209:. Edges between nodes represent interactions between the nodes, that can correspond to individual molecular reactions between DNA, mRNA, miRNA, proteins or molecular processes through which the products of one gene affect those of another, though the lack of experimentally obtained information often implies that some reactions are not modeled at such a fine level of detail. These interactions can be inductive (usually represented by arrowheads or the + sign), with an increase in the concentration of one leading to an increase in the other, inhibitory (represented with filled circles, blunt arrows or the minus sign), with an increase in one leading to a decrease in the other, or dual, when depending on the circumstances the regulator can activate or inhibit the target node. The nodes can regulate themselves directly or indirectly, creating feedback loops, which form cyclic chains of dependencies in the topological network. The network structure is an abstraction of the system's molecular or chemical dynamics, describing the manifold ways in which one substance affects all the others to which it is connected. In practice, such GRNs are inferred from the biological literature on a given system and represent a distillation of the collective knowledge about a set of related biochemical reactions. To speed up the manual curation of GRNs, some recent efforts try to use
412:
computational simulations. For example, fluctuations in the abundance of feed-forward loops in a model that simulates the evolution of gene regulatory networks by randomly rewiring nodes may suggest that the enrichment of feed-forward loops is a side-effect of evolution. In another model of gene regulator networks evolution, the ratio of the frequencies of gene duplication and gene deletion show great influence on network topology: certain ratios lead to the enrichment of feed-forward loops and create networks that show features of hierarchical scale free networks. De novo evolution of coherent type 1 feed-forward loops has been demonstrated computationally in response to selection for their hypothesized function of filtering out a short spurious signal, supporting adaptive evolution, but for non-idealized noise, a dynamics-based system of feed-forward regulation with different topology was instead favored.
1301:
used due to their simplicity and ability to handle noisy data but lose data information by having a binary representation of the genes. Also, artificial neural networks omit using a hidden layer so that they can be interpreted, losing the ability to model higher order correlations in the data. Using a model that is not constrained to be interpretable, a more accurate model can be produced. Being able to predict gene expressions more accurately provides a way to explore how drugs affect a system of genes as well as for finding which genes are interrelated in a process. This has been encouraged by the DREAM competition which promotes a competition for the best prediction algorithms. Some other recent work has used artificial neural networks with a hidden layer.
1284:), that can model GRNs where transcription and translation are modeled as multiple time delayed events and its dynamics is driven by a stochastic simulation algorithm (SSA) able to deal with multiple time delayed events. The time delays can be drawn from several distributions and the reaction rates from complex functions or from physical parameters. SGNSim can generate ensembles of GRNs within a set of user-defined parameters, such as topology. It can also be used to model specific GRNs and systems of chemical reactions. Genetic perturbations such as gene deletions, gene over-expression, insertions, frame shift mutations can also be modeled as well.
232:, typically AND, OR, and NOT). These functions have been interpreted as performing a kind of information processing within the cell, which determines cellular behavior. The basic drivers within cells are concentrations of some proteins, which determine both spatial (location within the cell or tissue) and temporal (cell cycle or developmental stage) coordinates of the cell, as a kind of "cellular memory". The gene networks are only beginning to be understood, and it is a next step for biology to attempt to deduce the functions for each gene "node", to help understand
364:. Network motifs can be regarded as repetitive topological patterns when dividing a big network into small blocks. Previous analysis found several types of motifs that appeared more often in gene regulatory networks than in randomly generated networks. As an example, one such motif is called feed-forward loops, which consist of three nodes. This motif is the most abundant among all possible motifs made up of three nodes, as is shown in the gene regulatory networks of fly, nematode, and human.
41:
1288:
input is assigned to an operator site and different transcription factors can be allowed, or not, to compete for the same operator site, while indirect inputs are given a target. Finally, a function is assigned to each gene, defining the gene's response to a combination of transcription factors (promoter state). The transfer functions (that is, how genes respond to a combination of inputs) can be assigned to each combination of promoter states as desired.
353:
334:
pathway. This suggests that the Hippo signaling pathway operates as a conserved regulatory module that can be used for multiple functions depending on context. Thus, changing network topology can allow a conserved module to serve multiple functions and alter the final output of the network. The second way networks can evolve is by changing the strength of interactions between nodes, such as how strongly a transcription factor may bind to a
1271:
403:, the feed-forward loop acts as a fold-change detector that responses to the fold change, rather than the absolute change, in the level of β-catenin, potentially increasing the resistance to fluctuations in β-catenin levels. Following the convergent evolution hypothesis, the enrichment of feed-forward loops would be an
33:
1287:
The GRN is created from a graph with the desired topology, imposing in-degree and out-degree distributions. Gene promoter activities are affected by other genes expression products that act as inputs, in the form of monomers or combined into multimers and set as direct or indirect. Next, each direct
1091:
From here, a set of reactions were proposed that allow generating GRNs. These are then simulated using a modified version of the
Gillespie algorithm, that can simulate multiple time delayed reactions (chemical reactions where each of the products is provided a time delay that determines when will it
1046:
Continuous network models of GRNs are an extension of the
Boolean networks described above. Nodes still represent genes and connections between them regulatory influences on gene expression. Genes in biological systems display a continuous range of activity levels and it has been argued that using a
123:
In multicellular animals the same principle has been put in the service of gene cascades that control body-shape. Each time a cell divides, two cells result which, although they contain the same genome in full, can differ in which genes are turned on and making proteins. Sometimes a 'self-sustaining
1291:
In other recent work, multiscale models of gene regulatory networks have been developed that focus on synthetic biology applications. Simulations have been used that model all biomolecular interactions in transcription, translation, regulation, and induction of gene regulatory networks, guiding the
1276:
Furthermore, there seems to be a trade-off between the noise in gene expression, the speed with which genes can switch, and the metabolic cost associated their functioning. More specifically, for any given level of metabolic cost, there is an optimal trade-off between noise and processing speed and
1037:
The validity of the model can be tested by comparing simulation results with time series observations. A partial validation of a
Boolean network model can also come from testing the predicted existence of a yet unknown regulatory connection between two particular transcription factors that each are
407:
for fast response and noise resistance. A recent research found that yeast grown in an environment of constant glucose developed mutations in glucose signaling pathways and growth regulation pathway, suggesting regulatory components responding to environmental changes are dispensable under constant
119:
In single-celled organisms, regulatory networks respond to the external environment, optimising the cell at a given time for survival in this environment. Thus a yeast cell, finding itself in a sugar solution, will turn on genes to make enzymes that process the sugar to alcohol. This process, which
1075:
Experimental results have demonstrated that gene expression is a stochastic process. Thus, many authors are now using the stochastic formalism, after the work by Arkin et al. Works on single gene expression and small synthetic genetic networks, such as the genetic toggle switch of Tim
Gardner and
1300:
Other work has focused on predicting the gene expression levels in a gene regulatory network. The approaches used to model gene regulatory networks have been constrained to be interpretable and, as a result, are generally simplified versions of the network. For example, Boolean networks have been
979:
in the equations correspond to critical cell states in which small state or parameter perturbations could switch the system between one of several stable differentiation fates. Trajectories correspond to the unfolding of biological pathways and transients of the equations to short-term biological
333:
provides a good example. The Hippo signaling pathway controls both mitotic growth and post-mitotic cellular differentiation. Recently it was found that the network the Hippo signaling pathway operates in differs between these two functions which in turn changes the behavior of the Hippo signaling
242:
of GRNs have been developed to capture the behavior of the system being modeled, and in some cases generate predictions corresponding with experimental observations. In some other cases, models have proven to make accurate novel predictions, which can be tested experimentally, thus suggesting new
371:, suggesting they are "optimal designs" for certain regulatory purposes. For example, modeling shows that feed-forward loops are able to coordinate the change in node A (in terms of concentration and activity) and the expression dynamics of node C, creating different input-output behaviors. The
1087:
Since some processes, such as gene transcription, involve many reactions and could not be correctly modeled as an instantaneous reaction in a single step, it was proposed to model these reactions as single step multiple delayed reactions in order to account for the time it takes for the entire
424:
to adapt to almost every environmental niche on earth. A network of interactions among diverse types of molecules including DNA, RNA, proteins and metabolites, is utilised by the bacteria to achieve regulation of gene expression. In bacteria, the principal function of regulatory networks is to
411:
On the other hand, some researchers hypothesize that the enrichment of network motifs is non-adaptive. In other words, gene regulatory networks can evolve to a similar structure without the specific selection on the proposed input-output behavior. Support for this hypothesis often comes from
275:
signal of histone modification are more correlated with transcription factor motifs at promoters in comparison to RNA level. Hence it is proposed that time-series histone modification ChIP-seq could provide more reliable inference of gene-regulatory networks in comparison to methods based on
1320:
There are three classes of multiple sclerosis: relapsing-remitting (RRMS), primary progressive (PPMS) and secondary progressive (SPMS). Gene regulatory network (GRN) plays a vital role to understand the disease mechanism across these three different multiple sclerosis classes.
1055:, e.g., but proteins do often control gene expression in a synergistic, i.e. non-linear, way. However, there is now a continuous network model that allows grouping of inputs to a node thus realizing another level of regulation. This model is formally closer to a higher order
323:
There are primarily two ways that networks can evolve, both of which can occur simultaneously. The first is that network topology can be changed by the addition or subtraction of nodes (genes) or parts of the network (modules) may be expressed in different contexts.
204:
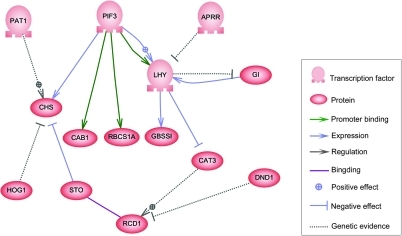
The nodes of this network can represent genes, proteins, mRNAs, protein/protein complexes or cellular processes. Nodes that are depicted as lying along vertical lines are associated with the cell/environment interfaces, while the others are free-floating and can
1112:
99:
to activate that sequence. The interaction can be direct or indirect (through transcribed RNA or translated protein). In general, each mRNA molecule goes on to make a specific protein (or set of proteins). In some cases this protein will be
1080:, provided additional experimental data on the phenotypic variability and the stochastic nature of gene expression. The first versions of stochastic models of gene expression involved only instantaneous reactions and were driven by the
396:
delays the activation of arabinose catabolism operon and transporters, potentially avoiding unnecessary metabolic transition due to temporary fluctuations in upstream signaling pathways. Similarly in the Wnt signaling pathway of
425:
control the response to environmental changes, for example nutritional status and environmental stress. A complex organization of networks permits the microorganism to coordinate and integrate multiple environmental signals.
