354:. This type of bond is much stronger than a "normal" hydrogen bond. The effective bond order is 0.5, so its strength is comparable to a covalent bond. It is seen in ice at high pressure, and also in the solid phase of many anhydrous acids such as
3399:
Khashayar
Rajabimoghadam Yousef Darwish Umyeena Bashir Dylan Pitman Sidney Eichelberger Maxime A. Siegler Marcel Swart Isaac Garcia-Bosch Aerobic Oxidation of Alcohols by Copper Complexes Bearing Redox-Active Ligands with Tunable H-Bonding
1179:. The energy preference of the bifurcated H-bond hydroxyl or thiol system is -3.4 kcal/mol or -2.6 kcal/mol, respectively. This type of bifurcated H-bond provides an intrahelical H-bonding partner for polar side-chains, such as
1063:
In these macromolecules, bonding between parts of the same macromolecule cause it to fold into a specific shape, which helps determine the molecule's physiological or biochemical role. For example, the double helical structure of
819:
Acceptor-type hydrogen bonds (terminating on an oxygen's lone pairs) are more likely to form bifurcation (it is called overcoordinated oxygen, OCO) than are donor-type hydrogen bonds, beginning on the same oxygen's hydrogens.
1255:, being a protein fibre, is held together by hydrogen bonds, causing wool to recoil when stretched. However, washing at high temperatures can permanently break the hydrogen bonds and a garment may permanently lose its shape.
2907:
1206:(CD) experiments have shown osmolyte to act through an enthalpic effect. The molecular mechanism for their role in protein stabilization is still not well established, though several mechanisms have been proposed. Computer
1202:, shift the protein folding equilibrium toward the folded state, in a concentration dependent manner. While the prevalent explanation for osmolyte action relies on excluded volume effects that are entropic in nature,
595:
structures in water can be probed by this OH stretching vibration. In the hydrogen bonding network in protic organic ionic plastic crystals (POIPCs), which are a type of phase change material exhibiting solid-solid
1318:, where hydrogen bonds stabilize the linear chains laterally. The chain axes are aligned along the fibre axis, making the fibres extremely stiff and strong. Hydrogen-bond networks make both polymers sensitive to
714:
credits T. S. Moore and T. F. Winmill with the first mention of the hydrogen bond, in 1912. Moore and
Winmill used the hydrogen bond to account for the fact that trimethylammonium hydroxide is a weaker base than
773:
such that every water molecule is H-bonded with up to four other molecules, as shown in the figure (two through its two lone pairs, and two through its two hydrogen atoms). Hydrogen bonding strongly affects the
781:, helping to create an open hexagonal lattice. The density of ice is less than the density of water at the same temperature; thus, the solid phase of water floats on the liquid, unlike most other substances.
807:
relative to hydrogen bonds between water and the donors and acceptors for hydrogen bonds on those solutes. Hydrogen bonds between water molecules have an average lifetime of 10 seconds, or 10 picoseconds.
3471:; Siegbahn, Per E. M.; Eisenstein, Odile; Rheingold, Arnold L.; Koetzle, Thomas F. (1996). "A New Intermolecular Interaction: Unconventional Hydrogen Bonds with Element-Hydride Bonds as Proton Acceptor".
1350:
is a special type of hydrogen bond in which the proton is spaced exactly halfway between two identical atoms. The strength of the bond to each of those atoms is equal. It is an example of a
727:, saying, "Mr. Huggins of this laboratory in some work as yet unpublished, has used the idea of a hydrogen kernel held between two atoms as a theory in regard to certain organic compounds."
259:
Hydrogen bonds arise from a combination of electrostatics (multipole-multipole and multipole-induced multipole interactions), covalency (charge transfer by orbital overlap), and dispersion (
1523:
Hapala, Prokop; Kichin, Georgy; Wagner, Christian; Tautz, F. Stefan; Temirov, Ruslan; Jelínek, Pavel (2014-08-19). "Mechanism of high-resolution STM/AFM imaging with functionalized tips".
2331:
Legon, A. C.; Millen, D. J. (1987). "Angular geometries and other properties of hydrogen-bonded dimers: a simple electrostatic interpretation of the success of the electron-pair model".
3072:"Hétérocycles à fonction quinone. V. Réaction anormale de la butanedione avec la diamino-1,2 anthraquinone; structure cristalline de la naphto \2,3-f] quinoxalinedione-7,12 obtenue"
907:
1714:
Arunan, Elangannan; Desiraju, Gautam R.; Klein, Roger A.; Sadlej, Joanna; Scheiner, Steve; Alkorta, Ibon; Clary, David C.; Crabtree, Robert H.; Dannenberg, Joseph J. (2011-07-08).
1033:
52:
molecules on silver-terminated silicon, interacting via hydrogen bonding, taken at 77 K. ("Hydrogen bonds" in the top image are exaggerated by artifacts of the imaging technique.)
2524:
2522:
575:
401:(commonly abbreviated as RAHB) is a strong type of hydrogen bond. It is characterized by the π-delocalization that involves the hydrogen and cannot be properly described by the
2519:
803:
increases to 3.69. Another study found a much smaller number of hydrogen bonds: 2.357 at 25 °C. Defining and counting the hydrogen bonds is not straightforward however.
1578:"Binding and Release between Polymeric Carrier and Protein Drug: pH-Mediated Interplay of Coulomb Forces, Hydrogen Bonding, van der Waals Interactions, and Entropy"
1370:. Due to severe steric constraint, the protonated form of Proton Sponge (1,8-bis(dimethylamino)naphthalene) and its derivatives also have symmetric hydrogen bonds (
1919:
Beijer, Felix H.; Kooijman, Huub; Spek, Anthony L.; Sijbesma, Rint P.; Meijer, E. W. (1998). "Self-Complementarity
Achieved through Quadruple Hydrogen Bonding".
1402:
remains unclear. Generally, the hydrogen bond is characterized by a proton acceptor that is a lone pair of electrons in nonmetallic atoms (most notably in the
1438:
the majority of orally active drugs have no more than five hydrogen bond donors and fewer than ten hydrogen bond acceptors. These interactions exist between
702:
demonstrated information transfer between hydrogen-bonded nuclei, a feat that would only be possible if the hydrogen bond contained some covalent character.
3614:
1322:
levels in the atmosphere because water molecules can diffuse into the surface and disrupt the network. Some polymers are more sensitive than others. Thus
1149:
288:
in which X is more electronegative than H, and an atom or a group of atoms in the same or another molecule, in which there is evidence of bond formation.
2693:
Ghanty, Tapan K.; Staroverov, Viktor N.; Koren, Patrick R.; Davidson, Ernest R. (2000-02-01). "Is the
Hydrogen Bond in Water Dimer and Ice Covalent?".
2067:"Gas-phase bihalide and pseudobihalide ions. An ion cyclotron resonance determination of hydrogen bond energies in XHY- species (X, Y = F, Cl, Br, CN)"
2288:
Lin, Xuhui; Zhang, Huaiyu; Jiang, Xiaoyu; Wu, Wei; Mo, Yirong (2017). "The Origin of the Non-Additivity in
Resonance-Assisted Hydrogen Bond Systems".
1011:
Hydrogen bonding plays an important role in determining the three-dimensional structures and the properties adopted by many proteins. Compared to the
1826:
1466:
Sweetman, A. M.; Jarvis, S. P.; Sang, Hongqian; Lekkas, I.; Rahe, P.; Wang, Yu; Wang, Jianbo; Champness, N.R.; Kantorovich, L.; Moriarty, P. (2014).
3921:
2532:"1,2,4-Triazolium perfluorobutanesulfonate as an archetypal pure protic organic ionic plastic crystal electrolyte for all-solid-state fuel cells"
3249:
Gilman-Politi, Regina; Harries, Daniel (2011). "Unraveling the
Molecular Mechanism of Enthalpy Driven Peptide Folding by Polyol Osmolytes".
1933:
1194:
The role of hydrogen bonds in protein folding has also been linked to osmolyte-induced protein stabilization. Protective osmolytes, such as
2471:
Cowan ML; Bruner BD; Huse N; et al. (2005). "Ultrafast memory loss and energy redistribution in the hydrogen bond network of liquid H
2151:
3286:"A hydrogen-bonding network in mammalian sorbitol dehydrogenase stabilizes the tetrameric state and is essential for the catalytic power"
2623:
Grunenberg, Jörg (2004). "Direct
Assessment of Interresidue Forces in Watson−Crick Base Pairs Using Theoretical Compliance Constants".
1625:
Hämäläinen, Sampsa K.; van der
Heijden, Nadine; van der Lit, Joost; den Hartog, Stephan; Liljeroth, Peter; Swart, Ingmar (2014-10-31).
1426:
of these complexes is similar to hydrogen bonds, in that the bond length is very adaptable to the metal complex/hydrogen donor system.
