640:
342:-regulatory modules are called billboards. Their transcriptional output is the summation effect of the bound transcription factors. Enhancers affect the probability of a gene being activated, but have little or no effect on rate. The Binary response model acts like an on/off switch for transcription. This model will increase or decrease the amount of cells that transcribe a gene, but it does not affect the rate of transcription. Rheostatic response model describes cis-regulatory modules as regulators of the initiation rate of transcription of its associated gene.
154:
3805:
289:– in this design two different regulatory factors are necessary to make sure that a positive output results. "Toggle Switches" – This design occurs when the signal ligand is absent while the transcription factor is present; this transcription factor ends up acting as a dominant repressor. However, once the signal ligand is present the transcription factor's role as repressor is eliminated and transcription can occur.
3817:
632:-regulatory modules in a genomic sequence have been difficult to identify. Problems in identification arise because often scientists find themselves with a small set of known transcription factors, so it makes it harder to identify statistically significant clusters of transcription factor binding sites. Additionally, high costs limit the use of large whole genome
532:-regulatory modules can regulate their target genes over large distances. Several models have been proposed to describe the way that these modules may communicate with their target gene promoter. These include the DNA scanning model, the DNA sequence looping model and the facilitated tracking model. In the DNA scanning model, the transcription factor and
383:. It has been found that a single gene can contain multiple promoter sites. In order to initiate transcription of the downstream gene, a host of DNA-binding proteins called transcription factors (TFs) must bind sequentially to this region. Only once this region has been bound with the appropriate set of TFs, and in the proper order, can
595:
user-friendly since it allows automatic retrieval of sequences and several visualizations and links to third-party tools in order to help users to find those instances that are more likely to be true regulatory sites. INSECT 2.0 algorithm was previously published and the algorithm and theory behind it explained in
624:
is applied to the data set to identify possible combinations of transcription factors, which have binding sites that are close to the promoter of the gene set of interest. The possible cis-regulatory modules are then statistically analyzed and the significant combinations are graphically represented
280:
Within the assumption of the
Boolean logic, principles guiding the operation of these modules includes the design of the module which determines the regulatory function. In relation to development, these modules can generate both positive and negative outputs. The output of each module is a product
267:
Additionally, the regulation of chromatin structure and nuclear organization also play a role in determining and controlling the function of cis-regulatory modules. Thus gene-regulation functions (GRF) provide a unique characteristic of a cis-regulatory module (CRM), relating the concentrations of
594:
is a web server that allows to search Cis-regulatory modules in a genome-wide manner. The program relies on the definition of strict restrictions among the
Transcription Factor Binding Sites (TFBSs) that compose the module in order to decrease the false positives rate. INSECT is designed to be
315:-regulatory module regulated by two transcription factors, experimentally determined gene-regulation functions can not be described by the 16 possible Boolean functions of two variables. Non-Boolean extensions of the gene-regulatory logic have been proposed to correct for this issue.
509:-regulatory modules can provide enough information to generate spatial and temporal patterns of gene expression. During development each domain, where each domain represents a different spatial regions of the embryo, of gene expression will be under the control of different
264:, which come into play only during specific situations during development. These inputs can come from different time points, can represent different signal ligands, or can come from different domains or lineages of cells. However, a lot still remains unknown.
215:-regulatory modules as a DNA sequence with transcription factor binding sites which are clustered into modular structures, including -but not limited to- locus control regions, promoters, enhancers, silencers, boundary control elements and other modulators.
405:, or even relatively far away from the gene they regulate. Multiple enhancers can act in a coordinated fashion to regulate transcription of one gene. A number of genome-wide sequencing projects have revealed that enhancers are often transcribed to
608:
use an algorithm that combines site predictions and tissue-specific expression data for transcription factors and target genes of interest. This model also uses regression trees to depict the relationship between the identified
165:
of an organism contains anywhere from a few hundred to thousands of different genes, all encoding a singular product or more. For numerous reasons, including organizational maintenance, energy conservation, and generating
1529:
Wrzodek, Clemens; Schröder, Adrian; Dräger, Andreas; Wanke, Dierk; Berendzen, Kenneth W.; Kronfeld, Marcel; Harter, Klaus; Zell, Andreas (2010). "ModuleMaster: A new tool to decipher transcriptional regulatory networks".
138:-regulatory modules carry out their function by integrating the active transcription factors and the associated co-factors at a specific time and place in the cell where this information is read and an output is given.
552:-regulatory module complex causes the looping of the DNA sequence slowly towards the target promoter and forms a stable looped configuration. The facilitated tracking model combines parts of the two previous models.
729:
To summarize, cis-regulatory elements are present on the same molecule of DNA as the gene they regulate whereas trans-regulatory elements can regulate genes distant from the gene from which they were transcribed.
296:-regulatory module lead to an output of zero. Additionally, besides influence from the different logic operations, the output of a "cis"-regulatory module will also be influenced by prior events. 4)
2327:
Prud'homme B, Gompel N, Rokas A, Kassner VA, Williams TM, Yeh SD, True JR, Carroll SB (April 2006). "Repeated morphological evolution through cis-regulatory changes in a pleiotropic gene".
268:
transcription factors (input) to the promoter activities (output). The challenge is to predict GRFs. This challenge still remains unsolved. In general, gene-regulation functions do not use
676:, which, in turn, prevents transcription of the adjacent genes on the same DNA molecule. The lac operator is, thus, considered to "act in cis" on the regulation of the nearby genes. The
616:
CRÈME examine clusters of target sites for transcription factors of interest. This program uses a database of confirmed transcription factor binding sites that were annotated across the
3821:
482:
arising within a CRE can generate expression variance by changing the way TFs bind. Tighter or looser binding of regulatory proteins will lead to up- or down-regulated transcription.
338:. The architecture and the arrangement of the transcription factor binding sites are critical because disruption of the arrangement could cancel out the function. Functional flexible
2282:
Gompel N, Prud'homme B, Wittkopp PJ, Kassner VA, Carroll SB (February 2005). "Chance caught on the wing: cis-regulatory evolution and the origin of pigment patterns in
Drosophila".
3048:
252:
serve as inputs, and the output of the module is the command given to the transcription machinery, which in turn determines the rate of gene transcription or whether it is
602:
to identify statistically significant clusters of transcription factor combinations. It also uses a second related genome to improve the prediction accuracy of the model.
540:-regulatory module and then continues to move along the DNA sequence until it finds the target gene promoter. In the looping model, the transcription factor binds to the
326:-regulatory modules can be characterized by the information processing that they encode and the organization of their transcription factor binding sites. Additionally,
695:
are diffusible factors, usually proteins, that may modify the expression of genes distant from the gene that was originally transcribed to create them. For example, a
356:
Promoters are CREs consisting of relatively short sequences of DNA which include the site where transcription is initiated and the region approximately 35 bp
330:-regulatory modules are also characterized by the way they affect the probability, proportion, and rate of transcription. Highly cooperative and coordinated
174:
is at the transcriptional level. CREs function to control transcription by acting nearby or within a gene. The most well characterized types of CREs are
260:. There are two types of transcription factor inputs: those that determine when the target gene is to be expressed and those that serve as functional
442:
of messenger RNA, that binds proteins which suppress translation of that mRNA molecule, but this usage is distinct from its use in describing a CRE.
401:
Enhancers are CREs that influence (enhance) the transcription of genes on the same molecule of DNA and can be found upstream, downstream, within the
207:
The original definition presented cis-regulatory modules as enhancers of cis-acting DNA, which increased the rate of transcription from a linked
2438:
300:-regulatory modules must interact with other regulatory elements. For the most part, even with the presence of functional overlap between
3383:
1351:
Wittkopp PJ, Kalay G (December 2011). "Cis-regulatory elements: molecular mechanisms and evolutionary processes underlying divergence".