727:
136:
gradients, which in effect provide a positioning system that tells a cell where in the body it is, and hence what sort of cell to become. A gene that is turned on in one cell may make a product that leaves the cell and
189:, interacting with molecules in the environment. Still others pass through cell membranes and mediate long range signals to other cells in a multi-cellular organism. These molecules and their interactions comprise a
4026:
BIOREL is a web-based resource for quantitative estimation of the gene network bias in relation to available database information about gene activity/function/properties/associations/interactio.
575:
197:
141:
through adjacent cells, entering them and turning on genes only when it is present above a certain threshold level. These cells are thus induced into a new fate, and may even generate other
443:. Specifically, on average, the response strength of a gene was predictable from the difference between the numbers of activating and repressing input transcription factors of that gene.
439:
After this environmental change, thousands of genes change expression level. However, these changes are predictable from the topology and logic of the gene network that is reported in
108:, i.e., a micro-machine that catalyses a certain reaction, such as the breakdown of a food source or toxin. Some proteins though serve only to activate other genes, and these are the
926:
177:
At one level, biological cells can be thought of as "partially mixed bags" of biological chemicals – in the discussion of gene regulatory networks, these chemicals are mostly the
1266:{\displaystyle {\text{RNAP}}+{\text{Pro}}_{i}{\overset {k_{i,bas}}{\longrightarrow }}{\text{Pro}}_{i}(\tau _{i}^{1})+{\text{RBS}}_{i}(\tau _{i}^{1})+{\text{RNAP}}(\tau _{i}^{2})}
1095:
For example, basic transcription of a gene can be represented by the following single-step reaction (RNAP is the RNA polymerase, RBS is the RNA ribosome binding site, and Pro
120:
we associate with wine-making, is how the yeast cell makes its living, gaining energy to multiply, which under normal circumstances would enhance its survival prospects.
185:
that arise from gene expression. These mRNA and proteins interact with each other with various degrees of specificity. Some diffuse around the cell. Others are bound to
1047:
continuous representation captures several properties of gene regulatory networks not present in the
Boolean model. Formally most of these approaches are similar to an
243:
approaches to explore in an experiment that sometimes wouldn't be considered in the design of the protocol of an experimental laboratory. Modeling techniques include
846:
811:
784:
757:
3427:
Roussel MR, Zhu R (December 2006). "Validation of an algorithm for delay stochastic simulation of transcription and translation in prokaryotic gene expression".
2234:
Mangan S, Itzkovitz S, Zaslaver A, Alon U (March 2006). "The incoherent feed-forward loop accelerates the response-time of the gal system of
Escherichia coli".
949:
615:
595:
489:
1972:
Lee TI, Rinaldi NJ, Robert F, Odom DT, Bar-Joseph Z, Gerber GK, et al. (October 2002). "Transcriptional regulatory networks in
Saccharomyces cerevisiae".
1420:
Lee TI, Rinaldi NJ, Robert F, Odom DT, Bar-Joseph Z, Gerber GK, et al. (October 2002). "Transcriptional regulatory networks in
Saccharomyces cerevisiae".
116:
region at the start of other genes they turn them on, initiating the production of another protein, and so on. Some transcription factors are inhibitory.
2984:
623:
2279:
Mangan S, Zaslaver A, Alon U (November 2003). "The coherent feedforward loop serves as a sign-sensitive delay element in transcription networks".
1530:
Leitner F, Krallinger M, Tripathi S, Kuiper M, Lægreid A, Valencia A (July 2013). "Mining cis-regulatory transcription networks from literature".
1547:"The combination of the functionalities of feedback circuits is determinant for the attractors' number and size in pathway-like Boolean networks"
104:, and will accumulate at the cell membrane or within the cell to give it particular structural properties. In other cases the protein will be an
3058:"Evolution and Morphogenesis of Differentiated Multicellular Organisms: Autonomously Generated Diffusion Gradients for Positional Information"
2832:"Boolean modelling reveals new regulatory connections between transcription factors orchestrating the development of the ventral spinal cord"
2375:"Whole genome, whole population sequencing reveals that loss of signaling networks is the major adaptive strategy in a constant environment"
4608:
951:, one obtains (possibly several) concentration profiles of proteins and mRNAs that are theoretically sustainable (though not necessarily
213:, curated databases, network inference from massive data, model checking and other information extraction technologies for this purpose.
17:
132:
modification may provide cellular memory by blocking or allowing transcription. A major feature of multicellular animals is the use of
4017:
3478:
Ribeiro A, Zhu R, Kauffman SA (November 2006). "A general modeling strategy for gene regulatory networks with stochastic dynamics".
338:. Such variation in strength of network edges has been shown to underlie between species variation in vulva cell fate patterning of
320:
to more highly connected genes. Recent work has also shown that natural selection tends to favor networks with sparse connectivity.
3870:"Gene Regulatory Networks in Peripheral Mononuclear Cells Reveals Critical Regulatory Modules and Regulators of Multiple Sclerosis"
1730:
Wagner GP, Zhang J (March 2011). "The pleiotropic structure of the genotype-phenotype map: the evolvability of complex organisms".
1008:
can model a GRN together with its gene products (the outputs) and the substances from the environment that affect it (the inputs).
1026:
For a gene, "on" corresponds to the gene being expressed; for inputs and outputs, "on" corresponds to the substance being present.
3392:
Gillespie DT (1976). "A general method for numerically simulating the stochastic time evolution of coupled chemical reactions".
4574:
4053:
360:
Another widely cited characteristic of gene regulatory network is their abundance of certain repetitive sub-networks known as
297:) and many poorly connected nodes nested within a hierarchical regulatory regime. Thus gene regulatory networks approximate a
3942:
3789:"Gene expression prediction by soft integration and the elastic net-best performance of the DREAM3 gene expression challenge"
2771:
2636:
2607:
2578:
1366:
428:
One example stress is when the environment suddenly becomes poor of nutrients. This triggers a complex adaptation process in
161:
processes, and the loss of such feedback because of a mutation can be responsible for the cell proliferation that is seen in
64:) is a collection of molecular regulators that interact with each other and with other substances in the cell to govern the
3065:
Artificial Life XI: Proceedings of the
Eleventh International Conference on the Simulation and Synthesis of Living Systems
224:. The value of the node depends on a function which depends on the value of its regulators in previous time steps (in the
4539:
3032:
3027:
Knabe JF, Nehaniv CL, Schilstra MJ (2006). "Evolutionary
Robustness of Differentiation in Genetic Regulatory Networks".
494:
388:, potentially facilitating the metabolic transition to galactose when glucose is depleted. The feed-forward loop in the
4384:
4112:
77:
4084:
3002:
Knabe JF, Nehaniv CL, Schilstra MJ, Quick T (2006). "Evolving Biological Clocks using Genetic Regulatory Networks".
4341:
4127:
4122:
468:
3186:"Stochastic kinetic analysis of developmental pathway bifurcation in phage lambda-infected Escherichia coli cells"
3341:
Gardner TS, Cantor CR, Collins JJ (January 2000). "Construction of a genetic toggle switch in Escherichia coli".
1545:
Azpeitia E, Muñoz S, González-Tokman D, Martínez-Sánchez ME, Weinstein N, Naldi A, et al. (February 2017).
1020:
in which there is an arrow from one node to another if and only if there is a causal link between the two nodes.
124:
feedback loop' ensures that a cell maintains its identity and passes it on. Less understood is the mechanism of
1012:
was amongst the first biologists to use the metaphor of Boolean networks to model genetic regulatory networks.
464:
95:
or any combination of two or more of these three that form a complex, such as a specific sequence of DNA and a
4518:
4046:
BEN: a web-based resource for exploring the connections between genes, diseases, and other biomedical entities
2988:
4369:
4041:
Tutorial: Genetic Algorithms and their Application to the Artificial Evolution of Genetic Regulatory Networks
4023:
1687:
Barabási AL, Oltvai ZN (February 2004). "Network biology: understanding the cell's functional organization".
4020:– regularly updated, contains hundreds of links to papers from bioinformatics, statistics, machine learning.
2504:"Feed-forward regulation adaptively evolves via dynamics rather than topology when there is intrinsic noise"
881:
972:
853:
1876:"Quantitative variation in autocrine signaling and pathway crosstalk in the Caenorhabditis vulval network"
967:
can usually be characterized by the sign of higher derivatives at critical points, and then correspond to
4336:
2467:
Cordero OX, Hogeweg P (October 2006). "Feed-forward loop circuits as a side effect of genome evolution".
1077:
968:
4528:
4394:
4226:
3290:
Elowitz MB, Leibler S (January 2000). "A synthetic oscillatory network of transcriptional regulators".
157:
from scratch through a series of sequential steps. They also control and maintain adult bodies through
145:
that signal back to the original cell. Over longer distances morphogens may use the active process of
4221:
3952:
Kauffman SA (March 1969). "Metabolic stability and epigenesis in randomly constructed genetic nets".
2787:
Kauffman SA (March 1969). "Metabolic stability and epigenesis in randomly constructed genetic nets".
2657:"The transcription factor network of E. coli steers global responses to shifts in RNAP concentration"
1048:
872:
471:, describing the reaction kinetics of the constituent parts. Suppose that our regulatory network has
267:. Conversely, techniques have been proposed for generating models of GRNs that best explain a set of
4035:
2293:
2248:
4603:
4593:
3997:
Plant Transcription Factor Database and Plant Transcriptional Regulation Data and Analysis Platform
3041:
3012:
1060:
1056:
3773:
3571:
2944:
2706:"Models of transcription factor binding: sensitivity of activation functions to model assumptions"
4462:
4399:
4257:
4177:
3079:
1825:
Jukam D, Xie B, Rister J, Terrell D, Charlton-Perkins M, Pistillo D, et al. (October 2013).