405:
model alone. This description of the hydrogen bond has been proposed to describe unusually short distances generally observed between
1946:
Desiraju, G. R. and Steiner, T. The Weak Hydrogen Bond: In Structural Chemistry and Biology, International Union of Crystallography;
4039:
3966:
3647:
2728:
Cordier, F; Rogowski, M; Grzesiek, S; Bax, A (1999). "Observation of through-hydrogen-bond (2h)J(HC') in a perdeuterated protein".
2261:
Gu, Yanliang; Kar, Tapas; Scheiner, Steve (1999). "Fundamental Properties of the CH···O Interaction: Is It a True Hydrogen Bond?".
270:
The definition of hydrogen bonding has gradually broadened over time to include these weaker attractive interactions. In 2011, an
1626:
1870:
1418:. In the dihydrogen bond, however, a metal hydride serves as a proton acceptor, thus forming a hydrogen-hydrogen interaction.
3054:
2833:
2607:
2362:
1283:
The properties of many polymers are affected by hydrogen bonds within and/or between the chains. Prominent examples include
2820:
The nature of the chemical bond and the structure of molecules and crystals; an introduction to modern structural chemistry
764:
molecules. In a discrete water molecule, there are two hydrogen atoms and one oxygen atom. The simplest case is a pair of
3617:(Audio slideshow from the National High Magnetic Field Laboratory explaining cohesion, surface tension and hydrogen bonds)
2971:
Zielkiewicz, Jan (2005). "Structural properties of water: Comparison of the SPC, SPCE, TIP4P, and TIP5P models of water".
3961:
45:
1390:
bonding interaction involving hydrogen atoms. These structures have been known for some time, and well characterized by
1134:
is formed. When two strands are joined by hydrogen bonds involving alternating residues on each participating strand, a
274:
Task Group recommended a modern evidence-based definition of hydrogen bonding, which was published in the IUPAC journal
49:
2935:
Jorgensen, W. L.; Madura, J. D. (1985). "Temperature and size dependence for Monte Carlo simulations of TIP4P water".
2571:
Weinhold, Frank; Klein, Roger A. (2014). "What is a hydrogen bond? Resonance covalency in the supramolecular domain".
2148:
as detailed in the reference and should be compared to 7.9 kJ/mol for bulk water, obtained using the same calculation.
1210:
simulations suggest that osmolytes stabilize proteins by modifying the hydrogen bonds in the protein hydration layer.
586:
stretching frequency to lower energy (i.e. the vibration frequency decreases). This shift reflects a weakening of the
4075:
3602:
2536:
230:
system, where the dots represent the hydrogen bond. Liquids that display hydrogen bonding (such as water) are called
3156:
Markovitch, Omer; Agmon, Noam (2008). "The Distribution of Acceptor and Donor Hydrogen-Bonds in Bulk Liquid Water".
382:
hydrogen bonds. The most important method for the identification of hydrogen bonds also in complicated molecules is
3886:
698:
suggested that the hydrogen bonds had a partial covalent nature. This interpretation remained controversial until
3710:
1351:
639:
146:
together, and for causing separate sheets of paper to stick together after becoming wet and subsequently drying.
3901:
3070:
Baron, Michel; Giorgi-Renault, Sylviane; Renault, Jean; Mailliet, Patrick; Carré, Daniel; Etienne, Jean (1984).
627:) revealed large differences between individual H bonds of the same type. For example, the central interresidue
96:, and the dotted or dashed line indicates the hydrogen bond. The most frequent donor and acceptor atoms are the
4080:
3737:
3698:
3688:
1975:
1955:
1815:
1791:
1771:
1311:
effectively link adjacent chains, which gives the material mechanical strength. Hydrogen bonds also affect the
798:
The number of hydrogen bonds formed by a molecule of liquid water fluctuates with time and temperature. From
748:
719:. The description of hydrogen bonding in its better-known setting, water, came some years later, in 1920, from
284:
The hydrogen bond is an attractive interaction between a hydrogen atom from a molecule or a molecular fragment
3214:
Politi, Regina; Harries, Daniel (2010). "Enthalpically driven peptide stabilization by protective osmolytes".
657:
of ordinary ice claim that the hydrogen bond is partly covalent. However, this interpretation was challenged.
3693:
2531:
940:
The fact that ice is less dense than liquid water is due to a crystal structure stabilized by hydrogen bonds.
716:
17:
3620:
842:
723:
and Rodebush. In that paper, Latimer and Rodebush cited the work of a fellow scientist at their laboratory,
3640:
1089:
553:
2658:
Isaacs, E.D.; et al. (1999). "Covalency of the Hydrogen Bond in Ice: A Direct X-Ray Measurement".
1435:
1139:
699:
276:
182:
4070:
4065:
3956:
3946:
3936:
3911:
3881:
643:
3015:
Jencks, William; Jencks, William P. (1986). "Hydrogen Bonding between Solutes in Aqueous Solution".
2226:
Biedermann F, Schneider HJ (May 2016). "Experimental Binding Energies in Supramolecular Complexes".
297:
Hydrogen bonds can vary in strength from weak (1–2 kJ/mol) to strong (161.5 kJ/mol in the
158:
that have much weaker hydrogen bonds. Intramolecular hydrogen bonding is partly responsible for the
3727:
2181:
2101:
720:
3988:
3891:
3863:
3633:
2357:
Friebolin, H., "Basic One- and Two- Dimensional NMR Spectroscopy, 4th ed.," VCH: Weinheim, 2008.
2066:
1347:
541:
260:
191:
2176:
1242:
1153:
2825:
88:
of electrons—the hydrogen bond acceptor (Ac). Such an interacting system is generally denoted
4032:
3993:
3103:
Laage, Damien; Hynes, James T. (2006). "A Molecular Jump Mechanism for Water Reorientation".
2202:
1921:
1858:
1391:
724:
155:
4027:
3951:
3842:
3705:
3664:
3507:
Lipinski CA (December 2004). "Lead- and drug-like compounds: the rule-of-five revolution".
3423:
3354:
3165:
3114:
2981:
2945:
2790:
2739:
2667:
2484:
2425:
2297:
2168:
1648:
1542:
1479:
1387:
672:
661:
624:
202:
119:
608:
Hydrogen bonding is of persistent theoretical interest. According to a modern description
241:
Examples of hydrogen bond donating (donors) and hydrogen bond accepting groups (acceptors)
8:
3853:
3717:
3683:
1419:
159:
123:
35:
4017:
3427:
3358:
3310:
3285:
3169:
3118:
2985:
2949:
2794:
2743:
2671:
2488:
2429:
2301:
2172:
1652:
1627:"Intermolecular Contrast in Atomic Force Microscopy Images without Intermolecular Bonds"
1546:
1483:
832:—which has three lone pairs on the F atom but only one H atom—can form only two bonds; (
791:
is due to the high number of hydrogen bonds each molecule can form, relative to its low
638:
Theoretically, the bond strength of the hydrogen bonds can be assessed using NCI index,
3772:
3568:
3543:
3468:
3323:
3181:
3138:
2763:
2553:
2508:
2448:
2413:
2145:
2047:
1745:
1680:
1638:
1558:
1532:
1500:
1467:
1423:
1207:
1203:
691:
654:
163:
3377:
3342:
1688:
930:, and viscosity of many compounds can be explained by the concept of hydrogen bonding.
4003:
3792:
3752:
3742:
3598:
3573:
3524:
3489:
3447:
3439:
3382:
3315:
3266:
3231:
3130:
3105:
3050:
3017:
2997:
2829:
2818:
2755:
2710:
2640:
2603:
2500:
2453:
2391:
2358:
2313:
2243:
2194:
2004:
1971:
1951:
1892:
Sabin, John R. (1971). "Hydrogen bonds involving sulfur. I. Hydrogen sulfide dimer".
1874:
1811:
1787:
1767:
1737:
1672:
1607:
1505:
1394:; however, an understanding of their relationship to the conventional hydrogen bond,
1355:
1024:
stronger than van der Waals forces) is relevant in the properties of many materials.
992:
829:
774:
135:
97:
81:
3327:
3185:
2557:
2051:
1836:
1749:
1562:
1263:
612:
integrates both the intermolecular O:H lone pair ":" nonbond and the intramolecular
138:
like DNA and proteins. Hydrogen bonds are responsible for holding materials such as
4044:
3784:
3757:
3563:
3555:
3516:
3481:
3431:
3372:
3362:
3305:
3297:
3258:
3223:
3173:
3142:
3122:
3083:
3025:
2989:
2953:
2889:
2858:
2798:
2747:
2702:
2675:
2632:
2580:
2545:
2512:
2492:
2443:
2433:
2383:
2340:
2305:
2270:
2235:
2186:
2159:
2110:
2078:
2037:
2029:
1996:
1929:
1901:
1866:
1840:
1831:
1727:
1684:
1664:
1660:
1656:
1597:
1589:
1550:
1495:
1487:
1246:
1238:
596:
30:
2781:
Needham, Paul (2013). "Hydrogen bonding: Homing in on a tricky chemical concept".