2014:"Molecular control of vertebrate iron metabolism: mRNA-based regulatory circuits operated by iron, nitric oxide, and oxidative stress"
2996:
311:, detailed studies show that in general the logic of gene regulation is not Boolean. This means, for example, that in the case of a
292:
Other
Boolean logic operations can occur as well, such as sequence specific transcriptional repressors, which when they bind to the
564:
algorithms for predicting them. Most algorithms try to search for significant combinations of transcription factor binding sites (
2965:
3164:
2416:
1332:
204:-regulatory modules are non-random clusters at their specified target site that contain transcription factor binding sites.
3408:
2930:
978:
81:
CREs are found in the vicinity of the genes that they regulate. CREs typically regulate gene transcription by binding to
1240:
Jeziorska DM, Jordan KW, Vance KW (2009). "A systems biology approach to understanding cis-regulatory module function".
170:
variance, it is important that genes are only expressed when they are needed. The most efficient way for an organism to
3727:
3430:
3028:
2775:
2503:
1963:
Mokrejs M, Vopálenský V, Kolenaty O, Masek T, Feketová Z, Sekyrová P, Skaloudová B, Kríz V, Pospísek M (January 2006).
1879:
Chung BY, Firth AE, Atkins JF (March 2010). "Frameshifting in alphaviruses: a diversity of 3' stimulatory structures".
75:
126:, which refers to effects on genes not located on the same strand or farther away, such as transcription factors. One
3719:
3659:
3136:
2475:
572:
sequences of co-expressed genes. More advanced methods combine the search for significant motifs with correlation in
2732:
2518:
2513:
3781:
3664:
3376:
3290:
3234:
439:
3229:
2909:
3786:
3776:
3763:
3317:
3248:
2989:
2760:
1002:
357:
171:
109:
3403:
3053:
966:
795:
548:
of the DNA sequence and allows for the interaction with the target gene promoter. The transcription factor-
380:
196:. Regulatory elements are binding sites for transcription factors, which are involved in gene regulation.
2727:
2443:
59:
2202:"A novel RNA structural motif in the selenocysteine insertion element of eukaryotic selenoprotein mRNAs"
3809:
3369:
3268:
2919:
2785:
2617:
85:. A single transcription factor may bind to many CREs, and hence control the expression of many genes (
2612:
2157:
Kortmann J, Narberhaus F (March 2012). "Bacterial RNA thermometers: molecular zippers and switches".
591:
470:
among organisms; yet different organisms display marked phenotypic diversity. It has been found that
438:, thereby preventing transcription of a gene. The term "silencer" can also refer to a region in the
3312:
3100:
3013:
2982:
2447:
761:
692:
639:
142:
121:
51:
120:
because they are typically located on the same DNA strand as the genes they control as opposed to
3858:
3848:
3448:
3273:
3090:
3075:
2853:
2790:
2765:
2648:
2568:
1012:
533:
494:
3203:
3095:
2737:
2722:
2691:
2592:
518:
471:
3607:
3602:
3544:
3458:
3193:
3178:
3058:
2949:
2843:
2770:
2669:
2607:
2602:
2533:
2468:
1322:
1191:
839:
819:
577:
498:
285:– this design indicates that in an output will be given when either input is given , and the
1172:
157:
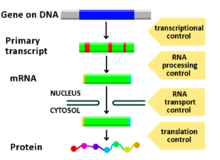
Diagram showing at which stages in the DNA-mRNA-protein pathway expression can be controlled
3707:
3564:
3413:
3300:
3198:
3116:
2858:
2701:
2696:
2578:
2573:
2560:
2336:
2291:
2115:
2025:
1282:"The RNA polymerase II core promoter: a key component in the regulation of gene expression"
1127:
696:
644:
581:
431:
245:
226:
105:
82:
2434:
Gene
Regulation Info – manually curated lists of resources, reviews, community discussions
718:
There are cis-regulatory and trans-regulatory elements. Cis-regulatory elements are often
130:-regulatory element can regulate several genes, and conversely, one gene can have several
8:
3576:
3571:
3549:
3495:
3126:
2954:
1027:
1007:
665:
648:
427:
422:
413:(eRNA), whose changes in levels frequently correlate with those of the target gene mRNA.
406:
396:
351:
253:
193:
179:
175:
97:
means "on this side", i.e. on the same molecule of DNA as the gene(s) to be transcribed.
2340:
2295:
2119:
2084:
2029:
1131:
304:-regulatory modules of a gene, the modules' inputs and outputs tend to not be the same.
3261:
3144:
2643:
2390:
2373:
2360:
2315:
2270:
2218:
2201:
2182:
2139:
1989:
1964:
1940:
1915:
1856:
1831:
1758:
1731:
1704:
1679:
1471:
1446:
1422:
1395:
1376:
1148:
1115:
1022:
961:
677:
569:
467:
208:
1807:
1782:
474:
occurring within non-coding sequences have a profound effect on phenotype by altering
272:, although in some cases the approximation of the Boolean logic is still very useful.
3748:
3392:
2904:
2684:
2679:
2588:
2412:
2395:
2352:
2307:
2262:
2223:
2174:
2131:
2088:
2053:
2048:
2013:
1994:
1945:
1896:
1861:
1812:
1763:
1709:
1657:
1606:
1555:
1547:
1543:
1511:
1507:
1476:
1427:
1368:
1328:
1303:
1257:
1195:
1153:
466:
CREs have an important evolutionary role. The coding regions of genes are often well
376:
200:-regulatory modules perform a large amount of developmental information processing.
141:
CREs are often but not always upstream of the transcription site. CREs contrast with
3345:
2888:
2319:
2186:
2143:
1965:"IRESite: the database of experimentally verified IRES structures (www.iresite.org)"
1642:
1625:
1591:
1574:
1494:
Aerts, S.; et al. (2003). "Computational detection of cis-regulatory modules".
1380:
2914:
2883:
2878:
2833:
2461:
2385:
2364:
2344:
2299:
2254:
2245:
Wray GA (March 2007). "The evolutionary significance of cis-regulatory mutations".
2213:
2166:
2123:
2080:
2071:
Platt T (1986). "Transcription termination and the regulation of gene expression".
2043:
2033:
1984:
1976:
1935:
1931:
1927:
1888:
1851:
1843:
1802:
1794:
1753:
1743:
1699:
1691:
1647:
1637:
1596:
1586:
1539:
1503:
1466:
1458:
1417:
1407:
1360:
1293:
1249:
1187:
1143:
1135:
605:
565:
372:
2274:
1830:
Bekaert M, Firth AE, Zhang Y, Gladyshev VN, Atkins JF, Baranov PV (January 2010).