952:
330:
313:
165:. In parallel with this process of building structure, the gene cascade turns on genes that make
72:
and proteins which, in turn, determine the function of the cell. GRN also play a central role in
2655:
Almeida BL, Bahrudeen MN, Chauhan V, Dash S, Kandavalli V, Häkkinen A, et al. (June 2022).
1604:"Computational inference of gene regulatory networks: Approaches, limitations and opportunities"
963:
solutions to the above equation to naturally cyclic cell types. Mathematical stability of these
4447:
4346:
4331:
4300:
4201:
3036:
3007:
2939:
2628:
2599:
2570:
2288:
2243:
335:
256:
3133:
Blake WJ, KAErn M, Cantor CR, Collins JJ (April 2003). "Noise in eukaryotic gene expression".
4558:
4452:
4379:
4278:
4216:
4211:
4142:
4077:
1335:
244:
109:
3776:. Columbia University Center for Multiscale Analysis Genomic and Cellular Networks (MAGNet).
4467:
4310:
4305:
4187:
4182:
4169:
3961:
3881:
3800:
3639:
3586:
3436:
3401:
3350:
3299:
3246:
3142:
3091:
2843:
2796:
2720:
2515:
2326:"Evidence that fold-change, and not absolute level, of beta-catenin dictates Wnt signaling"
2139:
2039:
1981:
1558:
1484:
1429:
1029:
Time is viewed as proceeding in discrete steps. At each step, the new state of a node is a
824:
818:
789:
762:
735:
368:
309:
217:
96:
236:
in increasing levels of complexity, from gene to signaling pathway, cell or tissue level.
8:
4563:
1081:
290:
146:
3965:
3885:
3804:
3643:
3590:
3440:
3405:
3354:
3303:
3250:
3146:
3095:
2847:
2800:
2724:
2519:
2143:
2043:
1985:
1562:
1488:
1433:
4252:
3902:
3869:
3823:
3788:
3750:
3723:
3663:
3629:
3547:
3522:
3503:
3460:
3374:
3323:
3267:
3234:
3210:
3185:
3166:
3115:
2965:
2866:
2831:
2744:
2681:
2656:
2536:
2503:
2449:
2424:
Lynch M (October 2007). "The evolution of genetic networks by non-adaptive processes".
2401:
2374:
2350:
2325:
2211:
2186:
2108:
2060:
2027:
2005:
1954:
1900:
1875:
1851:
1826:
1799:
1774:
1755:
1712:
1579:
1546:
1453:
1315:
989:
976:
934:
860:
for the real molecular dynamics. Such models are then studied using the mathematics of
600:
580:
474:
381:
contains a feed-forward loop which accelerates the activation of galactose utilization
301:
289:
Gene regulatory networks are generally thought to be made up of a few highly connected
239:
113:
4051:
Global protein-protein interaction and gene regulation network of Arabidopsis thaliana
2187:"The incoherent feedforward loop can provide fold-change detection in gene regulation"
2162:
2127:
2026:
Boyle AP, Araya CL, Brdlik C, Cayting P, Cheng C, Cheng Y, et al. (August 2014).
1507:
1472:
4598:
4513:
4293:
4288:
4197:
3977:
3973:
3938:
3907:
3828:
3755:
3704:
3655:
3602:
3598:
3552:
3495:
3452:
3413:
3366:
3315:
3272:
3215:
3158:
3107:
3057:
2957:
2912:
2871:
2812:
2808:
2767:
2736:
2686:
2632:
2603:
2574:
2541:
2484:
2441:
2406:
2355:
2306:
2261:
2216:
2167:
2100:
2065:
1997:
1946:
1905:
1874:
Hoyos E, Kim K, Milloz J, Barkoulas M, Pénigault JB, Munro E, Félix MA (April 2011).
1856:
1804:
1747:
1704:
1666:
1625:
1584:
1512:
1445:
1402:
1398:
985:
229:
166:
101:
4497:
3699:
3682:
3667:
3464:
3448:
3327:
3119:
2830:
Lovrics A, Gao Y, Juhász B, Bock I, Byrne HM, Dinnyés A, Kovács KA (November 2014).
2748:
2453:
2028:"Comparative analysis of regulatory information and circuits across distant species"
1716:
4523:
4492:
4487:
4442:
4070:
4050:
3969:
3897:
3889:
3847:
3818:
3808:
3745:
3735:
3694:
3647:
3594:
3572:"Optimal parameter settings for information processing in gene regulatory networks"
3542:
3534:
3507:
3487:
3444:
3409:
3358:
3307:
3262:
3254:
3205:
3197:
3170:
3150:
3099:
2969:
2949:
2902:
2861:
2851:
2804:
2728:
2676:
2668:
2531:
2523:
2476:
2433:
2396:
2386:
2345:
2337:
2298:
2253:
2206:
2198:
2157:
2147:
2092:
2055:
2047:
2009:
1989:
1958:
1936:
1895:
1887:
1846:
1838:
1794:
1786:
1759:
1739:
1696:
1656:
1615:
1574:
1566:
1502:
1492:
1457:
1437:
1394:
1052:
1030:
861:
434:
317:
298:
294:
264:
252:
3378:
3201:
2705:
2112:
1643:
Kumar V, Muratani M, Rayan NA, Kraus P, Lufkin T, Ng HH, Prabhakar S (July 2013).
4533:
4437:
4247:
4057:
3848:"Time Series Gene Expression Prediction using Neural Networks with Hidden Layers"
3813:
2856:
2391:
2341:
2202:
1620:
1603:
1361:
1009:
1005:
272:
233:
225:
221:
65:
722:{\displaystyle {\frac {dS_{j}}{dt}}=f_{j}\left(S_{1},S_{2},\ldots ,S_{N}\right)}
112:
that are the main players in regulatory networks or cascades. By binding to the
4062:
3930:
3893:
3651:
2732:
2527:
2132:
Proceedings of the National Academy of Sciences of the United States of America
1477:
Proceedings of the National Academy of Sciences of the United States of America
1017:
457:
340:
216:
Genes can be viewed as nodes in the network, with input being proteins such as
2930:
Geard N, Wiles J (2005). "A gene network model for developing cell lineages".
2302:
2257:
1925:"Network motifs in the transcriptional regulation network of Escherichia coli"
1891:
617:. Then the temporal evolution of the system can be described approximately by
4587:
4501:
4415:
4283:
4117:
4094:
3855:
Proceedings of the 7th Biotechnology and Bioinformatics Symposium (BIOT 2010)
3620:
Zabet NR (September 2011). "Negative feedback and physical limits of genes".
3491:
2953:
2562:
1356:
1064:
865:
814:
786:
on the concentrations of other substances present in the cell. The functions
361:
186:
178:
150:
73:
3258:
3103:
2985:"Modelling the Regulation of Gene Expression in Genetic Regulatory Networks"
2480:
2152:
1993:
1842:
1827:"Opposite feedbacks in the Hippo pathway for growth control and neural fate"
1544:
1497:
1441:
196:
40:
4326:
4147:
4137:
3911:
3832:
3759:
3740:
3708:
3659:
3606:
3556:
3538:
3499:
3456:
3370:
3319:
3276:
3162:
3111:
2961:
2916:
2907:
2890:
2875:
2740:
2690:
2545:
2488:
2445:
2410:
2359:
2310:
2265:
2220:
2171:
2104:
2069:
2001:
1950:
1909:
1860:
1808:
1751:
1708:
1670:
1629:
1588:
1516:
1449:
993:
981:
956:
4024:
https://web.archive.org/web/20060907074456/http://mips.gsf.de/proj/biorel/
3981:
3219:
2816:
2672:
2083:
Conant GC, Wagner A (July 2003). "Convergent evolution of gene circuits".
1406:
32:
4262:
4242:
1340:
960:
849:
268:
210:
125:
4011:
3154:
2051:
1790:
1051:, as inputs to a node are summed up and the result serves as input to a
304:
topology. This is consistent with the view that most genes have limited
4506:
4206:
4159:
3029:
Proceedings of the 7th German Workshop on Artificial Life 2006 (GWAL-7)
404:
326:
305:
260:
142:
1570:
1277:
increasing the metabolic cost leads to better speed-noise trade-offs.
4457:
4425:
4192:
4098:
3996:
3362:
3311:
1661:
1644:
1346:
1330:
1016:
Each gene, each input, and each output is represented by a node in a
964:
440:
389:
372:
352:
248:
206:
154:
138:
133:
129:
2437:
1743:
1700:
4420:
4389:
4045:
2987:. Biocomputation group, University of Hertfordshire. Archived from
2096:
1645:"Uniform, optimal signal processing of mapped deep-sequencing data"
429:
421:
158:
3634:
1941:
1924:
1385:
Latchman DS (September 1996). "Inhibitory transcription factors".
959:
of kinetic equations thus correspond to potential cell types, and
4152:
1775:"Survival of the sparsest: robust gene networks are parsimonious"
1033:
of the prior states of the nodes with arrows pointing towards it.
868:
constants and sensitivities, are encoded as constant parameters.
399:
377:
182:
92:
4029:
1608:
Biochimica et Biophysica Acta (BBA) - Gene Regulatory Mechanisms
980:
events. For a more mathematical discussion, see the articles on
4132:
4040:
4018:
A bibliography on learning causal networks of gene interactions
2128:"Structure and function of the feed-forward loop network motif"
1529:
1351:
857:
382:
193:. A typical gene regulatory network looks something like this:
162:
105:
76:, the creation of body structures, which in turn is central to
4001:
3235:"Noise in gene expression: origins, consequences, and control"
1059:. The same model has also been used to mimic the evolution of
1023:
Each node in the graph can be in one of two states: on or off.
456:
For an example of modelling of the cell cycle with ODEs, see
4030:
Evolving Biological Clocks using Genetic Regulatory Networks
2185:
Goentoro L, Shoval O, Kirschner MW, Alon U (December 2009).