2767:
590:
bond. Certain hydrogen bonds - improper hydrogen bonds - show a blue shift of the
4022:
3896:
3767:
3473:
3284:
Hellgren, M.; Kaiser, C.; de Haij, S.; Norberg, A.; Höög, J. O. (December 2007).
3044:
1602:
1415:
1383:
1241:. Furthermore, the dehydration stabilizes the hydrogen bond by destabilizing the
1143:
1077:
988:
977:
631:
hydrogen bond between guanine and cytosine is much stronger in comparison to the
383:
245:
226:
Lewis acid and the acceptor is the Lewis base. Hydrogen bonds are represented as
3520:
2849:
Moore, T. S.; Winmill, T. F. (1912). "The state of amines in aqueous solution".
2802:
2239:
623:
Quantum chemical calculations of the relevant interresidue potential constants (
3931:
3732:
2973:
2679:
1593:
1554:
1403:
792:
347:(21 kJ/mol or 5.0 kcal/mol), illustrated water-water, alcohol-alcohol
187:
115:
69:
3559:
3544:"A 'rule of 0.5' for the metabolite-likeness of approved pharmaceutical drugs"
3301:
3177:
2957:
1624:
1217:
A protein backbone hydrogen bond incompletely shielded from water attack is a
671:
between two or more intermolecular bonds. This is slightly different from the
4059:
3980:
3940:
3827:
3802:
3675:
3656:
3443:
3340:
2714:
2309:
2042:
2024:
1934:
10.1002/(SICI)1521-3773(19980202)37:1/2<75::AID-ANIE75>3.0.CO;2-R
1835:, 2nd ed. (the "Gold Book") (1997). Online corrected version: (2006–) "
1741:
1732:
1715:
1577:
1399:
1279:), showing the hydrogen bonds (dashed) within and between cellulose molecules
1226:
1100:
residues participating in a hydrogen bond occurs regularly between positions
996:
923:
919:
788:
739:
Crystal structure of hexagonal ice. Gray dashed lines indicate hydrogen bonds
711:
695:
679:
665:
545:
402:
206:
127:
93:
77:
3412:
3367:
3126:
2878:"Polarity and ionization from the standpoint of the Lewis theory of valence"
2438:
1844:
995:, which occur even in the gas phase, resulting in gross deviations from the
579: 15.5, which is about 10 ppm downfield of a conventional alcohol.
198:
3926:
3577:
3528:
3493:
3451:
3386:
3319:
3270:
3235:
3134:
3001:
2759:
2751:
2730:
2644:
2504:
2457:
2395:
2317:
2247:
2198:
2008:
1878:
1676:
1611:
1509:
1374:), although in the case of protonated Proton Sponge, the assembly is bent.
1076:
interactions), which link one complementary strand to the other and enable
1037:
694:
has a net negative sum. The initial theory of hydrogen bonding proposed by
675:
668:
171:
810:
134:. This type of bond can occur in inorganic molecules such as water and in
4012:
3762:
2862:
2344:
2114:
1359:
1112:
1073:
799:
769:
687:
3542:
o′Hagan, Steve; Swainston, Neil; Handl, Julia; Kell, Douglas B. (2015).
3283:
3029:
2893:
2496:
2082:
1905:
836:
has the opposite problem: three hydrogen atoms but only one lone pair).
3401:
3227:
2937:
2584:
2549:
2000:
1668:
1491:
1395:
1363:
1135:
1127:
1097:
927:
683:
650:
298:
131:
84:"donor" atom or group (Dn), and another electronegative atom bearing a
27:
Intermolecular attraction between a hydrogen-donor pair and an acceptor
3485:
3435:
3262:
2993:
2706:
2636:
2387:
2274:
2190:
1576:
De Luca, S.; Chen, F.; Seal, P.; Stenzel, M. H.; Smith, S. C. (2017).
3747:
3722:
3341:
Fernández, A.; Rogale K.; Scott Ridgway; Scheraga H. A. (June 2004).
1871:
10.1002/1521-3773(20020104)41:1<48::AID-ANIE48>3.0.CO;2-U
1407:
1284:
1195:
1184:
1069:
1056:
934:
912:
426:
353:(13 kJ/mol or 3.1 kcal/mol), illustrated by ammonia-ammonia
85:
57:
3088:
3071:
2877:
3821:
3467:
1849:
1450:–hydrogen centers. Many drugs do not, however, obey these "rules".
1443:
1439:
1335:
1319:
1300:
1271:
1234:
1218:
1199:
1188:
1052:
981:
768:
molecules with one hydrogen bond between them, which is called the
646:, as its name indicates, using the electron density of the system.
324:(161.5 kJ/mol or 38.6 kcal/mol), illustrated uniquely by
314:
237:
213:
108:
100:
73:
3595:
An Introduction to Hydrogen Bonding (Topics in Physical Chemistry)
3069:
1643:
1537:
194:
dimer complex. The hydrogen bonds are represented by dotted lines.
3625:
1411:
1331:
1048:
833:
167:
3343:"Inhibitor design by wrapping packing defects in HIV-1 proteins"
1987:
Nishio, M (2011). "The CH/ hydrogen bond in chemistry. "Title".
1856:
Steiner, Thomas (2002). "The Hydrogen Bond in the Solid State".
1032:
389:
Hydrogen bonds involving C-H bonds are both very rare and weak.
341:(29 kJ/mol or 6.9 kcal/mol), illustrated water-ammonia
2824:(3rd ed.). Ithaca (NY): Cornell University Press. p.
1447:
1327:
1312:
1288:
1222:
1180:
1043:
784:
104:
1716:"Definition of the hydrogen bond (IUPAC Recommendations 2011)"
540:
Strong hydrogen bonds are revealed by downfield shifts in the
41:
2934:
2692:
1323:
1315:
1304:
1296:
1230:
1221:. Dehydrons promote the removal of water through proteins or
1093:
765:
761:
359:(8 kJ/mol or 1.9 kcal/mol), illustrated water-amide
271:
151:
139:
3248:
3102:
2470:
2152:"Structure and energetics of the hydronium hydration shells"
1002:
Pentamer formation of water and alcohols in apolar solvents.
3155:
2149:
1434:
The Hydrogen bond is relevant to drug design. According to
1382:
The hydrogen bond can be compared with the closely related
1292:
1252:
743:
210:
143:
3833:
2848:
2727:
2407:
2405:
2374:
Hobza P, Havlas Z (2000). "Blue-Shifting Hydrogen Bonds".
1918:
1164:
as the H-bond acceptor and two H-bond donors from residue
1138:
is formed. Hydrogen bonds also play a part in forming the
760:
An ubiquitous example of a hydrogen bond is found between
735:
220:
3541:
2021:
1713:
1065:
778:
544:. For example, the acidic proton in the enol tautomer of
2411:
2131:(Eds.: E. Grushka, N. Grinberg), CRC Press, Boca Raton,
1465:
2528:
2402:
1522:
1468:"Mapping the force field of a hydrogen-bonded assembly"
1115:
is formed. When the spacing is less, between positions
1092:, hydrogen bonds form between the backbone oxygens and
811:
Bifurcated and over-coordinated hydrogen bonds in water
1575:
1410:
groups). In some cases, these proton acceptors may be
710:
The concept of hydrogen bonding once was challenging.
3213:
3014:
2600:
The Attribute of Water: Single Notion, Multiple Myths
845:
660:
Most generally, the hydrogen bond can be viewed as a
556:
730:
2783:
Studies in History and Philosophy of Science Part A
2225:
392:
2817:
913:Further manifestations of solvent hydrogen bonding
901:
569:
2597:
2099:Emsley, J. (1980). "Very Strong Hydrogen Bonds".
1970:; Wiley-VCH, New York, 1998. • Wiley-VCH; 1998)
686:. However, hydrogen bonding is generally still a
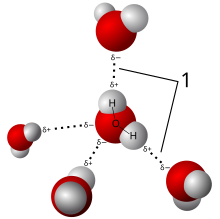
34:Model of hydrogen bonds (1) between molecules of
4057:
3597:. Oxford University Press, US (March 13, 1997).
2970:
2930:
2928:
2876:Latimer, Wendell M.; Rodebush, Worth H. (1920).
582:In the IR spectrum, hydrogen bonding shifts the
3347:Proceedings of the National Academy of Sciences
2875:
2418:Proceedings of the National Academy of Sciences
2287:
1068:is due largely to hydrogen bonding between its
2412:Feldblum, Esther S.; Arkin, Isaiah T. (2014).