1412:
3712:
3438:
3322:
3159:
3005:
2924:
2828:
2638:
1253:
989:
983:
881:
573:
475:
47:
1732:"Comparing sequences without using alignments: application to HIV/SIV subtyping"
3743:
3694:
3654:
3149:
3121:
2453:
2018:
Proceedings of the
National Academy of Sciences of the United States of America
909:
561:
384:
257:
234:
183:
43:
1892:
1139:
182:. Both of these sequence elements are structural regions of DNA that serve as
18:
Region of non-coding DNA that regulates the transcription of neighboring genes
3837:
3753:
3644:
3559:
3350:
3037:
2892:
2806:
2674:
2508:
2485:
1551:
1116:"Predicting Gene-Regulation Functions: Lessons from Temperate Bacteriophages"
900:
769:
719:
704:
673:
269:
71:
63:
2409:
From DNA to
Diversity: Molecular Genetics and the Evolution of Animal Design
2127:
2038:
1575:"INSECT 2.0: a web-server for genome-wide cis-regulatory modules prediction"
1094:
The
Regulatory Genome: Gene Regulatory Networks in Development and Evolution
3853:
3843:
3689:
3597:
3592:
3224:
3154:
3023:
2717:
2538:
2528:
2399:
2356:
2311:
2266:
2178:
2135:
1998:
1949:
1900:
1865:
1816:
1767:
1713:
1661:
1610:
1559:
1515:
1480:
1431:
1372:
1307:
1261:
1199:
1157:
723:
700:
633:
617:
599:
410:
335:
2227:
2092:
2057:
1748:
613:-regulatory module and the possible binding set of transcription factors.
580:
and target genes. Both methods have been implemented, for example, in the
153:
3771:
3500:
3327:
3256:
2653:
2633:
1980:
1847:
1798:
1695:
941:
281:
of the various operations performed on it. Common operations include the
249:
2348:
2303:
2170:
1298:
1281:
513:-regulatory modules. The design of regulatory modules help in producing
450:
Operators are CREs in prokaryotes and some eukaryotes that exist within
3527:
3522:
3517:
3285:
2897:
2597:
2550:
1680:"Stubb: a program for discovery and analysis of cis-regulatory modules"
1652:
1601:
1462:
872:
858:
669:
86:
3554:
2848:
2816:
2583:
2489:
1037:
956:
830:
810:
806:
786:
782:
621:
455:
435:
361:
222:
167:
3361:
2258:
1396:"Transcriptional enhancers: Transcription, function and flexibility"
1364:
3617:
3612:
3505:
3305:
3295:
3219:
2811:
2780:
2281:
951:
936:
891:
868:
849:
778:
652:
514:
485:
479:
365:
286:
2974:
1573:
Parra RG, Rohr CO, Koile D, Perez-Castro C, Yankilevich P (2015).
584:. Other programs created for the identification and prediction of
3723:
3698:
3673:
3669:
3649:
3462:
3041:
2543:
2374:"Evolutionary developmental biology and the problem of variation"
1626:"INSECT: IN-silico SEarch for Co-occurring Transcription factors"
946:
914:
774:
681:
282:
67:
2326:
1623:
3534:
3512:
2523:
1017:
451:
402:
162:
104:, usually 100–1000 DNA base pairs in length, where a number of
2433:
1962:
555:
3736:
3063:
1832:"Recode-2: new design, new search tools, and many more genes"
369:
90:
1783:"CREME: Cis-Regulatory Module Explorer for the human genome"
1780:
364:, promoters usually have the following four components: the
116:
and regulate their transcription rates. They are labeled as
3539:
2821:
1032:
560:
Besides experimentally determining CRMs, there are various
192:-regulatory modules are one of several types of functional
113:
55:
1829:
1572:
1528:
846:
Regulates transcription of associated genes and/or operons
3452:
3442:
3033:
1624:
Rohr CO, Parra RG, Yankilevich P, Perez-Castro C (2013).
1170:
685:
229:, which work indirectly by interacting with other nearby
101:
2406:
2199:
1916:"Frameshifting RNA pseudoknots: structure and mechanism"
1387:
307:
While the assumption of
Boolean logic is important for
221:-regulatory modules can be divided into three classes;
1173:"Gene regulation: gene control network in development"
803:
Initiates translation in the middle of a messenger RNA
2378:
Evolution; International
Journal of Organic Evolution
2200:
Walczak R, Westhof E, Carbon P, Krol A (April 1996).
1327:. Springer Science & Business Media. p. 78.
1781:
Sharan R, Ben-Hur A, Loots GG, Ovcharenko I (2004).
1239:
1394:Melamed P, Yosefzun Y, et al. (2 March 2016).
1346:
1344:
2156:
703:might itself have been transcribed from a gene on
2106:Breaker RR (March 2008). "Complex riboswitches".
1447:"Building Developmental Gene Regulatory Networks"
908:Directs the cell to translate UGA stop-codons as
827:Regulates the expression of iron associated genes
211:. However, this definition has changed to define
3835:
2483:
1878:
1677:
1341:
1393:
275:
2407:Weatherbee SD, Carroll SB, Grenier JK (2004).
1913:
1729:
1279:
501:, whose function is dependent on the multiple
3377:
2990:
2469:
1350:
489:-regulatory module in gene regulatory network
225:, which regulate gene expression positively;
2099:
1956:
1907:
1872:
1823:
1673:
1671:
1275:
1273:
1271:
1235:
1233:
1231:
1229:
2064:
2005:
1725:
1723:
1522:
1227:
1225:
1223:
1221:
1219:
1217:
1215:
1213:
1211:
1209:
1091:
556:Identification and computational prediction
3384:
3370:
3145:Precursor mRNA (pre-mRNA / hnRNA)
2997:
2983:
2476:
2462:
2011:
1444:
1109:
1107:
1105:
1103:
643:Binding sites of gene regulatory factors.
544:-regulatory module, which then causes the
2446:at the U.S. National Library of Medicine
2389:
2217:
2047:
2037:
1988:
1939:
1855:
1806:
1757:
1747:
1703:
1668:
1651:
1641:
1600:
1590:
1566:
1493:
1487:
1470:
1421:
1411:
1297:
1268:
1171:Ben-Tabou de-Leon S, Davidson EH (2007).
1147:
1113:
1914:Giedroc DP, Cornish PV (February 2009).
1720:
1617:
1206:
1192:10.1146/annurev.biophys.35.040405.102002
638:
152:
2411:. Cambridge, MA: Blackwell Publishers.
2105:
1314:
1280:Butler JE, Kadonaga JT (October 2002).
1100:
1087:
1085:
1083:
1081:
1079:
1077:
1075:
145:. TREs code for transcription factors.
3836:
2966:Index of evolutionary biology articles
1774:
1164:
1073:
1071:
1069:
1067:
1065:
1063:
1061:
1059:
1057:
1055:
454:, where they can bind proteins called
387:bind and begin transcribing the gene.
334:-regulatory modules are classified as
3391:
3365:
3165:Histone acetylation and deacetylation
2978:
2457:
2371:
2070:
1438:
768:Regulates alternative frame use with
3230:Ribosome-nascent chain complex (RNC)
2244:
1320:
672:. This DNA sequence is bound by the
461:
3816:
3004:
2085:10.1146/annurev.bi.55.070186.002011
1678:Sinha S, Liang Y, Siggia E (2006).
1052:
979:List of cis-regulatory RNA elements
711:is constructed from the Latin root
505:-regulatory modules. The layout of
497:depends on the architecture of the
237:that turn off expression of genes.
13:
2776:Evolutionary developmental biology
2391:10.1111/j.0014-3820.2000.tb00544.x
2237:
2012:Hentze MW, Kühn LC (August 1996).
733:
360:from the initiation site (bp). In
76:evolutionary developmental biology
14:
3870:
3660:Post-transcriptional modification
2427:
901:Selenocysteine insertion sequence
524:
318:
244:-regulatory modules is such that
3815:
3804:
3803:
2733:Evolution of sexual reproduction
1544:10.1016/j.biosystems.2009.09.005
432:transcription regulation factors
143:trans-regulatory elements (TREs)
3665:Post-translational modification
3235:Post-translational modification
2193:
2150:
1793:(Web Server issue): W253–W256.