1387:
The International Journal of Biochemistry & Cell Biology
4430:
2654:
2233:
2184:
463:
It is common to model such a network with a set of coupled
69:
4032:– Information page with model source code and Java applet.
3935:
Computational modeling of genetic and biochemical networks
3867:
3078:
Elowitz MB, Levine AJ, Siggia ED, Swain PS (August 2002).
3004:
Proceedings of the Artificial Life X Conference (Alife 10)
3001:
451:
3845:
2501:
88:
84:
4006:
3868:
Gnanakkumaar P, Murugesan R, Ahmed SS (September 2019).
3077:
3132:
2502:
Xiong K, Lancaster AK, Siegal ML, Masel J (June 2019).
2025:
1873:
1642:
821:
enzymatic kinetics. Hence, the functional forms of the
1922:
169:
that give each cell the physical properties it needs.
3055:
3026:
1824:
1115:
937:
884:
827:
792:
765:
738:
626:
603:
583:
497:
477:
3340:
2829:
1971:
1419:
1092:
be released in the system as a "finished product").
3786:
4014:– Inference of gene association networks with GGMs
3683:"SGN Sim, a stochastic genetic networks simulator"
3680:
3477:
2278:
1923:Shen-Orr SS, Milo R, Mangan S, Alon U (May 2002).
1265:
943:
920:
840:
805:
778:
751:
721:
609:
589:
570:{\displaystyle S_{1}(t),S_{2}(t),\ldots ,S_{N}(t)}
569:
483:
3846:Smith MR, Clement M, Martinez T, Snell Q (2010).
2703:
2323:
367:The enriched motifs have been proposed to follow
312:. This structure is thought to evolve due to the
4585:
4092:
3183:
2982:
415:
3232:
3289:
2466:
2372:
1686:
817:or simple expressions derived from these e.g.
271:observations. Recently it has been shown that
4078:
3080:"Stochastic gene expression in a single cell"
2557:
2555:
1470:
3928:
3569:
3391:
2823:
2587:
2082:
1367:Weighted gene co-expression network analysis
1280:A recent work proposed a simulator (SGNSim,
44:Control process of a gene regulatory network
3787:Gustafsson M, Hörnquist M (February 2010).
3570:Chu DF, Zabet NR, Hone AN (May–June 2011).
3184:Arkin A, Ross J, McAdams HH (August 1998).
3056:Knabe JF, Schilstra MJ, Nehaniv CL (2008).
2755:
2704:Chu D, Zabet NR, Mitavskiy B (April 2009).
1729:
1636:
1538:
1070:
4085:
4071:
3426:
2983:Schilstra MJ, Bolouri H (2 January 2002).
2929:
2593:
2552:
2324:Goentoro L, Kirschner MW (December 2009).
2125:
1378:
279:
4012:Graphical Gaussian models for genome data
4007:BIB: Yeast Biological Interaction Browser
3901:
3822:
3812:
3749:
3739:
3721:
3698:
3633:
3546:
3266:
3209:
3040:
3011:
2943:
2906:
2888:
2865:
2855:
2680:
2616:
2535:
2400:
2390:
2349:
2292:
2247:
2210:
2161:
2151:
2059:
1940:
1899:
1850:
1798:
1660:
1619:
1578:
1506:
1496:
4002:Open source web service for GRN analysis
3951:
3681:Ribeiro AS, Lloyd-Price J (March 2007).
3520:
2786:
2761:
1601:
1384:
1004:The following example illustrates how a
351:
195:
39:
31:
3527:Journal of the Royal Society, Interface
2622:
2561:
2373:Kvitek DJ, Sherlock G (November 2013).
1772:
452:Coupled ordinary differential equations
14:
4586:
4575:Index of evolutionary biology articles
3523:"Computational limits to binary genes"
3233:Raser JM, O'Shea EK (September 2005).
1041:
921:{\displaystyle {\frac {dS_{j}}{dt}}=0}
36:Structure of a gene regulatory network
4066:
3619:
2891:"Neural model of the genetic network"
2650:
2648:
2423:
2021:
2019:
1820:
1818:
1309:
1682:
1680:
864:. System-specific information, like
577:represent the concentrations of the
3613:
3563:
3514:
3033:Akademische Verlagsgesellschaft AKA
2895:The Journal of Biological Chemistry
24:
4609:Evolutionary developmental biology
4385:Evolutionary developmental biology
3921:
2645:
2016:
1815:
1471:Davidson E, Levin M (April 2005).
1282:Stochastic Gene Networks Simulator
999:
813:are ultimately derived from basic
78:evolutionary developmental biology
27:Collection of molecular regulators
25:
4620:
3990:
2596:Two-Component Systems in Bacteria
2126:Mangan S, Alon U (October 2003).
1677:
597:corresponding substances at time
284:
220:, and outputs being the level of
4342:Evolution of sexual reproduction
3599:10.1016/j.biosystems.2011.01.006
3480:Journal of Computational Biology
1602:Banf M, Rhee SY (January 2017).
848:are usually chosen as low-order
347:
3861:
3839:
3780:
3766:
3715:
3674:
3471:
3420:
3385:
3334:
3283:
3226:
3177:
3126:
3071:
3049:
3020:
2995:
2976:
2923:
2882:
2780:
2697:
2625:Stress Response in Microbiology
2594:Gross R, Beier D, eds. (2012).
2495:
2469:Molecular Biology and Evolution
2460:
2417:
2366:
2317:
2272:
2227:
2178:
2119:
2076:
1965:
1916:
1867:
1766:
1532:Proceedings of BioLINK SIG 2013
1304:
1102:is the promoter region of gene
815:principles of chemical kinetics
465:ordinary differential equations
200:Example of a regulatory network
4113:Genotype–phenotype distinction
3954:Journal of Theoretical Biology
3937:. Cambridge, Mass: MIT Press.
3724:"Models for synthetic biology"
3722:Kaznessis YN (November 2007).
3622:Journal of Theoretical Biology
3521:Zabet NR, Chu DF (June 2010).
2889:Vohradsky J (September 2001).
2789:Journal of Theoretical Biology
2713:Journal of Theoretical Biology
1723:
1595:
1523:
1464:
1413:
1260:
1242:
1231:
1213:
1195:
1177:
1138:
971:of the concentration profile.
564:
558:
536:
530:
514:
508:
308:and operate within regulatory
13:
1:
4370:Regulation of gene expression
3700:10.1093/bioinformatics/btm004
3006:. MIT Press. pp. 15–21.
2567:Bacterial Regulatory Networks
1372:
1295:
1292:design of synthetic systems.
416:Bacterial regulatory networks
4540:Endless Forms Most Beautiful
4320:Evolution of genetic systems
4128:Gene–environment correlation
4123:Gene–environment interaction
3974:10.1016/0022-5193(69)90015-0
3814:10.1371/journal.pone.0009134
3414:10.1016/0021-9991(76)90041-3
2857:10.1371/journal.pone.0111430
2809:10.1016/0022-5193(69)90015-0
2392:10.1371/journal.pgen.1003972
2342:10.1016/j.molcel.2009.11.017
2281:Journal of Molecular Biology
2236:Journal of Molecular Biology
2203:10.1016/j.molcel.2009.11.018
1621:10.1016/j.bbagrm.2016.09.003
1428:(5594). Young Lab: 799–804.
1399:10.1016/1357-2725(96)00039-8
446:
7:
4519:Christiane Nüsslein-Volhard
3202:10.1093/genetics/149.4.1633
2766:. Oxford University Press.
1324:
172:
149:. Such signalling controls
18:Genetic regulatory networks
10:
4625:
4395:Hedgehog signaling pathway
4272:Developmental architecture
3894:10.1038/s41598-019-49124-x
3652:10.1016/j.jtbi.2011.06.021
2733:10.1016/j.jtbi.2008.11.026
2528:10.1038/s41467-019-10388-6
1773:Leclerc RD (August 2008).
1473:"Gene regulatory networks"
1313:
759:express the dependence of
455:
420:Regulatory networks allow
247:(ODEs), Boolean networks,
234:the behavior of the system
228:described below these are
4572:
4551:
4480:
4408:
4362:
4355:
4319:
4271:
4235:
4222:Transgressive segregation
4168:
4105:
3449:10.1088/1478-3975/3/4/005
2303:10.1016/j.jmb.2003.09.049
2258:10.1016/j.jmb.2005.12.003
1892:10.1016/j.cub.2011.02.040
1779:Molecular Systems Biology
1049:artificial neural network
4036:Engineered Gene Networks
3492:10.1089/cmb.2006.13.1630
2954:10.1162/1064546054407202
2623:Requena JM, ed. (2012).
2426:Nature Reviews. Genetics
1732:Nature Reviews. Genetics
1689:Nature Reviews. Genetics
1088:process to be complete.
1071:Stochastic gene networks
1061:cellular differentiation
1057:recurrent neural network
4400:Notch signaling pathway
4375:Gene regulatory network
4258:Dual inheritance theory
3259:10.1126/science.1105891
3104:10.1126/science.1070919
2153:10.1073/pnas.2133841100
1994:10.1126/science.1075090
1843:10.1126/science.1238016
1498:10.1073/pnas.0502024102
1442:10.1126/science.1075090
1063:and even multicellular
392:utilization systems of
331:Hippo signaling pathway
314:preferential attachment
280:Structure and evolution
257:Gaussian network models
191:gene regulatory network
4448:cis-regulatory element
4356:Control of development
4236:Non-genetic influences
4202:evolutionary landscape
3741:10.1186/1752-0509-1-47
3539:10.1098/rsif.2009.0474
2908:10.1074/jbc.M104391200
2661:Nucleic Acids Research
2629:Caister Academic Press
2600:Caister Academic Press
2571:Caister Academic Press
1267:
945:
922:
842:
807:
780:
753:
723:
611:
591:
571:
485:
375:utilization system of
357:
336:cis-regulatory element
245:differential equations
201:
45:
37:
4559:Nature versus nurture
4463:Cell surface receptor
4380:Evo-devo gene toolkit
4279:Developmental biology
4217:Polygenic inheritance
4143:Quantitative genetics
4056:16 March 2016 at the
2508:Nature Communications
2481:10.1093/molbev/msl060
1336:Cis-regulatory module
1268:
969:biochemical stability
946:
923:
843:
841:{\displaystyle f_{j}}
808:
806:{\displaystyle f_{j}}
781:
779:{\displaystyle S_{j}}
754:
752:{\displaystyle f_{j}}
724:
612:
592:
572:
486:
355:
218:transcription factors
199:
110:transcription factors
83:The regulator can be
43:
35:
4468:Transcription factor
4183:Genetic assimilation
4170:Genetic architecture
2764:The Origins of Order
2762:Kauffman SA (1993).