1891:
1191:within the hydrophobic membrane environments.
3641:
3500:
3198:
2925:
2780:
2570:
2260:
2064:
1372:[N···H···N]
1368:[F···H···F]
1006:
603:
2657:
973:, and HCl, where hydrogen-bonding is absent.
3506:
2373:
2330:
2127:V. David, N. Grinberg, S. C. Moldoveanu in
1142:through interaction of R-groups. (See also
958:, and HF compared to the heavier analogues
751:, which features two linear hydrogen-bonds.
3648:
3634:
3251:Journal of Chemical Theory and Computation
3049:. Cambridge University Press. p. 37.
2908:"11.9: Water - An Extraordinary Substance"
2622:
2098:
2015:
1786:; Oxford university press New York, 1997.
1341:
118:(occurring between separate molecules) or
3567:
3376:
3366:
3309:
3087:
2573:Chemistry Education Research and Practice
2447:
2437:
2180:
2094:
2092:
2041:
1804:Hydrogen bonding in biological structures
1731:
1642:
1601:
1536:
1499:
1362:at high pressure. It is also seen in the
1225:. The exogenous dehydration enhances the
4040:Polyhedral skeletal electron pair theory
2882:Journal of the American Chemical Society
2695:Journal of the American Chemical Society
2625:Journal of the American Chemical Society
2263:Journal of the American Chemical Society
1270:
1262:
1042:
1031:
742:
734:
642:, which allows a visualization of these
244:
236:
197:
181:
40:
29:
3463:
3461:
2815:
1855:
1429:
1214:sorbitol dehydrogenase protein family.
902:{\displaystyle {\ce {H-F***H-F***H-F}}}
635:bond between the adenine-thymine pair.
221:Definitions and general characteristics
92:, where the solid line denotes a polar
14:
4058:
3042:
3036:
2150:Markovitch, Omer; Agmon, Noam (2007).
2089:
2065:Larson, J. W.; McMahon, T. B. (1984).
2058:
1986:
943:Dramatically higher boiling points of
3629:
3201:Hydrogen Bonding in Polymer Materials
2464:
1275:A strand of cellulose (conformation I
416:
377:(18 kJ/mol or 4.3 kcal/mol)
3458:
3402:https://doi.org/10.1021/jacs.8b08748
3290:Cellular and Molecular Life Sciences
3046:Biophysics: A Physiological Approach
2129:Advances in Chromatography Volume 54
1966:Nishio, M.; Hirota, M.; Umezawa, Y.
1709:
1707:
1705:
1152:systems are common in alpha-helical
616:polar-covalent bond associated with
600:or rotational disorder of the ions.
249:Cyclic dimer of acetic acid; dashed
154:(100 °C) compared to the other
3416:The Journal of Physical Chemistry A
2290:The Journal of Physical Chemistry A
1784:An introduction to hydrogen bonding
1175:and a side-chain hydroxyl or thiol
1096:hydrogens. When the spacing of the
24:
3655:
3587:
3509:Drug Discovery Today: Technologies
2537:Energy & Environmental Science
1832:Compendium of Chemical Terminology
1377:
570:{\displaystyle \delta _{\text{H}}}
50:naphthalenetetracarboxylic diimide
25:
4092:
3608:
2414:"Strength of a bifurcated H bond"
2025:"Definition of the hydrogen bond"
1702:
1258:
731:Hydrogen bonds in small molecules
3832:
3826:
3820:
3621:isotopic effect on bond dynamics
1287:and its derived fibers, such as
823:
425:distance is typically ≈110
399:resonance assisted hydrogen bond
393:Resonance assisted hydrogen bond
292:
3535:
3406:
3393:
3334:
3277:
3242:
3207:
3192:
3149:
3096:
3063:
3008:
2964:
2900:
2869:
2842:
2809:
2774:
2721:
2686:
2651:
2616:
2591:
2564:
2367:
2351:
2324:
2281:
2254:
2219:
2138:
2121:
1980:
1960:
1940:
1912:
1885:
1352:three-center four-electron bond
1090:secondary structure of proteins
1036:The structure of part of a DNA
640:non-covalent interactions index
535:
1820:
1796:
1776:
1756:
1661:10.1103/PhysRevLett.113.186102
1618:
1569:
1516:
1459:
749:nickel bis(dimethylglyoximate)
255:lines represent hydrogen bonds
72:force of attraction between a
13:
1:
1453:
1237:groups by de-shielding their
1140:tertiary structure of protein
717:tetramethylammonium hydroxide
280:. This definition specifies:
2598:Sun, C. Q.; Sun, Yi (2016).
1802:Jeffrey, G. A.; Saenger, W.
937:of mixtures of HF and water.
411:···O=C−C=C−OH
7:
3521:10.1016/j.ddtec.2004.11.007
2803:10.1016/j.shpsa.2012.04.001
2240:10.1021/acs.chemrev.5b00583
1156:between the backbone amide
1083:
10:
4097:
3738:Metal–ligand multiple bond
2680:10.1103/PhysRevLett.82.600
1766:Franklin Classics, 2018),
1720:Pure and Applied Chemistry
1594:10.1021/acs.biomac.7b00657
1555:10.1103/PhysRevB.90.085421
1007:Hydrogen bonds in polymers
705:
604:Theoretical considerations
277:Pure and Applied Chemistry
177:
4002:
3979:
3910:
3872:
3852:
3841:
3818:
3801:
3783:
3674:
3663:
3560:10.1007/s11306-014-0733-z
3302:10.1007/s00018-007-7318-1
3178:10.1080/00268970701877921
2958:10.1080/00268978500103111
1299:, hydrogen bonds between
1245:consisting of dehydrated
1047:Hydrogen bonding between
991:and hexamer formation in
644:non-covalent interactions
124:van der Waals interaction
4076:Supramolecular chemistry
2333:Chemical Society Reviews
2310:10.1021/acs.jpca.7b09425
2102:Chemical Society Reviews
2043:10.1351/PAC-REC-10-01-02
1733:10.1351/PAC-REC-10-01-02
1326:are more sensitive than
1229:interaction between the
755:
407:O=C−OH···
126:, and weaker than fully
90:Dn−H···Ac
3368:10.1073/pnas.0404641101
3127:10.1126/science.1122154
2660:Physical Review Letters
2439:10.1073/pnas.1319827111
1845:10.1351/goldbook.H02899
1631:Physical Review Letters
1436:Lipinski's rule of five
1348:symmetric hydrogen bond
1342:Symmetric hydrogen bond
976:Viscosity of anhydrous
649:Interpretations of the
454:HCN···HF
357:N−H···:O
351:N−H···:N
345:O−H···:O
339:O−H···:N
322:F−H···:F
3199:Shiao-Wei Kuo (2018).
3043:Dillon, P. F. (2012).
2752:10.1006/jmre.1999.1899
1989:Phys. Chem. Chem. Phys
1280:
1268:
1154:transmembrane proteins
1060:
1055:, one of two types of
1040:
1027:
903:
752:
740:
690:phenomenon, since the
633:N−H···N
629:N−H···N
571:
471:CO···HF
290:
256:
242:
217:
195:
190:hydrogen bonding in a
114:Hydrogen bonds can be
53:
38:
4081:Intermolecular forces
1968:The CH–π Interactions
1922:Angew. Chem. Int. Ed.
1859:Angew. Chem. Int. Ed.
1603:1959.4/unsworks_55160
1472:Nature Communications
1274:
1267:Para-aramid structure
1266:
1171:: the backbone amide
1046:
1035:
904:
746:
738:
725:Maurice Loyal Huggins
572:
505:S···HF
488:O···HF
372:···:OH
282:
248:
240:
201:
185:
44:
33:
3728:Coordinate (dipolar)
2912:Chemistry LibreTexts
2863:10.1039/CT9120101635
2816:Pauling, L. (1960).
2345:10.1039/CS9871600467
2144:Data obtained using
2115:10.1039/cs9800900091
1806:; Springer: Berlin,
1430:Application to drugs
1334:more sensitive than
843:
625:compliance constants
620:repulsive coupling.
554:
522:···HF
431:H···Y
228:H···Y
209:helps stabilize the
205:hydrogen bonding in
3902:C–H···O interaction
3684:Electron deficiency
3593:George A. Jeffrey.
3469:Crabtree, Robert H.
3428:2005JPCA..109.1637O
3359:2004PNAS..10111640F
3170:2008MolPh.106..485M
3119:2006Sci...311..832L
3030:10.1021/ja00274a058
2986:2005JChPh.123j4501Z
2950:1985MolPh..56.1381J
2894:10.1021/ja01452a015
2795:2013SHPSA..44...51N
2744:1999JMagR.140..510C
2672:1999PhRvL..82..600I
2497:10.1038/nature03383
2489:2005Natur.434..199C
2430:2014PNAS..111.4085F
2302:2017JPCA..121.8535L
2173:2007JPCA..111.2253M
2083:10.1021/ic00182a010
2071:Inorganic Chemistry
1995:(31): 13873–13900.