1690:(Web Server issue): W555–W559.
1538:(1). Ireland: Elsevier: 79–81.
1324:Introduction to Systems Biology
58:. CREs are vital components of
2504:Genotype–phenotype distinction
1932:10.1016/j.virusres.2008.06.008
1508:10.1093/bioinformatics/btg1052
1180:Annu Rev Biophys Biomol Struct
521:, and cross-regulatory loops.
1:
3787:Post-translational regulation
2761:Regulation of gene expression
2444:Regulation of Gene Expression
2073:Annual Review of Biochemistry
1730:Chen X, Blanchette M (2007).
1643:10.1093/bioinformatics/btt506
1592:10.1093/bioinformatics/btv726
1413:10.1080/21541264.2015.1128517
1046:
1003:Regulation of gene expression
715:, which means "across from".
680:itself does not code for any
588:-regulatory modules include:
3735:High-throughput technique ("
2931:Endless Forms Most Beautiful
2711:Evolution of genetic systems
2519:Gene–environment correlation
2514:Gene–environment interaction
2159:Nature Reviews. Microbiology
1975:(Database issue): D125–130.
1881:Journal of Molecular Biology
1254:10.1016/j.semcdb.2009.07.007
967:Upstream activation sequence
796:Internal ribosome entry site
445:
416:
390:
276:The Boolean logic assumption
7:
3613:Functional biology/medicine
2910:Christiane Nüsslein-Volhard
924:
664:An example of a cis-acting
659:
345:
148:
60:genetic regulatory networks
10:
3875:
2786:Hedgehog signaling pathway
2663:Developmental architecture
1842:(Database issue): D69–74.
1445:Li E, Davidson EH (2009).
1096:. Elsevier. pp. 1–86.
420:
394:
349:
184:transcriptional regulators
3799:
3762:
3682:
3637:
3630:
3585:
3472:
3429:
3422:
3399:
3338:
3247:
3212:
3186:
3177:
3135:
3109:
3083:
3074:
3012:
2963:
2942:
2871:
2799:
2753:
2746:
2710:
2662:
2626:
2613:Transgressive segregation
2559:
2496:
1893:10.1016/j.jmb.2010.01.044
1140:10.1016/j.bpj.2009.11.046
699:that regulates a gene on
693:trans-regulatory elements
458:to affect transcription.
233:-regulatory modules; and
3296:sequestration (P-bodies)
2448:Medical Subject Headings
2372:Stern DL (August 2000).
250:epigenetic modifications
172:regulate gene expression
62:, which in turn control
3274:Gene regulatory network
2791:Notch signaling pathway
2766:Gene regulatory network
2649:Dual inheritance theory
2247:Nature Reviews Genetics
2128:10.1126/science.1152621
2039:10.1073/pnas.93.16.8175
1353:Nature Reviews Genetics
1286:Genes & Development
1013:Gene regulatory network
668:is the operator in the
495:gene regulatory network
430:are CREs that can bind
70:, and other aspects of
3279:cis-regulatory element
2839:cis-regulatory element
2747:Control of development
2627:Non-genetic influences
2593:evolutionary landscape
1969:Nucleic Acids Research
1836:Nucleic Acids Research
1321:Choi S (17 May 2008).
656:
440:3' untranslated region
358:upstream or downstream
158:
100:CRMs are stretches of
3608:Developmental biology
3603:Computational biology
2950:Nature versus nurture
2854:Cell surface receptor
2771:Evo-devo gene toolkit
2670:Developmental biology
2608:Polygenic inheritance
2534:Quantitative genetics
1749:10.1186/1471-2105-8-1
1242:Semin. Cell Dev. Biol
820:Iron response element
645:Transcription factors
642:
578:transcription factors
381:core promoter element
379:, and the downstream
246:transcription factors
156:
134:-regulatory modules.
106:transcription factors
83:transcription factors
72:embryonic development
66:, the development of
3782:Post-transcriptional
3301:alternative splicing
3291:Post-transcriptional
3117:Transcription factor
2859:Transcription factor
2574:Genetic assimilation
2561:Genetic architecture
1092:Davidson EH (2006).
697:transcription factor
649:RNA-binding proteins
536:complex form at the
25:-regulatory elements
3577:Histone methylation
3225:Transfer RNA (tRNA)
2955:Morphogenetic field
2872:Influential figures
2349:10.1038/nature04597
2341:2006Natur.440.1050P
2335:(7087): 1050–1053.
2304:10.1038/nature03235
2296:2005Natur.433..481G
2171:10.1038/nrmicro2730
2120:2008Sci...319.1795B
2114:(5871): 1795–1797.
2030:1996PNAS...93.8175H
1502:(Suppl 2): ii5–14.
1299:10.1101/gad.1026202
1132:2010BpJ....98.1247T
1120:Biophysical Journal
1028:Trans-acting factor
1008:Cis-trans isomerism
740:
666:regulatory sequence
423:Silencer (genetics)
407:long non-coding RNA
397:Enhancer (genetics)
352:Promoter (genetics)
194:regulatory elements
110:regulate expression
36:-regulatory modules
3339:Influential people
3318:Post-translational
3137:Post-transcription
2644:Genomic imprinting
2439:Cellular Darwinism
1981:10.1093/nar/gkj081
1848:10.1093/nar/gkp788
1799:10.1093/nar/gkh385
1736:BMC Bioinformatics
1696:10.1093/nar/gkl224
1463:10.1002/bdrc.20152
1114:Teif V.B. (2010).
962:Operator (biology)
762:Frameshift element
738:
657:
598:Stubb uses hidden
493:The function of a
434:(proteins) called
159:
3831:
3830:
3810:Molecular biology
3795:
3794:
3749:Mass spectrometry
3626:
3625:
3393:Molecular biology
3359:
3358:
3243:
3242:
3173:
3172:
3049:Special transfers
2972:
2971:
2905:Eric F. Wieschaus
2867:
2866:
2685:Pattern formation
2589:Fitness landscape
2418:978-1-4051-1950-4
2290:(7025): 481–487.
2024:(16): 8175–8182.
1787:Nucleic Acids Res
1684:Nucleic Acids Res
1451:Birth Defects Res
1334:978-1-59745-531-2
1292:(20): 2583–2592.
922:
921:
606:Bayesian Networks
576:datasets between
566:DNA binding sites
462:Evolutionary role
42:) are regions of
3866:
3819:
3818:
3807:
3806:
3740:
3635:
3634:
3488:
3483:
3427:
3426:
3386:
3379:
3372:
3363:
3362:
3184:
3183:
3081:
3080:
2999:
2992:
2985:
2976:
2975:
2915:William McGinnis
2884:Richard Lewontin
2879:C. H. Waddington
2751:
2750:
2728:Neutral networks
2478:
2471:
2464:
2455:
2454:
2422:
2403:
2393:
2384:(4): 1079–1091.