1649:Nature Biotechnology
1113:
1038:nodes of the model.
935:
882:
825:
790:
763:
736:
732:where the functions
624:
601:
581:
495:
475:
369:convergent evolution
153:, the building of a
97:transcription factor
4564:Morphogenetic field
4481:Influential figures
3966:1969JThBi..22..437K
3886:2019NatSR...912732G
3805:2010PLoSO...5.9134G
3774:"The DREAM Project"
3728:BMC Systems Biology
3644:2011JThBi.284...82Z
3591:2011BiSys.104...99C
3441:2006PhBio...3..274R
3406:1976JCoPh..22..403G
3355:2000Natur.403..339G
3304:2000Natur.403..335E
3251:2005Sci...309.2010R
3245:(5743): 2010–2013.
3155:10.1038/nature01546
3147:2003Natur.422..633B
3096:2002Sci...297.1183E
3090:(5584): 1183–1186.
2991:on 13 October 2007.
2901:(39): 36168–36173.
2848:2014PLoSO...9k1430L
2801:1969JThBi..22..437K
2725:2009JThBi.257..419C
2673:10.1093/nar/gkac540
2520:2019NatCo..10.2418X
2144:2003PNAS..10011980M
2138:(21): 11980–11985.
2052:10.1038/nature13668
2044:2014Natur.512..453B
1986:2002Sci...298..799L
1791:10.1038/msb.2008.52
1563:2017NatSR...742023A
1489:2005PNAS..102.4935D
1434:2002Sci...298..799L
1259:
1230:
1194:
1082:Gillespie algorithm
1042:Continuous networks
871:By solving for the
276:expression levels.
240:Mathematical models
167:structural proteins
147:signal transduction
4253:Genomic imprinting
3874:Scientific Reports
3035:. pp. 75–84.
1551:Scientific Reports
1316:Multiple sclerosis
1310:Multiple sclerosis
1263:
1245:
1216:
1180:
990:bifurcation theory
941:
918:
862:nonlinear dynamics
838:
803:
776:
749:
719:
607:
587:
567:
481:
358:
302:scale free network
202:
58:regulatory network
46:
38:
4581:
4580:
4514:Eric F. Wieschaus
4476:
4475:
4294:Pattern formation
4198:Fitness landscape
3944:978-0-262-02481-5
3857:. pp. 67–69.
3349:(6767): 339–342.
3298:(6767): 335–338.
3141:(6932): 633–637.
2773:978-0-19-505811-6
2667:(12): 6801–6819.
2638:978-1-908230-04-1
2609:978-1-908230-08-9
2580:978-1-908230-03-4
2475:(10): 1931–1936.
2038:(7515): 453–456.
1980:(5594): 799–804.
1837:(6155): 1238016.
1571:10.1038/srep42023
1240:
1205:
1169:
1163:
1128:
1119:
986:dynamical systems
944:{\displaystyle j}
910:
856:that serve as an
652:
610:{\displaystyle t}
590:{\displaystyle N}
484:{\displaystyle N}
356:Feed-forward loop
253:Bayesian networks
230:Boolean functions
16:(Redirected from
4616:
4524:William McGinnis
4493:Richard Lewontin
4488:C. H. Waddington
4360:
4359:
4337:Neutral networks
4087:
4080:
4073:
4064:
4063:
3985:
3948:
3916:
3915:
3905:
3865:
3859:
3858:
3852:
3843:
3837:
3836:
3826:
3816:
3784:
3778:
3777:
3770:
3764:
3763:
3753:
3743:
3719:
3713:
3712:
3702:
3678:
3672:
3671:
3637:
3617:
3611:
3610:
3576:
3567:
3561:
3560:
3550:
3518:
3512:
3511:
3486:(9): 1630–1639.
3475:
3469:
3468:
3429:Physical Biology
3424:
3418:
3417:
3389:
3383:
3382:
3363:10.1038/35002131
3338:
3332:
3331:
3312:10.1038/35002125
3287:
3281:
3280:
3270:
3230:
3224:
3223:
3213:
3196:(4): 1633–1648.
3181:
3175:
3174:
3130:
3124:
3123:
3075:
3069:
3068:
3062:
3053:
3047:
3046:
3044:
3024:
3018:
3017:
3015:
2999:
2993:
2992:
2980:
2974:
2973:
2947:
2927:
2921:
2920:
2910:
2886:
2880:
2879:
2869:
2859:
2827:
2821:
2820:
2784:
2778:
2777:
2759:
2753:
2752:
2710:
2701:
2695:
2694:
2684:
2652:
2643:
2642:
2620:
2614:
2613:
2591:
2585:
2584:
2559:
2550:
2549:
2539:
2499:
2493:
2492:
2464:
2458:
2457:
2421:
2415:
2414:
2404:
2394:
2385:(11): e1003972.
2370:
2364:
2363:
2353:
2321:
2315:
2314:
2296:
2276:
2270:
2269:
2251:
2242:(5): 1073–1081.
2231:
2225:
2224:
2214:
2182:
2176:
2175:
2165:
2155:
2123:
2117:
2116:
2080:
2074:
2073:
2063:
2023:
2014:
2013:
1969:
1963:
1962:
1944:
1920:
1914:
1913:
1903:
1871:
1865:
1864:
1854:
1822:
1813:
1812:
1802:
1770:
1764:
1763:
1727:
1721:
1720:
1684:
1675:
1674:
1664:
1662:10.1038/nbt.2596
1640:
1634:
1633:
1623:
1599:
1593:
1592:
1582:
1542:
1536:
1535:
1527:
1521:
1520:
1510:
1500:
1468:
1462:
1461:
1417:
1411:
1410:
1382:
1272:
1270:
1269:
1264:
1258:
1253:
1241:
1238:
1229:
1224:
1212:
1211:
1206:
1203:
1193:
1188:
1176:
1175:
1170:
1167:
1164:
1162:
1161:
1137:
1135:
1134:
1129:
1126:
1120:
1117:
1053:sigmoid function
1031:Boolean function
950:
948:
947:
942:
927:
925:
924:
919:
911:
909:
901:
900:
899:
886:
847:
845:
844:
839:
837:
836:
819:Michaelis–Menten
812:
810:
809:
804:
802:
801:
785:
783:
782:
777:
775:
774:
758:
756:
755:
750:
748:
747:
728:
726:
725:
720:
718:
714:
713:
712:
694:
693:
681:
680:
666:
665:
653:
651:
643:
642:
641:
628:
616:
614:
613:
608:
596:
594:
593:
588:
576:
574:
573:
568:
557:
556:
529:
528:
507:
506:
490:
488:
487:
482:
318:duplicated genes
21:
4624:
4623:
4619:
4618:
4617:
4615:
4614:
4613:
4604:Systems biology
4594:Gene expression
4584:
4583:
4582:
4577:
4568:
4547:
4534:Sean B. Carroll
4472:
4404:
4351:
4315:
4267:
4248:Maternal effect
4231:
4164:
4101:
4091:
4058:Wayback Machine
3993:
3988:
3945:
3924:
3922:Further reading
3919:
3866:
3862:
3850:
3844:
3840:
3785:
3781:
3772:
3771:
3767:
3720:
3716:
3679:
3675:
3618:
3614:
3585:(2–3): 99–108.
3574:
3568:
3564:
3533:(47): 945–954.
3519:
3515:
3476:
3472:
3425:
3421:
3394:J. Comput. Phys
3390:
3386:
3339:
3335:
3288:
3284:
3231:
3227:
3182:
3178:
3131:
3127:
3076:
3072:
3060:
3054:
3050:
3025:
3021:
3000:
2996:
2981:
2977:
2932:Artificial Life
2928:
2924:
2887:
2883:
2842:(11): e111430.
2828:
2824:
2785:
2781:
2774:
2760:
2756:
2708:
2702:
2698:
2653:
2646:
2639:
2621:
2617:
2610:
2592:
2588:
2581:
2560:
2553:
2500:
2496:
2465:
2461:
2438:10.1038/nrg2192
2432:(10): 803–813.