1906:10.1021/ja00744a012
1653:2014PhRvL.113r6102H
1547:2014PhRvB..90h5421H
1484:2014NatCo...5.3931S
1422:has shown that the
1420:Neutron diffraction
1386:, which is also an
987:Dimer formation in
3887:Resonance-assisted
3228:10.1039/C0CC01763A
2585:10.1039/c4rp00030g
2550:10.1039/C4EE02280G
2146:molecular dynamics
2001:10.1039/c1cp20404a
1492:10.1038/ncomms4931
1424:molecular geometry
1281:
1269:
1208:molecular dynamics
1204:circular dichroism
1061:
1041:
899:
753:
741:
692:interaction energy
567:
417:Structural details
317:in vapor include:
257:
243:
232:associated liquids
218:
196:
76:(H) atom which is
68:) is primarily an
54:
39:
4053:
4052:
4004:Electron counting
3975:
3974:
3864:London dispersion
3816:
3815:
3793:Metal aromaticity
3486:10.1021/ar950150s
3436:10.1021/jp040618l
3263:10.1021/ct200455n
3257:(11): 3816–3828.
3222:(35): 6449–6451.
3158:Molecular Physics
3056:978-1-139-50462-1
3018:J. Am. Chem. Soc.
2994:10.1063/1.2018637
2835:978-0-8014-0333-0
2707:10.1021/ja9937019
2637:10.1021/ja046282a
2609:978-981-10-0178-9
2483:(7030): 199–202.
2424:(11): 4085–4090.
2388:10.1021/cr990050q
2382:(11): 4253–4264.
2363:978-3-527-31233-7
2296:(44): 8535–8541.
2275:10.1021/ja991795g
2269:(40): 9411–9422.
2191:10.1021/jp068960g
2167:(12): 2253–2256.
2077:(14): 2029–2033.
1900:(15): 3613–3620.
1764:The Hydrogen Bond
1588:(11): 3665–3677.
1582:Biomacromolecules
1525:Physical Review B
1356:hydrofluoric acid
1150:Bifurcated H-bond
993:hydrogen fluoride
897:
889:
877:
869:
857:
849:
830:hydrogen fluoride
775:crystal structure
678:of, for example,
597:phase transitions
564:
533:
532:
156:group-16 hydrides
136:organic molecules
98:period 2 elements
78:covalently bonded
16:(Redirected from
4088:
4071:Hydrogen physics
4066:Chemical bonding
4045:Jemmis mno rules
3897:Dihydrogen bonds
3850:
3849:
3836:
3830:
3824:
3758:Hyperconjugation
3672:
3671:
3650:
3643:
3636:
3627:
3626:
3582:
3581:
3571:
3539:
3533:
3532:
3504:
3498:
3497:
3465:
3456:
3455:
3422:(8): 1637–1642.
3410:
3404:
3397:
3391:
3390:
3380:
3370:
3338:
3332:
3331:
3313:
3281:
3275:
3274:
3246:
3240:
3239:
3211:
3205:
3204:
3196:
3190:
3189:
3153:
3147:
3146:
3100:
3094:
3093:
3091:
3067:
3061:
3060:
3040:
3034:
3033:
3012:
3006:
3005:
2968:
2962:
2961:
2932:
2923:
2922:
2920:
2919:
2904:
2898:
2897:
2888:(7): 1419–1433.
2873:
2867:
2866:
2846:
2840:
2839:
2823:
2813:
2807:
2806:
2778:
2772:
2771:
2725:
2719:
2718:
2701:(6): 1210–1214.
2690:
2684:
2683:
2655:
2649:
2648:
2620:
2614:
2613:
2595:
2589:
2588:
2568:
2562:
2561:
2526:
2517:
2516:
2468:
2462:
2461:
2451:
2441:
2409:
2400:
2399:
2371:
2365:
2355:
2349:
2348:
2328:
2322:
2321:
2285:
2279:
2278:
2258:
2252:
2251:
2228:Chemical Reviews
2223:
2217:
2216:
2214:
2213:
2207:
2201:. Archived from
2184:
2160:J. Phys. Chem. A
2156:
2142:
2136:
2125:
2119:
2118:
2096:
2087:
2086:
2062:
2056:
2055:
2045:
2036:(8): 1637–1641.
2030:Pure Appl. Chem.
2019:
2013:
2012:
1984:
1978:
1964:
1958:
1944:
1938:
1937:
1916:
1910:
1909:
1894:J. Am. Chem. Soc
1889:
1883:
1882:
1853:
1847:
1824:
1818:
1800:
1794:
1782:Jeffrey, G. A.;
1780:
1774:
1760:
1754:
1753:
1735:
1726:(8): 1637–1641.
1711:
1700:
1699:
1697:
1696:
1687:. Archived from
1646:
1622:
1616:
1615:
1605:
1573:
1567:
1566:
1540:
1520:
1514:
1513:
1503:
1463:
1373:
1369:
1247:isolated charges
1178:
1174:
1170:
1159:
1125:
1110:
1022:
1018:
1014:
989:carboxylic acids
972:
964:
957:
949:
918:Increase in the
908:
906:
905:
900:
898:
895:
894:
887:
875:
874:
867:
855:
854:
847:
634:
630:
619:
615:
611:
593:
589:
585:
578:
576:
574:
573:
568:
566:
565:
562:
523:
506:
489:
472:
455:
442:
441:Acceptor···donor
437:
436:
432:
424:
412:
408:
376:
371:
370:
367:
358:
352:
346:
340:
335:
334:
333:
330:
323:
312:
311:
310:
307:
287:
254:
229:
91:
21:
4096:
4095:
4091:
4090:
4089:
4087:
4086:
4085:
4056:
4055:
4054:
4049:
3998:
3971:
3914:
3906:
3868:
3855:
3845:
3837:
3831:
3825:
3812:
3797:
3779:
3667:
3659:
3654:
3615:The Bubble Wall
3611:
3590:
3588:Further reading
3585:
3540:
3536:
3505:
3501:
3474:Acc. Chem. Res.
3466:
3459:
3411:
3407:
3398:
3394:
3353:(32): 11640–5.
3339:
3335:
3296:(23): 3129–38.
3282:
3278:
3247:
3243:
3212:
3208:
3197:
3193:
3154:
3150:
3113:(5762): 832–5.
3101:
3097:
3089:10.1139/v84-087
3068:
3064:
3057:
3041:
3037:
3013:
3009:
2969:
2965:
2933:
2926:
2917:
2915:
2906:
2905:
2901:
2874:
2870:
2847:
2843:
2836:
2814:
2810:
2779:
2775:
2726:
2722:
2691:
2687:
2656:
2652:
2631:(50): 16310–1.
2621:
2617:
2610:
2596:
2592:
2569:
2565:
2527:
2520:
2474:
2469:
2465:
2410:
2403:
2372:
2368:
2356:
2352:
2329:
2325:
2286:
2282:
2259:
2255:
2234:(9): 5216–300.
2224:
2220:
2211:
2209:
2205:
2154:
2143:
2139:
2126:
2122:
2097:
2090:
2063:
2059:
2020:
2016:
1985:
1981:
1965:
1961:
1945:
1941:
1917:
1913:
1890:
1886:
1854:
1850:
1825:
1821:
1801:
1797:
1781:
1777:
1761:
1757:
1712:
1703:
1694:
1692:
1623:
1619:
1574:
1570:
1521:
1517:
1464:
1460:
1456:
1432:
1416:metal complexes
1392:crystallography
1384:dihydrogen bond
1380:
1378:Dihydrogen bond
1371:
1367:
1344:
1278:
1261:
1243:nonbonded state
1239:partial charges
1176:
1172:
1165:
1157:
1144:protein folding
1131:
1120:
1105:
1086:
1030:
1020:
1016:
1012:
1009:
978:phosphoric acid
970:
966:
963:
959:
955:
951:
948:
944:
915:
890:
870:
850:
846:
844:
841:
840:
826:
813:
758:
733:
708:
655:Compton profile
632:
628:
617:
613:
609:
606:
591:
587:
583:
561:
557:
555:
552:
551:
549:
538:
521:
517:
504:
500:
487:
483:
475:trigonal planar
470:
466:
453:
440:
430:
422:
419:
410:
406:
395:
384:crystallography
375:
368:
365:
364:
362:
356:
350:
344:
338:
331:
328:
327:
325:
321:
308:
305:
304:
302:
295:
285:
250:
227:
223:
180:
89:
82:electronegative
28:
23:
22:
15:
12:
11:
5:
4094:
4084:
4083:
4078:
4073:
4068:
4051:
4050:
4048:
4047:
4042:
4037:
4036:
4035:
4030:
4025:
4020:
4009:
4007:
4000:
3999:
3997:
3996:
3991:
3985:
3983:
3977:
3976:
3973:
3972:
3970:
3969:
3964:
3959:
3954:
3949:
3944:
3934:
3929:
3924:
3918:
3916:
3908:
3907:
3905:
3904:
3899:
3894:
3889:
3884:
3878:
3876:
3870:
3869:
3867:
3866:
3860:
3858:
3847:
3843:Intermolecular
3839:
3838:
3819:
3817:
3814:
3813:
3811:
3810:
3807:
3805:
3799:
3798:
3796:
3795:
3789:
3787:
3781:
3780:
3778:
3777:
3776:
3775:
3770:
3760:
3755:
3750:
3745:
3740:
3735:
3730:
3725:
3720:
3715:
3714:
3713:
3703:
3702:
3701:
3696:
3691:
3680:
3678:
3669:
3665:Intramolecular
3661:
3660:
3657:Chemical bonds
3653:
3652:
3645:
3638:
3630:
3624:
3623:
3618:
3610:
3609:External links
3607:
3606:
3605:
3589:
3586:
3584:
3583:
3554:(2): 323–339.