2368:
2323:
2278:
2232:
2231:
2221:
2197:
2191:
2190:
2154:
2148:
2147:
2103:
2097:
2096:
2068:
2062:
2061:
2051:
2041:
2009:
2003:
2002:
1992:
1960:
1954:
1953:
1943:
1911:
1905:
1904:
1876:
1870:
1869:
1859:
1827:
1821:
1820:
1810:
1778:
1772:
1771:
1761:
1751:
1727:
1718:
1717:
1707:
1675:
1666:
1665:
1655:
1645:
1621:
1615:
1614:
1604:
1594:
1570:
1564:
1563:
1526:
1520:
1519:
1491:
1485:
1484:
1474:
1442:
1436:
1435:
1425:
1415:
1391:
1385:
1384:
1348:
1339:
1338:
1318:
1312:
1311:
1301:
1277:
1266:
1265:
1237:
1204:
1203:
1177:
1168:
1162:
1161:
1151:
1111:
1098:
1097:
1089:
741:
737:
722:for one or more
709:trans-regulatory
373:recognition site
3874:
3873:
3869:
3868:
3867:
3865:
3864:
3863:
3834:
3833:
3832:
3827:
3791:
3764:Gene regulation
3758:
3734:
3695:Model organisms
3678:
3655:Cell signalling
3622:
3581:
3486:
3481:
3468:
3439:DNA replication
3418:
3395:
3390:
3360:
3355:
3334:
3269:Transcriptional
3239:
3208:
3169:
3160:Polyadenylation
3131:
3105:
3070:
3064:Protein→Protein
3015:
3008:
3006:Gene expression
3003:
2973:
2968:
2959:
2938:
2925:Sean B. Carroll
2863:
2795:
2742:
2706:
2658:
2639:Maternal effect
2622:
2555:
2492:
2482:
2430:
2425:
2419:
2259:10.1038/nrg2063
2240:
2238:Further reading
2235:
2198:
2194:
2155:
2151:
2104:
2100:
2069:
2065:
2010:
2006:
1961:
1957:
1912:
1908:
1877:
1873:
1828:
1824:
1779:
1775:
1728:
1721:
1676:
1669:
1622:
1618:
1571:
1567:
1527:
1523:
1492:
1488:
1443:
1439:
1392:
1388:
1365:10.1038/nrg3095
1349:
1342:
1335:
1319:
1315:
1278:
1269:
1238:
1207:
1175:
1169:
1165:
1112:
1101:
1090:
1053:
1049:
1044:
990:AU-rich element
984:Polyadenylation
927:
910:selenocysteines
888:Gene regulation
882:RNA thermometer
865:Gene regulation
736:
734:Examples in RNA
662:
574:gene expression
558:
527:
491:
476:gene expression
464:
448:
425:
419:
399:
393:
354:
348:
321:
309:systems biology
278:
151:
54:of neighboring
19:
12:
11:
5:
3872:
3862:
3861:
3859:Non-coding DNA
3856:
3851:
3849:Non-coding RNA
3846:
3829:
3828:
3826:
3825:
3813:
3800:
3797:
3796:
3793:
3792:
3790:
3789:
3784:
3779:
3774:
3768:
3766:
3760:
3759:
3757:
3756:
3751:
3746:
3744:DNA microarray
3741:
3731:
3730:
3717:
3716:
3715:
3710:
3702:
3692:
3686:
3684:
3680:
3679:
3677:
3676:
3667:
3662:
3657:
3652:
3647:
3641:
3639:
3632:
3628:
3627:
3624:
3623:
3621:
3620:
3615:
3610:
3605:
3600:
3595:
3589:
3587:
3583:
3582:
3580:
3579:
3574:
3569:
3568:
3567:
3562:
3552:
3547:
3542:
3537:
3532:
3531:
3530:
3525:
3520:
3510:
3509:
3508:
3503:
3492:
3491:
3490:
3489:
3484:
3476:
3474:
3470:
3469:
3467:
3466:
3456:
3446:
3435:
3433:
3424:
3420:
3419:
3417:
3416:
3411:
3406:
3400:
3397:
3396:
3389:
3388:
3381:
3374:
3366:
3357:
3356:
3354:
3353:
3348:
3346:François Jacob
3342:
3340:
3336:
3335:
3333:
3332:
3331:
3330:
3325:
3315:
3310:
3309:
3308:
3303:
3298:
3288:
3283:
3282:
3281:
3276:
3266:
3265:
3264:
3253:
3251:
3245:
3244:
3241:
3240:
3238:
3237:
3232:
3227:
3222:
3216:
3214:
3210:
3209:
3207:
3206:
3201:
3196:
3190:
3188:
3181:
3175:
3174:
3171:
3170:
3168:
3167:
3162:
3157:
3152:
3147:
3141:
3139:
3133:
3132:
3130:
3129:
3124:
3122:RNA polymerase
3119:
3113:
3111:
3107:
3106:
3104:
3103:
3098:
3093:
3087:
3085:
3078:
3072:
3071:
3069:
3068:
3067:
3066:
3061:
3056:
3046:
3045:
3044:
3026:
3020:
3018:
3010:
3009:
3002:
3001:
2994:
2987:
2979:
2970:
2969:
2964:
2961:
2960:
2958:
2957:
2952:
2946:
2944:
2940:
2939:
2937:
2936:
2935:
2934:
2922:
2917:
2912:
2907:
2902:
2901:
2900:
2889:François Jacob
2886:
2881:
2875:
2873:
2869:
2868:
2865:
2864:
2862:
2861:
2856:
2851:
2846:
2841:
2836:
2831:
2826:
2825:
2824:
2814:
2809:
2803:
2801:
2797:
2796:
2794:
2793:
2788:
2783:
2778:
2773:
2768:
2763:
2757:
2755:
2748:
2744:
2743:
2741:
2740:
2735:
2730:
2725:
2720:
2714:
2712:
2708:
2707:
2705:
2704:
2699:
2694:
2689:
2688:
2687:
2682:
2672:
2666:
2664:
2660:
2659:
2657:
2656:
2651:
2646:
2641:
2636:
2630:
2628:
2624:
2623:
2621:
2620:
2618:Sequence space
2615:
2610:
2605:
2600:
2595:
2586:
2581:
2576:
2571:
2565:
2563:
2557:
2556:
2554:
2553:
2548:
2547:
2546:
2536:
2531:
2526:
2521:
2516:
2511:
2506:
2500:
2498:
2494:
2493:
2481:
2480:
2473:
2466:
2458:
2452:
2451:
2441:
2436:
2429:
2428:External links
2426:
2424:
2423:
2417:
2404:
2369:
2324:
2279:
2253:(3): 206–216.
2241:
2239:
2236:
2234:
2233:
2212:(4): 367–379.
2192:
2165:(4): 255–265.
2149:
2098:
2063:
2004:
1955:
1926:(2): 193–208.
1920:Virus Research
1906:
1887:(2): 448–456.
1871:
1822:
1773:
1719:
1667:
1636:(22): 2852–8.
1630:Bioinformatics
1616:
1585:(8): 1229–31.
1579:Bioinformatics
1565:
1521:
1496:Bioinformatics
1486:
1457:(2): 123–130.
1437:
1386:
1340:
1333:
1313:
1267:
1248:(7): 856–862.
1205:
1163:
1126:(7): 1247–56.