2422:
2418:
2371:
2367:
2322:
2318:
2294:10.1.1.110.4629
2277:
2273:
2249:10.1.1.184.8360
2232:
2228:
2183:
2179:
2124:
2120:
2085:Nature Genetics
2081:
2077:
2024:
2017:
1970:
1966:
1929:Nature Genetics
1921:
1917:
1880:Current Biology
1872:
1868:
1823:
1816:
1771:
1767:
1744:10.1038/nrg2949
1728:
1724:
1701:10.1038/nrg1272
1685:
1678:
1641:
1637:
1600:
1596:
1543:
1539:
1528:
1524:
1469:
1465:
1418:
1414:
1383:
1379:
1375:
1362:Systems biology
1327:
1318:
1312:
1307:
1298:
1254:
1249:
1237:
1225:
1220:
1207:
1202:
1201:
1189:
1184:
1171:
1166:
1165:
1145:
1141:
1136:
1130:
1125:
1124:
1116:
1114:
1111:
1110:
1101:
1073:
1044:
1010:Stuart Kauffman
1006:Boolean network
1002:
1000:Boolean network
973:Critical points
936:
933:
932:
902:
895:
891:
887:
885:
883:
880:
879:
875:of the system:
832:
828:
826:
823:
822:
797:
793:
791:
788:
787:
770:
766:
764:
761:
760:
743:
739:
737:
734:
733:
708:
704:
689:
685:
676:
672:
671:
667:
661:
657:
644:
637:
633:
629:
627:
625:
622:
621:
602:
599:
598:
582:
579:
578:
552:
548:
524:
520:
502:
498:
496:
493:
492:
491:nodes, and let
476:
473:
472:
461:
454:
449:
418:
350:
287:
282:
265:Process Calculi
226:Boolean network
222:gene expression
175:
66:gene expression
28:
23:
22:
15:
12:
11:
5:
4622:
4612:
4611:
4606:
4601:
4596:
4579:
4578:
4573:
4570:
4569:
4567:
4566:
4561:
4555:
4553:
4549:
4548:
4546:
4545:
4544:
4543:
4531:
4526:
4521:
4516:
4511:
4510:
4509:
4498:François Jacob
4495:
4490:
4484:
4482:
4478:
4477:
4474:
4473:
4471:
4470:
4465:
4460:
4455:
4450:
4445:
4440:
4435:
4434:
4433:
4423:
4418:
4412:
4410:
4406:
4405:
4403:
4402:
4397:
4392:
4387:
4382:
4377:
4372:
4366:
4364:
4357:
4353:
4352:
4350:
4349:
4344:
4339:
4334:
4329:
4323:
4321:
4317:
4316:
4314:
4313:
4308:
4303:
4298:
4297:
4296:
4291:
4281:
4275:
4273:
4269:
4268:
4266:
4265:
4260:
4255:
4250:
4245:
4239:
4237:
4233:
4232:
4230:
4229:
4227:Sequence space
4224:
4219:
4214:
4209:
4204:
4195:
4190:
4185:
4180:
4174:
4172:
4166:
4165:
4163:
4162:
4157:
4156:
4155:
4145:
4140:
4135:
4130:
4125:
4120:
4115:
4109:
4107:
4103:
4102:
4090:
4089:
4082:
4075:
4067:
4061:
4060:
4048:
4043:
4038:
4033:
4027:
4021:
4015:
4009:
4004:
3999:
3992:
3991:External links
3989:
3987:
3986:
3960:(3): 437–467.
3949:
3943:
3925:
3923:
3920:
3918:
3917:
3860:
3838:
3779:
3765:
3714:
3693:(6): 777–779.
3687:Bioinformatics
3673:
3612:
3562:
3513:
3470:
3435:(4): 274–284.
3419:
3384:
3333:
3282:
3225:
3176:
3125:
3070:
3048:
3042:10.1.1.71.8768
3019:
3013:10.1.1.72.5016
2994:
2975:
2938:(3): 249–267.
2922:
2881:
2822:
2795:(3): 437–467.
2779:
2772:
2754:
2719:(3): 419–429.
2696:
2644:
2637:
2615:
2608:
2586:
2579:
2565:, ed. (2012).
2551:
2494:
2459:
2416:
2365:
2336:(5): 872–884.
2330:Molecular Cell
2316:
2287:(2): 197–204.
2271:
2226:
2197:(5): 894–899.
2191:Molecular Cell
2177:
2118:
2097:10.1038/ng1181
2091:(3): 264–266.
2075:
2015:
1964:
1915:
1886:(7): 527–538.
1866:
1814:
1765:
1738:(3): 204–213.
1722:
1695:(2): 101–113.
1676:
1655:(7): 615–622.
1635:
1594:
1537:
1522:
1463:
1412:
1393:(9): 965–974.
1376:
1374:
1371:
1370:
1369:
1364:
1359:
1354:
1349:
1344:
1338:
1333:
1326:
1323:
1314:Main article:
1311:
1308:
1306:
1303:
1297:
1294:
1274:
1273:
1262:
1257:
1252:
1248:
1244:
1236:
1233:
1228:
1223:
1219:
1215:
1210:
1200:
1197:
1192:
1187:
1183:
1179:
1174:
1160:
1157:
1154:
1151:
1148:
1144:
1140:
1133:
1123:
1096:
1072:
1069:
1043:
1040:
1035:
1034:
1027:
1024:
1021:
1018:directed graph
1001:
998:
940:
929:
928:
917:
914:
908:
905:
898:
894:
890:
854:Hill functions
835:
831:
800:
796:
773:
769:
746:
742:
730:
729:
717:
711:
707:
703:
700:
697:
692:
688:
684:
679:
675:
670:
664:
660:
656:
650:
647:
640:
636:
632:
606:
586:
566:
563:
560:
555:
551:
547:
544:
541:
538:
535:
532:
527:
523:
519:
516:
513:
510:
505:
501:
480:
458:cellular model
453:
450:
448:
445:
417:
414:
362:network motifs
349:
346:
341:Caenorhabditis
286:
285:Global feature
283:
281:
278:
187:cell membranes
179:messenger RNAs
174:
171:
26:
9:
6:
4:
3:
2:
4621:
4610:
4607:
4605:
4602:
4600:
4597:
4595:
4592:
4591:
4589:
4576:
4571:
4565:
4562:
4560:
4557:
4556:
4554:
4550:
4542:
4541:
4537:
4536:
4535:
4532:
4530:
4527:
4525:
4522:
4520:
4517:
4515:
4512:
4508:
4505:
4504:
4503:
4502:Jacques Monod
4499:
4496:
4494:
4491:
4489:
4486:
4485:
4483:
4479:
4469:
4466:
4464:
4461:
4459:
4456:
4454:
4451:
4449:
4446:
4444:
4441:
4439:
4436:
4432:
4429:
4428:
4427:
4424:
4422:
4419:
4417:
4416:Homeotic gene
4414:
4413:
4411:
4407:
4401:
4398:
4396:
4393:
4391:
4388:
4386:
4383:
4381:
4378:
4376:
4373:
4371:
4368:
4367:
4365:
4361:
4358:
4354:
4348:
4345:
4343:
4340:
4338:
4335:
4333:
4330:
4328:
4325:
4324:
4322:
4318:
4312:
4309:
4307:
4304:
4302:
4299:
4295:
4292:
4290:
4287:
4286:
4285:
4284:Morphogenesis
4282:
4280:
4277:
4276:
4274:
4270:
4264:
4261:
4259:
4256:
4254:
4251:
4249:
4246:
4244:
4241:
4240:
4238:
4234:
4228:
4225:
4223:
4220:
4218:
4215:
4213:
4210:
4208:
4205:
4203:
4199:
4196:
4194:
4191:
4189:
4186:
4184:
4181:
4179:
4176:
4175:
4173:
4171:
4167:
4161:
4158:
4154:
4151:
4150:
4149:
4146:
4144:
4141:
4139:
4136:
4134:
4131:
4129:
4126:
4124:
4121:
4119:
4118:Reaction norm
4116:
4114:
4111:
4110:
4108:
4104:
4100:
4096:
4088:
4083:
4081:
4076:
4074:
4069:
4068:
4065:
4059:
4055:
4052:
4049:
4047:
4044:
4042:
4039:
4037:
4034:
4031:
4028:
4025:
4022:
4019:
4016:
4013:
4010:
4008:
4005:
4003:
4000:
3998:
3995:
3994:
3983:
3979:
3975:
3971:
3967:
3963:
3959:
3955:
3950:
3946:
3940:
3936:
3932:
3927:
3926:
3913:
3909:
3904:
3899:
3895:
3891:
3887:
3883:
3879:
3875:
3871:
3864:
3856:
3849:
3842:
3834:
3830:
3825:
3820:
3815:
3810:
3806:
3802:
3798:
3794:
3790:
3783:
3775:
3769:
3761:
3757:
3752:
3747:
3742:
3737:
3733:
3729:
3725:
3718:
3710:
3706:
3701:
3696:
3692:
3688:
3684:
3677:
3669:
3665:
3661:
3657:
3653:
3649:
3645:
3641:
3636:
3631:
3627:
3623:
3616:
3608:
3604:
3600:
3596:
3592:
3588:
3584:
3580:
3573:
3566:
3558:
3554:
3549:
3544:
3540:
3536:
3532:
3528:
3524:
3517:
3509:
3505:
3501:
3497:
3493:
3489:
3485:
3481:
3474:
3466:
3462:
3458:
3454:
3450:
3446:
3442:
3438:
3434:
3430:
3423:
3415:
3411:
3407:
3403:
3400:(4): 403–34.