3534:
3515:(4): 337–341.
3499:
3480:(7): 348–354.
3457:
3405:
3392:
3333:
3276:
3241:
3206:
3191:
3148:
3095:
3082:(3): 526–530.
3062:
3055:
3035:
3007:
2980:(10): 104501.
2974:J. Chem. Phys.
2963:
2924:
2899:
2868:
2841:
2834:
2808:
2773:
2720:
2685:
2666:(3): 600–603.
2650:
2615:
2608:
2590:
2579:(3): 276–285.
2563:
2518:
2472:
2463:
2401:
2366:
2350:
2323:
2280:
2253:
2218:
2182:10.1.1.76.9448
2137:
2120:
2088:
2057:
2014:
1979:
1959:
1939:
1928:(1–2): 75–78.
1911:
1884:
1848:
1819:
1795:
1775:
1755:
1701:
1637:(18): 186102.
1617:
1568:
1515:
1457:
1455:
1452:
1431:
1428:
1388:intermolecular
1379:
1376:
1343:
1340:
1276:
1260:
1259:Other polymers
1257:
1223:ligand binding
1129:
1085:
1082:
1029:
1026:
1008:
1005:
1004:
1003:
1000:
985:
974:
968:
961:
953:
946:
941:
938:
931:
914:
911:
910:
909:
893:
886:
883:
880:
873:
866:
863:
860:
853:
825:
822:
812:
809:
793:molecular mass
757:
754:
732:
729:
707:
704:
700:NMR techniques
673:intramolecular
605:
602:
560:
542:H NMR spectrum
537:
534:
531:
530:
527:
524:
519:
514:
513:
510:
507:
502:
497:
496:
493:
490:
485:
480:
479:
476:
473:
468:
463:
462:
459:
456:
450:
449:
446:
445:VSEPR geometry
443:
429:, whereas the
418:
415:
394:
391:
379:
378:
373:
360:
354:
348:
342:
336:
294:
291:
222:
219:
203:Intramolecular
192:self-assembled
188:intermolecular
186:An example of
179:
176:
166:structures of
120:intramolecular
116:intermolecular
26:
9:
6:
4:
3:
2:
4093:
4082:
4079:
4077:
4074:
4072:
4069:
4067:
4064:
4063:
4061:
4046:
4043:
4041:
4038:
4034:
4031:
4029:
4026:
4024:
4021:
4019:
4018:Hückel's rule
4016:
4015:
4014:
4011:
4010:
4008:
4005:
4001:
3995:
3992:
3990:
3987:
3986:
3984:
3982:
3981:Bond cleavage
3978:
3968:
3965:
3963:
3960:
3958:
3955:
3953:
3950:
3948:
3947:Intercalation
3945:
3942:
3938:
3937:Metallophilic
3935:
3933:
3930:
3928:
3925:
3923:
3920:
3919:
3917:
3913:
3909:
3903:
3900:
3898:
3895:
3893:
3890:
3888:
3885:
3883:
3880:
3879:
3877:
3875:
3871:
3865:
3862:
3861:
3859:
3857:
3854:Van der Waals
3851:
3848:
3844:
3840:
3835:
3829:
3823:
3809:
3808:
3806:
3804:
3800:
3794:
3791:
3790:
3788:
3786:
3782:
3774:
3771:
3769:
3766:
3765:
3764:
3761:
3759:
3756:
3754:
3751:
3749:
3746:
3744:
3741:
3739:
3736:
3734:
3731:
3729:
3726:
3724:
3721:
3719:
3716:
3712:
3709:
3708:
3707:
3704:
3700:
3697:
3695:
3692:
3690:
3687:
3686:
3685:
3682:
3681:
3679:
3677:
3673:
3670:
3666:
3662:
3658:
3651:
3646:
3644:
3639:
3637:
3632:
3631:
3628:
3622:
3619:
3616:
3613:
3612:
3604:
3603:0-19-509549-9
3600:
3596:
3592:
3591:
3579:
3575:
3570:
3565:
3561:
3557:
3553:
3549:
3545:
3538:
3530:
3526:
3522:
3518:
3514:
3510:
3503:
3495:
3491:
3487:
3483:
3479:
3476:
3475:
3470:
3464:
3462:
3453:
3449:
3445:
3441:
3437:
3433:
3429:
3425:
3421:
3417:
3409:
3403:
3396:
3388:
3384:
3379:
3374:
3369:
3364:
3360:
3356:
3352:
3348:
3344:
3337:
3329:
3325:
3321:
3317:
3312:
3307:
3303:
3299:
3295:
3291:
3287:
3280:
3272:
3268:
3264:
3260:
3256:
3252:
3245:
3237:
3233:
3229:
3225:
3221:
3217:
3210:
3202:
3195:
3187:
3183:
3179:
3175:
3171:
3167:
3163:
3159:
3152:
3144:
3140:
3136:
3132:
3128:
3124:
3120:
3116:
3112:
3108:
3107:
3099:
3090:
3085:
3081:
3077:
3073:
3066:
3058:
3052:
3048:
3047:
3039:
3031:
3027:
3023:
3020:
3019:
3011:
3003:
2999:
2995:
2991:
2987:
2983:
2979:
2976:
2975:
2967:
2959:
2955:
2951:
2947:
2943:
2940:
2939:
2931:
2929:
2913:
2909:
2903:
2895:
2891:
2887:
2883:
2879:
2872:
2864:
2860:
2856:
2852:
2845:
2837:
2831:
2827:
2822:
2821:
2812:
2804:
2800:
2796:
2792:
2788:
2784:
2777:
2769:
2765:
2761:
2757:
2753:
2749:
2745:
2741:
2737:
2733:
2732:
2724:
2716:
2712:
2708:
2704:
2700:
2696:
2689:
2681:
2677:
2673:
2669:
2665:
2661:
2654:
2646:
2642:
2638:
2634:
2630:
2626:
2619:
2611:
2605:
2601:
2594:
2586:
2582:
2578:
2574:
2567:
2559:
2555:
2551:
2547:
2543:
2539:
2538:
2533:
2525:
2523:
2514:
2510:
2506:
2502:
2498:
2494:
2490:
2486:
2482:
2478:
2467:
2459:
2455:
2450:
2445:
2440:
2435:
2431:
2427:
2423:
2419:
2415:
2408:
2406:
2397:
2393:
2389:
2385:
2381:
2377:
2370:
2364:
2360:
2354:
2346:
2342:
2338:
2334:
2327:
2319:
2315:
2311:
2307:
2303:
2299:
2295:
2291:
2284:
2276:
2272:
2268:
2264:
2257:
2249:
2245:
2241:
2237:
2233:
2229:
2222:
2208:on 2014-08-13
2204:
2200:
2196:
2192:
2188:
2183:
2178:
2174:
2170:
2166:
2162:
2161:
2153:
2147:
2141:
2134:
2130:
2124:
2116:
2112:
2109:(1): 91–124.
2108:
2104:
2103:
2095:
2093:
2084:
2080:
2076:
2072:
2068:
2061:
2053:
2049:
2044:
2039:
2035:
2032:
2031:
2026:
2018:
2010:
2006:
2002:
1998:
1994:
1990:
1983:
1977:
1973:
1969:
1963:
1957:
1953:
1949:
1943:
1935:
1931:
1927:
1924:
1923:
1915:
1907:
1903:
1899:
1895:
1888:
1880:
1876:
1872:
1868:
1864:
1861:
1860:
1852:
1846:
1842:
1838:
1837:hydrogen bond
1834:
1833:
1828:
1823:
1817:
1813:
1809:
1805:
1799:
1793:
1789:
1785:
1779:
1773:
1769:
1765:
1762:Pimentel, G.