1099:
1050:
1048:
1045:
1043:
1042:
1041:
1040:
1035:
1030:
1025:
1020:
1015:
1010:
1005:
995:
994:
993:
987:
981:
971:
970:
969:
964:
959:
954:
949:
944:
939:
928:
926:
923:
920:
919:
917:
912:
906:
903:
897:
896:
894:
889:
886:
884:
878:
877:
875:
866:
863:
861:
855:
854:
852:
847:
844:
842:
840:Leader peptide
836:
835:
833:
828:
825:
822:
816:
815:
813:
804:
801:
798:
792:
791:
789:
772:
770:messenger RNAs
766:
764:
758:
757:
754:
751:
748:
745:
735:
732:
661:
658:
562:bioinformatics
557:
554:
526:
525:Mode of action
523:
490:
484:
463:
460:
447:
444:
421:Main article:
418:
415:
395:Main article:
392:
389:
385:RNA polymerase
350:Main article:
347:
344:
320:
319:Classification
317:
277:
274:
240:The design of
150:
147:
44:non-coding DNA
17:
9:
6:
4:
3:
2:
3871:
3860:
3857:
3855:
3852:
3850:
3847:
3845:
3842:
3841:
3839:
3824:
3823:
3814:
3812:
3811:
3802:
3801:
3798:
3788:
3785:
3783:
3780:
3778:
3775:
3773:
3770:
3769:
3767:
3765:
3761:
3755:
3754:Lab-on-a-chip
3752:
3750:
3747:
3745:
3742:
3738:
3733:
3732:
3729:
3728:Radioactivity
3725:
3721:
3718:
3714:
3711:
3709:
3706:
3705:
3703:
3700:
3696:
3693:
3691:
3688:
3687:
3685:
3681:
3675:
3671:
3668:
3666:
3663:
3661:
3658:
3656:
3653:
3651:
3648:
3646:
3645:Cultured meat
3643:
3642:
3640:
3636:
3633:
3629:
3619:
3616:
3614:
3611:
3609:
3606:
3604:
3601:
3599:
3596:
3594:
3591:
3590:
3588:
3584:
3578:
3575:
3573:
3570:
3566:
3565:trp repressor
3563:
3561:
3560:lac repressor
3558:
3557:
3556:
3553:
3551:
3548:
3546:
3543:
3541:
3538:
3536:
3533:
3529:
3526:
3524:
3521:
3519:
3516:
3515:
3514:
3511:
3507:
3504:
3502:
3499:
3498:
3497:
3494:
3493:
3485:
3480:
3479:
3478:
3477:
3475:
3471:
3464:
3460:
3457:
3454:
3450:
3449:Transcription
3447:
3444:
3440:
3437:
3436:
3434:
3432:
3431:Central dogma
3428:
3425:
3421:
3415:
3412:
3410:
3407:
3405:
3402:
3401:
3398:
3394:
3387:
3382:
3380:
3375:
3373:
3368:
3367:
3364:
3352:
3351:Jacques Monod
3349:
3347:
3344:
3343:
3341:
3337:
3329:
3326:
3324:
3321:
3320:
3319:
3316:
3314:
3313:Translational
3311:
3307:
3304:
3302:
3299:
3297:
3294:
3293:
3292:
3289:
3287:
3284:
3280:
3277:
3275:
3272:
3271:
3270:
3267:
3263:
3260:
3259:
3258:
3255:
3254:
3252:
3250:
3246:
3236:
3233:
3231:
3228:
3226:
3223:
3221:
3218:
3217:
3215:
3211:
3205:
3202:
3200:
3197:
3195:
3192:
3191:
3189:
3185:
3182:
3180:
3176:
3166:
3163:
3161:
3158:
3156:
3153:
3151:
3148:
3146:
3143:
3142:
3140:
3138:
3134:
3128:
3125:
3123:
3120:
3118:
3115:
3114:
3112:
3108:
3102:
3099:
3097:
3094:
3092:
3089:
3088:
3086:
3082:
3079:
3077:
3076:Transcription
3073:
3065:
3062:
3060:
3057:
3055:
3052:
3051:
3050:
3047:
3043:
3039:
3035:
3032:
3031:
3030:
3029:Central dogma
3027:
3025:
3022:
3021:
3019:
3017:
3011:
3007:
3000:
2995:
2993:
2988:
2986:
2981:
2980:
2977:
2967:
2962:
2956:
2953:
2951:
2948:
2947:
2945:
2941:
2933:
2932:
2928:
2927:
2926:
2923:
2921:
2918:
2916:
2913:
2911:
2908:
2906:
2903:
2899:
2896:
2895:
2894:
2893:Jacques Monod
2890:
2887:
2885:
2882:
2880:
2877:
2876:
2874:
2870:
2860:
2857:
2855:
2852:
2850:
2847:
2845:
2842:
2840:
2837:
2835:
2832:
2830:
2827:
2823:
2820:
2819:
2818:
2815:
2813:
2810:
2808:
2807:Homeotic gene
2805:
2804:
2802:
2798:
2792:
2789:
2787:
2784:
2782:
2779:
2777:
2774:
2772:
2769:
2767:
2764:
2762:
2759:
2758:
2756:
2752:
2749:
2745:
2739:
2736:
2734:
2731:
2729:
2726:
2724:
2721:
2719:
2716:
2715:
2713:
2709:
2703:
2700:
2698:
2695:
2693:
2690:
2686:
2683:
2681:
2678:
2677:
2676:
2675:Morphogenesis
2673:
2671:
2668:
2667:
2665:
2661:
2655:
2652:
2650:
2647:
2645:
2642:
2640:
2637:
2635:
2632:
2631:
2629:
2625:
2619:
2616:
2614:
2611:
2609:
2606:
2604:
2601:
2599:
2596:
2594:
2590:
2587:
2585:
2582:
2580:
2577:
2575:
2572:
2570:
2567:
2566:
2564:
2562:
2558:
2552:
2549:
2545:
2542:
2541:
2540:
2537:
2535:
2532:
2530:
2527:
2525:
2522:
2520:
2517:
2515:
2512:
2510:
2509:Reaction norm
2507:
2505:
2502:
2501:
2499:
2495:
2491:
2487:
2479:
2474:
2472:
2467:
2465:
2460:
2459:
2456:
2449:
2445:
2442:
2440:
2437:
2435:
2432:
2431:
2420:
2414:
2410:
2405:
2401:
2397:
2392:
2387:
2383:
2379:
2375:
2370:
2366:
2362:
2358:
2354:
2350:
2346:
2342:
2338:
2334:
2330:
2325:
2321:
2317:
2313:
2309:
2305:
2301:
2297:
2293:
2289:
2285:
2280:
2276:
2272:
2268:
2264:
2260:
2256:
2252:
2248:
2243:
2242:
2229:
2225:
2220:
2215:
2211:
2207:
2203:
2196:
2188:
2184:
2180:
2176:
2172:
2168:
2164:
2160:
2153:
2145:
2141:
2137:
2133:
2129:
2125:
2121:
2117:
2113:
2109:
2102:
2094:
2090:
2086:
2082:
2078:
2074:
2067:
2059:
2055:
2050:
2045:
2040:
2035:
2031:
2027:
2023:
2019:
2015:
2008:
2000:
1996:
1991:
1986:
1982:
1978:
1974:
1970:
1966:
1959:
1951:
1947:
1942:
1937:
1933:
1929:
1925:
1921:
1917:
1910:
1902:
1898:
1894:
1890:
1886:
1882:
1875:
1867:
1863:
1858:
1853:
1849:
1845:
1841:
1837:
1833:
1826:
1818:
1814:
1809:
1804:
1800:
1796:
1792:
1788:
1784:
1777:
1769:
1765:
1760:
1755:
1750:
1745:
1741:
1737:
1733:
1726:
1724:
1715:
1711:
1706:
1701:
1697:
1693:
1689:
1685:
1681:
1674:
1672:
1663:
1659:
1654:
1649:
1644:
1639:
1635:
1631:
1627:
1620:
1612:
1608:
1603:
1598:
1593:
1588:
1584:
1580:
1576:
1569:
1561:
1557:
1553:
1549:
1545:
1541:
1537:
1533:
1525:
1517:
1513:
1509:
1505:
1501:
1497:
1490:
1482:
1478:
1473:
1468:
1464:
1460:
1456:
1452:
1448:
1441:
1433:
1429:
1424:
1419:
1414:
1409:
1405:
1401:
1400:Transcription
1397:
1390:
1382:
1378:
1374:
1370:
1366:
1362:
1358:
1354:
1347:
1345:
1336:
1330:
1326:
1325:
1317:
1309:
1305:
1300:
1295:
1291:
1287:
1283:
1276:
1274:
1272:
1263:
1259:
1255:
1251:
1247:
1243:
1236:
1234:
1232:
1230:
1228:
1226:
1224:
1222:
1220:
1218:
1216:
1214:
1212:
1210:
1201:
1197:
1193:
1189:
1185:
1181:
1174:
1167:
1159:
1155:
1150:
1145:
1141:
1137:
1133:
1129:
1125:
1121:
1117:
1110:
1108:
1106:
1104:
1095:
1088:
1086:
1084:
1082:
1080:
1078:
1076:
1074:
1072:
1070:
1068:
1066:
1064:
1062:
1060:
1058:
1056:
1051:
1039:
1036:
1034:
1031:
1029:
1026:
1024:
1021:
1019:
1016:
1014:
1011:
1009:
1006:
1004:
1001:
1000:
999:
996:
991:
988:
986:signals, mRNA
985:
982:
980:
977:
976:
975:
972:
968:
965:
963:
960:
958:
955:
953:
950:
948:
945:
943:
940:
938:
935:
934:
933:
930:
929:
918:
916:
913:
911:
907:
904:
902:
899:
898:
895:
893:
890:
887:
885:
883:
880:
879:
876:
874:
870:
867:
864:
862:
860:
857:
856:
853:
851:
848:
845:
843:
841:
838:
837:
834:
832:
829:
826:
823:
821:
818:
817:
814:
812:
808:
805:
802:
799:
797:
794:
793:
790:
788:
784:
780:
776:
773:
771:
767:
765:
763:
760:
759:
755:
752:
749:
746:
743:
742:
739:RNA elements
731:
727:
725:
721:
720:binding sites
716:
714:
710:
706:
705:chromosome 11
702:
698:
694:
691:In contrast,
689:
687:
683:
679:
675:
674:lac repressor
671:
667:
654:
650:
647:binding DNA,
646:
641:
637:
635:
634:tiling arrays
631:
626:
623:
619:
614:
612:
607:
603:
601:
600:Markov models
596:
593:
589:
587:
583:
579:
575:
571:
567:
563:
553:
551:
547:
543:
539:
535:
531:
522:
520:
516:
512:
508:
504:
500:
496:
488:
483:
481:
477:
473:
472:polymorphisms
469:
459:
457:
453:
443:
441:
437:
433:
429:
424:
414:
412:
408:
404:
398:
388:
386:
382:
378:
374:
371:
367:
363:
359:
353:
343:
341:
337:
336:enhanceosomes
333:
329:
325:
316:
314:
310:
305:
303:
299:
295:
290:
288:
284:
273:
271:
270:Boolean logic
265:
263:
259:
255:
251:
247:
243:
238:
236:
232:
228:
224:
220:
216:
214:
210:
205:
203:
199:
195:
191:
187:
185:
181:
177:
173:
169:
164:
155:
146:
144:
139:
137:
133:
129:
125:
124:
119:
115:
111:
108:can bind and
107:
103:
98:
96:
92:
88:
84:
79:
77:
74:, studied in
73:
69:
65:
64:morphogenesis
61:
57:
53:
52:transcription
49:
45:
41:
37:
35:
30:
26:
24:
16:
3820:
3808:
3720:Fluorescence
3708:Nucleic acid
3699:C57BL/6 mice
3690:Cell culture
3598:Biochemistry
3593:Cell biology
3328:irreversible
3278:
3213:Key elements
3110:Key elements
3024:Genetic code
3014:Introduction
2929:
2838:
2822:eyeless gene
2718:Evolvability
2692:Segmentation
2569:Canalisation
2539:Heterochrony
2529:Heritability
2497:Key concepts
2408:
2381:
2377:
2332:
2328:
2287:
2283:
2250:
2246:
2209:
2205:
2195:
2162:
2158:
2152:
2111:
2107:
2101:
2076:
2072:
2066:
2021:
2017:
2007:
1972:
1968:
1958:
1923:
1919:
1909:
1884:
1880:
1874:
1839:
1835:
1825:
1790:
1786:
1776:
1739:
1735:
1687:
1683:
1633:
1629:
1619:
1582:
1578:
1568:
1535:
1531:
1524:
1499:
1495:
1489:
1454:
1450:
1440:
1406:(1): 26–31.
1403:
1399:
1389:
1359:(1): 59–69.
1356:
1352:
1323:
1316:
1289:
1285:
1245:
1241:
1183:
1179:
1166:
1123:
1119:
1093:
997:
973:
931:
753:Distribution
728:
724:trans-acting
717:
712:
708:
701:chromosome 6
690:
663:
629:
627:
618:human genome
615:
610:
604:
597:
590:
585:
582:ModuleMaster
559:
549:
545:
541:
537:
529:
528:
519:feed forward
510:
506:
502:
492:
486:
465:
449:
426:
411:enhancer RNA
409:(lncRNA) or
400:
355:
339:
331:
327:
323:
322:
312:
308:
306:
301:
297:
293:
291:
279:
266:
261:
241:
239:
230:
218:
217:
212:
206:
201:
197:
189:
188:
160:
140:
135:
131:
127:
122:
117:
99:
94:
80:
39:
33:
32:
28:
22:
21:
20:
15:
3822:WikiProject
3631:Engineering
3586:Linked life
3501:Pribnow box
3459:Translation
3179:Translation
3016:to genetics
2920:Mike Levine
2829:Distal-less
2654:Polyphenism
2634:Epigenetics
2486:development
2079:: 339–372.
1653:11336/12301
1602:11336/37980
1186:: 191–212.
942:Pribnow box
787:RNA viruses
707:. The term
655:binding RNA
620:. A search
3838:Categories
3772:Epigenetic
3683:Techniques
3545:Terminator
3528:trp operon
3523:lac operon
3518:gal operon
3323:reversible
3286:lac operon
3262:imprinting
3257:Epigenetic
3249:Regulation
3204:Eukaryotic
3150:5' capping
3101:Eukaryotic
2898:Lac operon
2723:Robustness
2702:Modularity
2697:Metamerism
2603:Plasticity
2598:Pleiotropy
2551:Heterotopy
1532:Biosystems
1047:References
859:Riboswitch
670:lac operon
592:INSECT 2.0
456:repressors
436:repressors
362:eukaryotes
227:insulators
168:phenotypic
112:of nearby
87:pleiotropy
3697:(such as
3555:Repressor
3194:Bacterial
3091:Bacterial
2849:Morphogen
2834:Engrailed
2817:Pax genes
2738:Tinkering
2584:Epistasis
2579:Dominance
2490:phenotype
1552:0303-2647
1038:Transterm
957:CCAAT box
873:Eukaryota
831:Eukaryota
811:Eukaryota
807:RNA virus
783:Eukaryota
726:factors.
653:microRNAs
622:algorithm
480:Mutations
468:conserved
446:Operators
428:Silencers
417:Silencers
391:Enhancers
377:initiator
254:turned on
235:silencers
223:enhancers
180:promoters
176:enhancers
3704:Methods
3638:Concepts
3618:Genetics
3572:Silencer
3550:Enhancer
3506:TATA box
3496:Promoter
3487:Heredity
3423:Overview
3414:Glossary
3306:microRNA
3220:Ribosome
3199:Archaeal
3155:Splicing
3127:Promoter
3096:Archaeal
3040: →
3036: →
2812:Hox gene
2800:Elements
2781:Homeobox
2400:11005278
2357:16625197
2320:16422483
2312:15690032
2267:17304246
2187:29414695
2179:22421878
2144:45588146
2136:18369140
1999:16381829
1950:18621088
1901:20114053
1866:19783826
1817:15215390
1768:17199892
1742:: 1–17.