3399:
3395:
3388:
3380:
3376:
3372:
3368:
3364:
3360:
3356:
3352:
3348:
3344:
3337:
3329:
3325:
3321:
3317:
3313:
3309:
3305:
3301:
3297:
3293:
3286:
3278:
3274:
3269:
3264:
3260:
3256:
3252:
3248:
3244:
3240:
3236:
3229:
3221:
3217:
3212:
3207:
3203:
3199:
3195:
3191:
3187:
3180:
3172:
3168:
3164:
3160:
3156:
3152:
3148:
3144:
3140:
3136:
3129:
3121:
3117:
3113:
3109:
3105:
3101:
3097:
3093:
3089:
3085:
3081:
3074:
3066:
3059:
3052:
3043:
3038:
3034:
3030:
3023:
3014:
3009:
3005:
2998:
2990:
2986:
2979:
2971:
2967:
2963:
2959:
2955:
2951:
2946:
2945:10.1.1.1.4742
2941:
2937:
2933:
2926:
2918:
2914:
2909:
2904:
2900:
2896:
2892:
2885:
2877:
2873:
2868:
2863:
2858:
2853:
2849:
2845:
2841:
2837:
2833:
2826:
2818:
2814:
2810:
2806:
2802:
2798:
2794:
2790:
2783:
2775:
2769:
2765:
2758:
2750:
2746:
2742:
2738:
2734:
2730:
2726:
2722:
2718:
2714:
2707:
2700:
2692:
2688:
2683:
2678:
2674:
2670:
2666:
2662:
2658:
2651:
2649:
2640:
2634:
2630:
2626:
2619:
2611:
2605:
2601:
2597:
2590:
2582:
2576:
2572:
2568:
2564:
2558:
2556:
2547:
2543:
2538:
2533:
2529:
2525:
2521:
2517:
2513:
2509:
2505:
2498:
2490:
2486:
2482:
2478:
2474:
2470:
2463:
2455:
2451:
2447:
2443:
2439:
2435:
2431:
2427:
2420:
2412:
2408:
2403:
2398:
2393:
2388:
2384:
2380:
2379:PLOS Genetics
2376:
2369:
2361:
2357:
2352:
2347:
2343:
2339:
2335:
2331:
2327:
2320:
2312:
2308:
2304:
2300:
2295:
2290:
2286:
2282:
2275:
2267:
2263:
2259:
2255:
2250:
2245:
2241:
2237:
2230:
2222:
2218:
2213:
2208:
2204:
2200:
2196:
2192:
2188:
2181:
2173:
2169:
2164:
2159:
2154:
2149:
2145:
2141:
2137:
2133:
2129:
2122:
2114:
2110:
2106:
2102:
2098:
2094:
2090:
2086:
2079:
2071:
2067:
2062:
2057:
2053:
2049:
2045:
2041:
2037:
2033:
2029:
2022:
2020:
2011:
2007:
2003:
1999:
1995:
1991:
1987:
1983:
1979:
1975:
1968:
1960:
1956:
1952:
1948:
1943:
1942:10.1038/ng881
1938:
1934:
1930:
1926:
1919:
1911:
1907:
1902:
1897:
1893:
1889:
1885:
1881:
1877:
1870:
1862:
1858:
1853:
1848:
1844:
1840:
1836:
1832:
1828:
1821:
1819:
1810:
1806:
1801:
1796:
1792:
1788:
1784:
1780:
1776:
1769:
1761:
1757:
1753:
1749:
1745:
1741:
1737:
1733:
1726:
1718:
1714:
1710:
1706:
1702:
1698:
1694:
1690:
1683:
1681:
1672:
1668:
1663:
1658:
1654:
1650:
1646:
1639:
1631:
1627:
1622:
1617:
1613:
1609:
1605:
1598:
1590:
1586:
1581:
1576:
1572:
1568:
1564:
1560:
1556:
1552:
1548:
1541:
1533:
1526:
1518:
1514:
1509:
1504:
1499:
1494:
1490:
1486:
1482:
1478:
1474:
1467:
1459:
1455:
1451:
1447:
1443:
1439:
1435:
1431:
1427:
1423:
1416:
1408:
1404:
1400:
1396:
1392:
1388:
1381:
1377:
1368:
1365:
1363:
1360:
1358:
1357:Synexpression
1355:
1353:
1350:
1348:
1345:
1342:
1339:
1337:
1334:
1332:
1329:
1328:
1322:
1317:
1302:
1293:
1289:
1285:
1283:
1278:
1255:
1250:
1246:
1234:
1226:
1221:
1217:
1208:
1198:
1190:
1185:
1181:
1172:
1158:
1155:
1152:
1149:
1146:
1142:
1131:
1121:
1109:
1108:
1107:
1105:
1100:
1093:
1089:
1085:
1083:
1079:
1068:
1066:
1065:morphogenesis
1062:
1058:
1054:
1050:
1039:
1032:
1028:
1025:
1022:
1019:
1015:
1014:
1013:
1011:
1007:
997:
995:
991:
987:
983:
978:
974:
970:
966:
962:
958:
957:Steady states
954:
938:
915:
912:
906:
903:
896:
892:
888:
878:
877:
876:
874:
869:
867:
866:reaction rate
863:
859:
855:
851:
833:
829:
820:
816:
798:
794:
771:
767:
744:
740:
715:
709:
705:
701:
698:
695:
690:
686:
682:
677:
673:
668:
662:
658:
654:
648:
645:
638:
634:
630:
620:
619:
618:
604:
584:
561:
553:
549:
545:
542:
539:
533:
525:
521:
517:
511:
503:
499:
478:
470:
466:
459:
444:
442:
438:
436:
431:
426:
423:
413:
409:
408:environment.
406:
402:
401:
395:
391:
387:
384:
380:
379:
374:
370:
365:
363:
354:
348:Local feature
345:
343:
342:
337:
332:
329:
328:
321:
319:
315:
311:
307:
303:
300:
296:
292:
277:
274:
270:
266:
262:
258:
254:
250:
246:
241:
237:
235:
231:
227:
223:
219:
214:
212:
208:
198:
194:
192:
188:
184:
180:
170:
168:
164:
160:
156:
152:
151:embryogenesis
148:
144:
140:
135:
131:
127:
121:
117:
115:
111:
107:
103:
98:
94:
90:
86:
81:
79:
75:
74:morphogenesis
71:
67:
63:
59:
55:
51:
42:
34:
30:
19:
4538:
4431:eyeless gene
4374:
4327:Evolvability
4301:Segmentation
4178:Canalisation
4148:Heterochrony
4138:Heritability
4106:Key concepts
3957:
3953:
3934:
3880:(1): 12732.
3877:
3873:
3863:
3854:
3841:
3799:(2): e9134.
3796:
3792:
3782:
3768:
3731:
3727:
3717:
3690:
3686:
3676:
3628:(1): 82–91.
3625:
3621:
3615:
3582:
3578:
3565:
3530:
3526:
3516:
3483:
3479:
3473:
3432:
3428:
3422:
3397:
3393:
3387:
3346:
3342:
3336:
3295:
3291:
3285:
3242:
3238:
3228:
3193:
3189:
3179:
3138:
3134:
3128:
3087:
3083:
3073:
3067:. MIT Press.
3064:
3051:
3028:
3022:
3003:
2997:
2989:the original
2978:
2935:
2931:
2925:
2898:
2894:
2884:
2839:
2835:
2825:
2792:
2788:
2782:
2763:
2757:
2716:
2712:
2699:
2664:
2660:
2624:
2618:
2595:
2589:
2566:
2511:
2507:
2497:
2472:
2468:
2462:
2429:
2425:
2419:
2382:
2378:
2368:
2333:
2329:
2319:
2284:
2280:
2274:
2239:
2235:
2229:
2194:
2190:
2180:
2135:
2131:
2121:
2088:
2084:
2078:
2035:
2031:
1977:
1973:
1967:
1935:(1): 64–68.
1932:
1928:
1918:
1883:
1879:
1869:
1834:
1830:
1782:
1778:
1768:
1735:
1731:
1725:
1692:
1688:
1652:
1648:
1638:
1614:(1): 41–52.
1611:
1607:
1597:
1554:
1550:
1540:
1531:
1525:
1483:(14): 4935.
1480:
1476:
1466:
1425:
1421:
1415:
1390:
1386:
1380:
1319:
1305:Applications
1299:
1290:
1286:
1281:
1279:
1275:
1103:
1098:
1094:
1090:
1086:
1074:
1045:
1036:
1003:
994:chaos theory
982:nonlinearity
977:bifurcations
930:
870:
731:
462:
433:
427:
419:
410:
398:
393:
385:
376:
366:
359:
339:
325:
322:
299:hierarchical
288:
255:, graphical
238:
215:
203:
190:
181:(mRNAs) and
176:
122:
118:
82:
80:(evo-devo).
61:
57:
53:
49:
47:
29:
4529:Mike Levine
4438:Distal-less
4263:Polyphenism
4243:Epigenetics
4095:development
3929:Bolouri H,
3579:Bio Systems
2514:(1): 2418.
1341:Genenetwork
1078:Jim Collins
961:oscillatory
873:fixed point
850:polynomials
269:time series
211:text mining
126:epigenetics
4588:Categories
4507:Lac operon
4332:Robustness
4311:Modularity
4306:Metamerism
4212:Plasticity
4207:Pleiotropy
4160:Heterotopy
3031:. Berlin:
2563:Filloux AA
1785:(1): 213.
1373:References
1343:(database)
1296:Prediction
965:attractors
467:(ODEs) or
432:, such as
405:adaptation
327:Drosophila
306:pleiotropy
261:Stochastic
249:Petri nets
143:morphogens
102:structural
68:levels of
4458:Morphogen
4443:Engrailed
4426:Pax genes
4347:Tinkering
4193:Epistasis
4188:Dominance
4099:phenotype
3635:1408.1869
3037:CiteSeerX
3008:CiteSeerX
2940:CiteSeerX
2289:CiteSeerX
2244:CiteSeerX
1557:: 42023.
1347:Morphogen
1331:Body plan
1247:τ
1218:τ
1182:τ
1139:⟶
699:…
543:…
447:Modelling
441:RegulonDB
390:arabinose
373:galactose
155:body plan
134:morphogen
130:chromatin
128:by which
4599:Networks
4421:Hox gene
4409:Elements
4390:Homeobox
4054:Archived
3933:(2001).
3931:Bower JM
3912:31484947
3833:20169069
3793:PLOS ONE
3760:17986347
3709:17267430
3668:14274912
3660:21723295
3607:21256918
3557:20007173
3500:17147485
3465:21456299
3457:17200603
3371:10659857
3328:41632754
3320:10659856
3277:16179466
3190:Genetics
3163:12687005
3120:10845628
3112:12183631
2962:16053570
2917:11395518
2876:25398016
2836:PLOS ONE
2749:12809260
2741:19121637
2691:35748858
2546:31160574
2489:16840361
2454:11839414
2446:17878896
2411:24278038
2360:20005849
2311:14607112
2266:16406067
2221:20005851
2172:14530388
2105:12819781
2070:25164757
2002:12399584
1951:11967538
1910:21458263
1861:23989952
1809:18682703
1752:21331091
1717:10950726
1709:14735121
1671:23770639
1630:27641093
1589:28186191
1517:15809445
1450:12399584
1325:See also
931:for all
430:bacteria
422:bacteria
273:ChIP-seq
183:proteins
173:Overview
159:feedback
139:diffuses
114:promoter
4552:Debates
4363:Systems
4289:Eyespot
4153:Neoteny
3982:5803332
3962:Bibcode
3903:6726613
3882:Bibcode
3824:2821917
3801:Bibcode
3751:2194732
3640:Bibcode
3587:Bibcode
3548:2871807
3508:6629364
3437:Bibcode
3402:Bibcode
3351:Bibcode
3300:Bibcode
3268:1360161
3247:Bibcode
3239:Science
3220:9691025
3211:1460268
3171:4347106
3143:Bibcode
3092:Bibcode
3084:Science
2970:8664677
2867:4232242
2844:Bibcode
2817:5803332
2797:Bibcode
2721:Bibcode
2682:9262627
2537:6546794
2516:Bibcode
2402:3836717
2351:2921914
2212:2896310
2140:Bibcode
2061:4336544
2040:Bibcode
2010:4841222
1982:Bibcode
1974:Science
1959:2180121
1901:3084603
1852:3796000
1831:Science
1800:2538912
1760:8612268
1580:5301197
1559:Bibcode
1534:: 5–12.