1759:
1751:
1747:
1743:
1739:
1734:
1729:
1725:
1721:
1717:
1710:
1708:
1706:
1691:on 2018-01-20
1690:
1686:
1682:
1678:
1674:
1670:
1666:
1662:
1658:
1654:
1650:
1645:
1640:
1636:
1632:
1628:
1621:
1613:
1609:
1604:
1599:
1595:
1591:
1587:
1583:
1579:
1572:
1564:
1560:
1556:
1552:
1548:
1544:
1539:
1534:
1531:(8): 085421.
1530:
1526:
1519:
1511:
1507:
1502:
1497:
1493:
1489:
1485:
1481:
1477:
1473:
1469:
1462:
1458:
1451:
1449:
1445:
1441:
1437:
1427:
1425:
1421:
1417:
1413:
1409:
1405:
1401:
1400:covalent bond
1397:
1393:
1389:
1385:
1375:
1365:
1361:
1357:
1353:
1349:
1339:
1337:
1333:
1329:
1325:
1321:
1317:
1314:
1310:
1306:
1302:
1298:
1294:
1290:
1286:
1273:
1265:
1256:
1254:
1250:
1248:
1244:
1240:
1236:
1232:
1228:
1227:electrostatic
1224:
1220:
1215:
1211:
1209:
1205:
1201:
1197:
1192:
1190:
1186:
1182:
1168:
1163:
1155:
1151:
1147:
1145:
1141:
1137:
1133:
1123:
1118:
1114:
1108:
1103:
1099:
1095:
1091:
1081:
1079:
1075:
1071:
1067:
1058:
1054:
1050:
1045:
1039:
1034:
1025:
1001:
998:
997:ideal gas law
994:
990:
986:
983:
979:
975:
942:
939:
936:
932:
929:
925:
924:boiling point
921:
920:melting point
917:
916:
891:
884:
881:
878:
871:
864:
861:
858:
851:
839:
838:
837:
835:
831:
828:For example,
824:Other liquids
821:
817:
808:
804:
801:
796:
794:
790:
789:boiling point
787:water's high
786:
782:
780:
776:
771:
767:
763:
750:
747:Structure of
745:
737:
728:
726:
722:
718:
713:
712:Linus Pauling
703:
701:
697:
696:Linus Pauling
693:
689:
685:
681:
677:
674:
670:
667:
666:electrostatic
663:
658:
656:
652:
647:
645:
641:
636:
626:
621:
601:
598:
580:
558:
547:
546:acetylacetone
543:
528:
525:
516:
515:
511:
508:
499:
498:
494:
491:
482:
481:
477:
474:
465:
464:
460:
457:
452:
451:
447:
444:
439:
438:
435:
428:
414:
404:
403:electrostatic
400:
390:
387:
385:
361:
355:
349:
343:
337:
320:
319:
318:
316:
300:
293:Bond strength
289:
281:
279:
278:
273:
268:
264:
262:
261:London forces
253:
247:
239:
235:
233:
215:
212:
208:
207:acetylacetone
204:
200:
193:
189:
184:
175:
173:
172:nucleic acids
169:
165:
161:
157:
153:
147:
145:
141:
137:
133:
129:
125:
121:
117:
112:
110:
106:
102:
99:
95:
94:covalent bond
87:
83:
79:
75:
71:
70:electrostatic
67:
63:
62:hydrogen bond
59:
51:
47:
43:
37:
32:
19:
18:Hydrogen-bond
4023:Baird's rule
3873:
3743:Charge-shift
3706:Hypervalence
3594:
3551:
3548:Metabolomics
3547:
3537:
3512:
3508:
3502:
3477:
3472:
3419:
3415:
3408:
3395:
3350:
3346:
3336:
3293:
3289:
3279:
3254:
3250:
3244:
3219:
3215:
3209:
3203:. Wiley-VCH.
3200:
3194:
3161:
3157:
3151:
3110:
3104:
3098:
3079:
3076:Can. J. Chem
3075:
3065:
3045:
3038:
3024:(14): 4196.
3021:
3016:
3010:
2977:
2972:
2966:
2941:
2936:
2916:. Retrieved
2914:. 2017-06-29
2911:
2902:
2885:
2881:
2871:
2854:
2851:J. Chem. Soc
2850:
2844:
2819:
2811:
2789:(1): 51–65.
2786:
2782:
2776:
2738:(2): 510–2.
2735:
2731:J Magn Reson
2729:
2723:
2698:
2694:
2688:
2663:
2659:
2653:
2628:
2624:
2618:
2602:. Springer.
2599:
2593:
2576:
2572:
2566:
2541:
2535:
2480:
2476:
2466:
2421:
2417:
2379:
2375:
2369:
2353:
2336:
2332:
2326:
2293:
2289:
2283:
2266:
2262:
2256:
2231:
2227:
2221:
2210:. Retrieved
2203:the original
2164:
2158:
2140:
2135:, chapter 3.
2132:
2128:
2123:
2106:
2100:
2074:
2070:
2060:
2033:
2028:
2017:
1992:
1988:
1982:
1967:
1962:
1947:
1942:
1925:
1920:
1914:
1897:
1893:
1887:
1865:(1): 48–76.
1862:
1857:
1851:
1830:
1822:
1807:
1803:
1798:
1783:
1778:
1763:
1758:
1723:
1719:
1693:. Retrieved
1689:the original
1634:
1630:
1620:
1585:
1581:
1571:
1528:
1524:
1518:
1475:
1471:
1461:
1433:
1381:
1345:
1308:
1282:
1251:
1216:
1212:
1193:
1166:
1161:
1148:
1121:
1116:
1106:
1101:
1087:
1072:(as well as
1062:
1038:double helix
1010:
827:
818:
814:
805:
797:
783:
759:
709:
676:bound states
669:scalar field
659:
651:anisotropies
648:
637:
622:
607:
581:
539:
536:Spectroscopy
420:
398:
396:
388:
380:
296:
283:
275:
269:
265:
258:
251:
231:
224:
148:
113:
65:
61:
55:
4013:Aromaticity
3989:Heterolysis
3967:Salt bridge
3912:Noncovalent
3882:Low-barrier
3763:Aromaticity
3753:Conjugation
3733:Pi backbond
2944:(6): 1381.
2544:(4): 1276.
1669:1874/307996
1360:formic acid
1160:of residue
1113:alpha helix
1078:replication
1074:pi stacking
770:water dimer
688:bound state
684:ionic bonds
664:-dependent
548:appears at
313:). Typical
144:felted wool
132:ionic bonds
4060:Categories
3941:aurophilic
3922:Mechanical
3164:(2): 485.
2938:Mol. Phys.
2918:2024-06-22
2212:2017-10-25
1976:0471252905
1956:0198509707
1816:3540579036
1810:Springer;
1808:1994, 2012
1792:0195095499
1772:0343171600
1695:2017-08-30
1454:References
1396:ionic bond
1364:bifluoride
1136:beta sheet
1098:amino acid
1070:base pairs
1057:base pairs
928:solubility
448:Angle (°)
315:enthalpies
299:bifluoride
80:to a more
4033:spherical
3994:Homolysis
3957:Cation–pi
3932:Chalcogen
3892:Symmetric
3748:Hapticity
3444:1089-5639
2715:0002-7863
2376:Chem. Rev
2177:CiteSeerX
1742:1365-3075
1644:1410.1933
1538:1406.3562
1408:chalcogen
1285:cellulose
1196:trehalose
1185:threonine
1126:, then a
935:azeotropy
933:Negative
892:−
885:⋅
882:⋅
879:⋅
872:−
865:⋅
862:⋅
859:⋅
852:−
559:δ
509:pyramidal
492:pyramidal
160:secondary
107:(O), and
86:lone pair
58:chemistry
48:image of
3962:Anion–pi
3952:Stacking
3874:Hydrogen
3785:Metallic
3676:Covalent
3668:(strong)
3578:25750602
3529:24981612
3494:19904922
3452:16833488
3387:15289598
3328:22090973
3320:17952367
3311:11136444
3271:26598272
3236:20657920
3216:ChemComm
3186:17648714
3135:16439623
3002:16178604
2857:: 1635.
2760:10497060
2645:15600318
2558:84176511
2505:15758995
2458:24591597
2396:11749346
2318:29048895
2248:27136957
2199:17388314
2052:97688573
2009:21611676
1879:12491444
1750:97688573
1677:25396382
1612:28880549
1563:53610973
1510:24875276
1478:: 3931.