1714:16845069
1662:24008418
1611:26656931
1560:19819296
1516:14534164
1481:19530131
1432:26934309
1381:13513643
1373:22143240
1308:12381658
1262:19660565
1200:17291181
1158:20371324
1023:Promoter
952:CAAT box
937:TATA box
925:See also
892:Bacteria
869:Bacteria
850:Bacteria
779:bacteria
750:Function
678:operator
660:Examples
570:promoter
534:cofactor
515:feedback
366:TATA box
346:Promoter
287:AND gate
209:promoter
149:Overview
48:regulate
3777:Genetic
3724:Pigment
3713:Protein
3674:Wet lab
3670:Dry lab
3650:Mitosis
3482:Genetic
3473:Element
3463:protein
3404:History
3059:RNA→DNA
3054:RNA→RNA
3042:Protein
2943:Debates
2754:Systems
2680:Eyespot
2544:Neoteny
2365:9581516
2337:Bibcode
2292:Bibcode
2228:8634917
2219:1369379
2116:Bibcode
2108:Science
2093:3527045
2058:8710843
2026:Bibcode
1990:1347444
1941:2670756
1857:2808893
1759:1766362
1705:1538799
1472:2747644
1423:4802784
1149:2849075
1128:Bibcode
947:SOS box
915:Metazoa
775:Archaea
682:protein
628:Active
546:looping
452:operons
403:introns
283:OR gate
262:drivers
93:prefix
89:). The
68:anatomy
3737:-omics
3726:&
3535:Intron
3513:Operon
2844:Ligand
2524:Operon
2450:(MeSH)
2415:
2398:
2363:
2355:
2329:Nature
2318:
2310:
2284:Nature
2275:560067
2273:
2265:
2226:
2216:
2185:
2177:
2142:
2134:
2091:
2056:
2046:
1997:
1987:
1948:
1938:
1899:
1864:
1854:
1815:
1808:441523
1805:
1766:
1756:
1712:
1702:
1660:
1609:
1558:
1550:
1514:
1479:
1469:
1430:
1420:
1379:
1371:
1331:
1306:
1260:
1198:
1156:
1146:
1018:Operon
992:, mRNA
163:genome
46:which
3409:Index
3187:Types
3084:Types
2361:S2CID
2316:S2CID
2271:S2CID
2183:S2CID
2140:S2CID
2049:38642
1377:S2CID
1176:(PDF)
998:Other
905:SECIS
756:Ref.
747:Abbr.
713:trans
568:) in
499:nodes
375:, an
370:TFIIB
123:trans
114:genes
91:Latin
56:genes
31:) or
3540:Exon
2484:The
2413:ISBN
2396:PMID
2353:PMID
2308:PMID
2263:PMID
2224:PMID
2175:PMID
2132:PMID
2089:PMID
2054:PMID
1995:PMID
1946:PMID
1897:PMID
1862:PMID
1813:PMID
1764:PMID
1710:PMID
1658:PMID
1607:PMID
1556:PMID
1548:ISSN
1512:PMID
1477:PMID
1428:PMID
1369:PMID
1329:ISBN
1304:PMID
1258:PMID
1196:PMID
1154:PMID
1033:Rfam
800:IRES
744:Type
651:and
368:, a
248:and
178:and
161:The
50:the
40:CRMs
29:CREs
3854:DNA
3844:RNA
3453:RNA
3443:DNA
3038:RNA
3034:DNA
2488:of
2386:doi
2345:doi
2333:440
2300:doi
2288:433
2255:doi
2214:PMC
2206:RNA
2167:doi
2124:doi
2112:319
2081:doi
2044:PMC
2034:doi
1985:PMC
1977:doi
1936:PMC
1928:doi
1924:139
1889:doi
1885:397
1852:PMC
1844:doi
1803:PMC
1795:doi
1754:PMC
1744:doi
1700:PMC
1692:doi
1648:hdl
1638:doi
1597:hdl
1587:doi
1540:doi
1504:doi
1467:PMC
1459:doi
1418:PMC
1408:doi
1361:doi
1294:doi
1250:doi
1188:doi
1144:PMC
1136:doi
974:RNA
932:DNA
824:IRE
686:RNA
684:or
630:cis
611:cis
586:cis
550:cis
542:cis
538:cis
530:Cis
511:cis
507:cis
503:cis
487:Cis
340:cis
332:cis
328:cis
324:Cis
313:cis
302:cis
298:Cis
294:cis
258:off
256:or
242:cis
231:cis
219:Cis
213:cis
202:Cis
198:Cis
190:Cis
136:Cis
132:cis
128:cis
118:cis
102:DNA
95:cis
34:cis
23:Cis
3840::
3739:")
3722:,
3672:/
2891:+
2394:.
2382:54
2380:.
2376:.
2359:.
2351:.
2343:.
2331:.
2314:.
2306:.
2298:.
2286:.
2269:.
2261:.
2249:.
2222:.
2208:.
2204:.
2181:.
2173:.
2163:10
2161:.
2138:.
2130:.
2122:.
2110:.
2087:.
2077:55
2075:.
2052:.
2042:.
2032:.
2022:93
2020:.
2016:.
1993:.
1983:.
1973:34
1971:.
1967:.
1944:.
1934:.
1922:.
1918:.
1895:.
1883:.
1860:.
1850:.
1840:38
1838:.
1834:.
1811:.
1801:.
1791:32
1789:.
1785:.
1762:.
1752:.
1738:.
1734:.
1722:^
1708:.
1698:.
1688:34
1686:.
1682:.
1670:^
1656:.
1646:.
1634:29
1632:.
1628:.
1605:.
1595:.
1583:32
1581:.
1577:.
1554:.
1546:.
1536:99
1534:.
1510:.
1500:19
1498:.
1475:.
1465:.
1455:87
1453:.
1449:.
1426:.
1416:.
1402:.
1398:.
1375:.
1367:.
1357:13
1355:.
1343:^
1302:.
1290:16
1288:.
1284:.
1270:^
1256:.
1246:20
1244:.
1208:^
1194:.
1184:36
1182:.
1178:.
1152:.
1142:.
1134:.
1124:98
1122:.
1118:.
1102:^
1054:^
871:,
809:,
785:,
781:,
777:,
688:.
636:.
517:,
478:.
186:.
78:.
3701:)
3465:)
3461:(
3455:)
3451:(
3445:)
3441:(
3385:e
3378:t
3371:v
2998:e
2991:t
2984:v
2591:/
2477:e
2470:t
2463:v
2421:.
2402:.
2388::
2367:.
2347::
2339::
2322:.
2302::
2294::
2277:.
2257::
2251:8
2230:.
2210:2
2189:.
2169::
2146:.
2126::
2118::
2095:.
2083::
2060:.
2036::
2028::
2001:.
1979::
1952:.
1930::
1903:.
1891::
1868:.
1846::
1819:.
1797::
1770:.
1746::
1740:8
1716:.
1694::
1664:.
1650::
1640::
1613:.
1599::
1589::
1562:.
1542::
1518:.
1506::
1483:.
1461::
1434:.
1410::
1404:7
1383:.
1363::
1337:.
1310:.
1296::
1264:.
1252::
1202:.
1190::
1160:.
1138::
1130::
38:(
27:(
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.