1485:Bibcode
1458:4841222
1430:Bibcode
1422:Science
1407:8930119
435:E. coli
400:Xenopus
378:E. coli
344:worms.
310:modules
207:diffuse
93:protein
54:genetic
4453:Ligand
4133:Operon
3980:
3941:
3910:
3900:
3831:
3821:
3758:
3748:
3734:: 47.
3707:
3666:
3658:
3605:
3555:
3545:
3506:
3498:
3463:
3455:
3379:345059
3377:
3369:
3343:Nature
3326:
3318:
3292:Nature
3275:
3265:
3218:
3208:
3169:
3161:
3135:Nature
3118:
3110:
3039:
3010:
2968:
2960:
2942:
2915:
2874:
2864:
2815:
2770:
2747:
2739:
2689:
2679:
2635:
2606:
2577:
2544:
2534:
2487:
2452:
2444:
2409:
2399:
2358:
2348:
2309:
2291:
2264:
2246:
2219:
2209:
2170:
2163:218699
2160:
2113:959172
2111:
2103:
2068:
2058:
2032:Nature
2008:
2000:
1957:
1949:
1908:
1898:
1859:
1849:
1807:
1797:
1758:
1750:
1715:
1707:
1669:
1628:
1587:
1577:
1515:
1508:556010
1505:
1456:
1448:
1405:
1352:Operon
1097:
992:, and
953:stable
858:ansatz
394:E.coli
386:galETK
383:operon
263:, and
163:cancer
106:enzyme
3851:(PDF)
3664:S2CID
3630:arXiv
3575:(PDF)
3504:S2CID
3461:S2CID
3375:S2CID
3324:S2CID
3167:S2CID
3116:S2CID
3061:(PDF)
2966:S2CID
2745:S2CID
2709:(PDF)
2450:S2CID
2109:S2CID
2006:S2CID
1955:S2CID
1756:S2CID
1713:S2CID
1454:S2CID
291:nodes
4093:The
3978:PMID
3939:ISBN
3908:PMID
3829:PMID
3756:PMID
3705:PMID
3656:PMID
3603:PMID
3553:PMID
3496:PMID
3453:PMID
3367:PMID
3316:PMID
3273:PMID
3216:PMID
3159:PMID
3108:PMID
2958:PMID
2913:PMID
2872:PMID
2813:PMID
2768:ISBN
2737:PMID
2687:PMID
2633:ISBN
2604:ISBN
2575:ISBN
2542:PMID
2485:PMID
2442:PMID
2407:PMID
2356:PMID
2307:PMID
2262:PMID
2217:PMID
2168:PMID
2101:PMID
2066:PMID
1998:PMID
1947:PMID
1906:PMID
1857:PMID
1805:PMID
1748:PMID
1705:PMID
1667:PMID
1626:PMID
1612:1860
1585:PMID
1513:PMID
1446:PMID
1403:PMID
1239:RNAP
1118:RNAP
975:and
469:SDEs
295:hubs
70:mRNA
52:(or
50:gene
4097:of
3970:doi
3898:PMC
3890:doi
3819:PMC
3809:doi
3746:PMC
3736:doi
3695:doi
3648:doi
3626:284
3595:doi
3583:104
3543:PMC
3535:doi
3488:doi
3445:doi
3410:doi
3359:doi
3347:403
3308:doi
3296:403
3263:PMC
3255:doi
3243:309
3206:PMC
3198:doi
3194:149
3151:doi
3139:422
3100:doi
3088:297
2950:doi
2903:doi
2899:276
2862:PMC
2852:doi
2805:doi
2729:doi
2717:257
2677:PMC
2669:doi
2532:PMC
2524:doi
2477:doi
2434:doi
2397:PMC
2387:doi
2346:PMC
2338:doi
2299:doi
2285:334
2254:doi
2240:356
2207:PMC
2199:doi
2158:PMC
2148:doi
2136:100
2093:doi
2056:PMC
2048:doi
2036:512
1990:doi
1978:298
1937:doi
1896:PMC
1888:doi
1847:PMC
1839:doi
1835:342
1795:PMC
1787:doi
1740:doi
1697:doi
1657:doi
1616:doi
1575:PMC
1567:doi
1503:PMC
1493:doi
1481:102
1438:doi
1426:298
1395:doi
1204:RBS
1168:Pro
1127:Pro
1106:):
955:).
852:or
324:The
316:of
89:RNA
85:DNA
62:GRN
4590::
4500:+
3976:.
3968:.
3958:22
3956:.
3906:.
3896:.
3888:.
3876:.
3872:.
3853:.
3827:.
3817:.
3807:.
3795:.
3791:.
3754:.
3744:.
3730:.
3726:.
3703:.
3691:23
3689:.
3685:.
3662:.
3654:.
3646:.
3638:.
3624:.
3601:.
3593:.
3581:.
3577:.
3551:.
3541:.
3529:.
3525:.
3502:.
3494:.
3484:13
3482:.
3459:.
3451:.
3443:.
3431:.
3408:.
3398:22
3396:.
3373:.
3365:.
3357:.
3345:.
3322:.
3314:.
3306:.
3294:.
3271:.
3261:.
3253:.
3241:.
3237:.
3214:.
3204:.
3192:.
3188:.
3165:.
3157:.
3149:.
3137:.
3114:.
3106:.
3098:.
3086:.
3082:.
3063:.
2964:.
2956:.
2948:.
2936:11
2934:.
2911:.
2897:.
2893:.
2870:.
2860:.
2850:.
2838:.
2834:.
2811:.
2803:.
2793:22
2791:.
2743:.
2735:.
2727:.
2715:.
2711:.
2685:.
2675:.
2665:50
2663:.
2659:.
2647:^
2631:.
2627:.
2602:.
2598:.
2573:.
2569:.
2554:^
2540:.
2530:.
2522:.
2512:10
2510:.
2506:.
2483:.
2473:23
2471:.
2448:.
2440:.
2428:.
2405:.
2395:.
2381:.
2377:.
2354:.
2344:.
2334:36
2332:.
2328:.
2305:.
2297:.
2283:.
2260:.
2252:.
2238:.
2215:.
2205:.
2195:36
2193:.
2189:.
2166:.
2156:.
2146:.
2134:.
2130:.
2107:.
2099:.
2089:34
2087:.
2064:.
2054:.
2046:.
2034:.
2030:.
2018:^
2004:.
1996:.
1988:.
1976:.
1953:.
1945:.
1933:31
1931:.
1927:.
1904:.
1894:.
1884:21
1882:.
1878:.
1855:.
1845:.
1833:.
1829:.
1817:^
1803:.
1793:.
1781:.
1777:.
1754:.
1746:.
1736:12
1734:.
1711:.
1703:.
1691:.
1679:^
1665:.
1653:31
1651:.
1647:.
1624:.
1610:.
1606:.
1583:.
1573:.
1565:.
1553:.
1549:.
1511:.
1501:.
1491:.
1479:.
1475:.
1452:.
1444:.
1436:.
1424:.
1401:.
1391:28
1389:.
1084:.
1067:.
996:.
988:,
984:,
259:,
251:,
91:,
87:,
56:)
48:A
4200:/
4086:e
4079:t
4072:v
3984:.
3972::
3964::
3947:.
3914:.
3892::
3884::
3878:9
3835:.
3811::
3803::
3797:5
3762:.
3738::
3732:1
3711:.
3697::
3670:.
3650::
3642::
3632::
3609:.
3597::
3589::
3559:.
3537::
3531:7
3510:.
3490::
3467:.
3447::
3439::
3433:3
3416:.
3412::
3404::
3381:.
3361::
3353::
3330:.
3310::
3302::
3279:.
3257::
3249::
3222:.
3200::
3173:.
3153::
3145::
3122:.
3102::
3094::
3045:.
3016:.
2972:.
2952::
2919:.
2905::
2878:.
2854::
2846::
2840:9
2819:.
2807::
2799::
2776:.
2751:.
2731::
2723::
2693:.
2671::
2641:.
2612:.
2583:.
2548:.
2526::
2518::
2491:.
2479::
2456:.
2436::
2430:8
2413:.
2389::
2383:9
2362:.
2340::
2313:.
2301::
2268:.
2256::
2223:.
2201::
2174:.
2150::
2142::
2115:.
2095::
2072:.
2050::
2042::
2012:.
1992::
1984::
1961:.
1939::
1912:.
1890::
1863:.
1841::
1811:.
1789::
1783:4
1762:.
1742::
1719:.
1699::
1693:5
1673:.
1659::
1632:.
1618::
1591:.
1569::
1561::
1555:7
1519:.
1495::
1487::
1460:.
1440::
1432::
1409:.
1397::
1261:)
1256:2
1251:i
1243:(
1235:+
1232:)
1227:1
1222:i
1214:(
1209:i
1199:+
1196:)
1191:1
1186:i
1178:(
1173:i
1159:s
1156:a
1153:b
1150:,
1147:i
1143:k
1132:i
1122:+
1104:i
1099:i
939:j
916:0
913:=
907:t
904:d
897:j
893:S
889:d
834:j
830:f
799:j
795:f
772:j
768:S
745:j
741:f
716:)
710:N
706:S
702:,
696:,
691:2
687:S
683:,
678:1
674:S
669:(
663:j
659:f
655:=
649:t
646:d
639:j
635:S
631:d
605:t
585:N
565:)
562:t
559:(
554:N
550:S
546:,
540:,
537:)
534:t
531:(
526:2
522:S
518:,
515:)
512:t
509:(
504:1
500:S
479:N
460:.
437:.
293:(
60:(
20:)
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.