1444:hydrogen
1440:nitrogen
1412:pi-bonds
1404:nitrogen
1336:nylon-11
1320:humidity
1303:and the
1301:carbonyl
1235:carbonyl
1219:dehydron
1200:sorbitol
1189:cysteine
1084:Proteins
1053:cytosine
982:glycerol
680:covalent
526:trigonal
214:tautomer
168:proteins
164:tertiary
128:covalent
109:fluorine
101:nitrogen
74:hydrogen
3927:Halogen
3773:bicyclo
3718:Agostic
3569:4342520
3424:Bibcode
3355:Bibcode
3166:Bibcode
3143:6707413
3115:Bibcode
3106:Science
2982:Bibcode
2946:Bibcode
2791:Bibcode
2740:Bibcode
2668:Bibcode
2513:4396493
2485:Bibcode
2449:3964065
2426:Bibcode
2339:: 467.
2298:Bibcode
2169:Bibcode
1685:8309018
1649:Bibcode
1543:Bibcode
1501:4050271
1480:Bibcode
1332:nylon 6
1328:aramids
1088:In the
1049:guanine
980:and of
834:ammonia
721:Latimer
706:History
653:in the
577:
550:
178:Bonding
4028:Möbius
3856:forces
3846:(weak)
3601:
3576:
3566:
3527:
3492:
3450:
3442:
3385:
3378:511032
3375:
3326:
3318:
3308:
3269:
3234:
3184:
3141:
3133:
3053:
3000:
2832:
2768:121429
2766:
2758:
2713:
2643:
2606:
2556:
2511:
2503:
2477:Nature
2456:
2446:
2394:
2361:
2316:
2246:
2197:
2179:
2050:
2007:
1974:
1954:
1877:
1814:
1790:
1770:
1748:
1740:
1683:
1675:
1610:
1561:
1508:
1498:
1448:oxygen
1406:, and
1398:, and
1330:, and
1324:nylons
1313:aramid
1289:cotton
1187:, and
1181:serine
1059:in DNA
1019:, and
785:Liquid
662:metric
458:linear
105:oxygen
66:H-bond
4006:rules
3915:other
3803:Ionic
3711:3c–4e
3699:8c–2e
3694:4c–2e
3689:3c–2e
3324:S2CID
3182:S2CID
3139:S2CID
2764:S2CID
2554:S2CID
2509:S2CID
2206:(PDF)
2155:(PDF)
2048:S2CID
1827:IUPAC
1746:S2CID
1681:S2CID
1639:arXiv
1559:S2CID
1533:arXiv
1316:fibre
1305:amide
1297:nylon
1295:. In
1231:amide
1132:helix
1111:, an
1094:amide
800:TIP4P
766:water
762:water
756:Water
610:O:H−O
301:ion,
272:IUPAC
252:green
152:water
140:paper
111:(F).
103:(N),
36:water
3768:homo
3723:Bent
3599:ISBN
3574:PMID
3525:PMID
3490:PMID
3448:PMID
3440:ISSN
3383:PMID
3316:PMID
3267:PMID
3232:PMID
3131:PMID
3051:ISBN
2998:PMID
2830:ISBN
2756:PMID
2711:ISSN
2641:PMID
2604:ISBN
2501:PMID
2475:O".
2454:PMID
2392:PMID
2359:ISBN
2314:PMID
2244:PMID
2195:PMID
2133:2018
2005:PMID
1972:ISBN
1952:ISBN
1948:2001
1875:PMID
1812:ISBN
1788:ISBN
1768:ISBN
1738:ISSN
1673:PMID
1608:PMID
1506:PMID
1446:and
1366:ion
1358:and
1293:flax
1291:and
1253:Wool
1233:and
1198:and
1119:and
1104:and
1051:and
529:142
478:120
461:180
421:The
397:The
211:enol
170:and
162:and
142:and
64:(or
60:, a
3564:PMC
3556:doi
3517:doi
3482:doi
3432:doi
3420:109
3373:PMC
3363:doi
3351:101
3306:PMC
3298:doi
3259:doi
3224:doi
3174:doi
3162:106
3123:doi
3111:311
3084:doi
3026:doi
3022:108
2990:doi
2978:123
2954:doi
2890:doi
2859:doi
2855:101
2826:450
2799:doi
2748:doi
2736:140
2703:doi
2699:122
2676:doi
2633:doi
2629:126
2581:doi
2546:doi
2493:doi
2481:434
2444:PMC
2434:doi
2422:111
2384:doi
2380:100
2341:doi
2306:doi
2294:121
2271:doi
2267:121
2236:doi
2232:116
2187:doi
2165:111
2111:doi
2079:doi
2038:doi
1997:doi
1930:doi
1902:doi
1867:doi
1841:doi
1839:".
1728:doi
1665:hdl
1657:doi
1635:113
1598:hdl
1590:doi
1551:doi
1496:PMC
1488:doi
1414:or
1173:N−H
1169:+ 4
1158:C=O
1146:).
1124:+ 3
1109:+ 4
1066:DNA
1028:DNA
1021:C−N
1017:C−O
1013:C−C
779:ice
777:of
682:or
618:O−O
614:H−O
592:X−H
588:X−H
584:X−H
512:89
495:46
423:X−H
409:or
286:X−H
263:).
130:or
56:In
46:AFM
4062::
3572:.
3562:.
3552:11
3550:.
3546:.
3523:.
3511:.
3488:.
3478:29
3460:^
3446:.
3438:.
3430:.
3418:.
3381:.
3371:.
3361:.
3349:.
3345:.
3322:.
3314:.
3304:.
3294:64
3292:.
3288:.
3265:.
3253:.
3230:.
3220:46
3218:.
3180:.
3172:.
3160:.
3137:.
3129:.
3121:.
3109:.
3080:62
3078:.
3074:.
2996:.
2988:.
2952:.
2942:56
2927:^
2910:.
2886:42
2884:.
2880:.
2853:.
2828:.
2797:.
2787:44
2785:.
2762:.
2754:.
2746:.
2734:.
2709:.
2697:.
2674:.
2664:82
2662:.
2639:.
2627:.
2577:15
2575:.
2552:.
2540:.
2534:.
2521:^
2507:.
2499:.
2491:.
2479:.
2452:.
2442:.
2432:.
2420:.
2416:.
2404:^
2390:.
2378:.
2337:16
2335:.
2312:.
2304:.
2292:.
2265:.
2242:.
2230:.
2193:.
2185:.
2175:.
2163:.
2157:.
2105:.
2091:^
2075:23
2073:.
2069:.
2046:.
2034:83
2027:.
2003:.
1993:13
1991:.
1950:,
1926:37
1898:93
1896:.
1873:.
1863:41
1829:,
1744:.
1736:.
1724:83
1722:.
1718:.
1704:^
1679:.
1671:.
1663:.
1655:.
1647:.
1633:.
1629:.
1606:.
1596:.
1586:18
1584:.
1580:.
1557:.
1549:.
1541:.
1529:90
1527:.
1504:.
1494:.
1486:.
1474:.
1470:.
1346:A
1338:.
1249:.
1183:,
1130:10
1080:.
1015:,
965:,
960:PH
950:,
945:NH
926:,
922:,
518:SO
427:pm
413:.
363:OH
326:HF
303:HF
234:.
174:.
3943:)
3939:(
3649:e
3642:t
3635:v
3580:.
3558::
3531:.
3519::
3513:1
3496:.
3484::
3454:.
3434::
3426::
3389:.
3365::
3357::
3330:.
3300::
3273:.
3261::
3255:7
3238:.
3226::
3188:.
3176::
3168::
3145:.
3125::
3117::
3092:.
3086::
3059:.
3032:.
3028::
3004:.
2992::
2984::
2960:.
2956::
2948::
2921:.
2896:.
2892::
2865:.
2861::
2838:.
2805:.
2801::
2793::
2770:.
2750::
2742::
2717:.
2705::
2682:.
2678::
2670::
2647:.
2635::
2612:.
2587:.
2583::
2560:.
2548::
2542:8
2515:.
2495::
2487::
2473:2
2460:.
2436::
2428::
2398:.
2386::
2347:.
2343::
2320:.
2308::
2300::
2277:.
2273::
2250:.
2238::
2215:.
2189::
2171::
2117:.
2113::
2107:9
2085:.
2081::
2054:.
2040::
2011:.
1999::
1936:.
1932::
1908:.
1904::
1881:.
1869::
1843::
1752:.
1730::
1698:.
1667::
1659::
1651::
1641::
1614:.
1600::
1592::
1565:.
1553::
1545::
1535::
1512:.
1490::
1482::
1476:5
1442:–
1309:H
1307:N
1277:α
1177:H
1167:i
1162:i
1128:3
1122:i
1117:i
1107:i
1102:i
999:.
984:.
971:S
969:2
967:H
962:3
956:O
954:2
952:H
947:3
896:F
888:H
876:F
868:H
856:F
848:H
563:H
520:2
503:2
501:H
486:2
484:H
469:2
467:H
374:2
369:3
366:+
332:2
329:−
309:2
306:−
216:.
20:)
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.