388:
which can be used to build consensus libraries of all repeated sequences, but De novo repeat finding approaches will only identify
Helitrons that are present in multiple relatively homogeneous copies in the genome. Therefore, the low copy and older Helitrons will tend to be fragmented and have poorly defined ends. These approaches are limited by the quality of the genome assembly and the homogeneity of the repeats. Another approach is structure based which relies on the structural features of canonical Helitrons and utilizes programs such as Helitronfinder, HelSearch, Helraizer, and HelitronScanner. As these programs are trained on known Helitron elements, they may not be efficient at identifying divergent families and they generate many false positives. This approach does not create consensus sequences of the candidate Helitrons, resulting in large data sets.
380:
392:
sensitive and/or more error prone: A Rep/helicase protein-based search yields a large number of false negatives, because the majority of
Helitrons are non-autonomous elements. A similarity-based search will not identify any new families and will thus work poorly in newly studied genomes. A repeat-based search requires extensive manual curation to identify Helitron families, an overwhelming task in large genomes with substantial DNA repetition. On the basis of the overall sensitivity and specificity, the structure-based approach to identify Helitron elements is quite successful and especially useful to identify Helitron elements in a newly characterized genome. However, because at least 2 copies are needed to make an alignment, single copy Helitrons will be missed.
201:
143:
404:
of DNA not conserved for function is lost) is quite rapid. In contrast to other DNA transposons, Helitrons from some species have been reported to exhibit long-term activity probably due to the mechanism of transposition or inability of the host to recognize
Helitrons because of either sequence heterogeneity or host gene capture. In contrast, to the relatively faster unconstrained DNA half-life (2.5–14 my) of the plant and insect genomes, the mammalian DNA half-life is estimated to be much slower (884 my) which along with the minimal requirements of Helitron transposition and the slow rate of decay in mammals have caused this pattern of vertical persistence.
163:
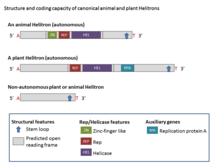
initiator (Rep) and DNA helicase (Hel) domains, which are present in a protein comprising 1000–3000 amino acids (aa) (Rep/Hel) encoded by all autonomous
Helitron elements. The Rep/Helicase protein includes zinc finger motifs, the Rep domain (which is a ~100-aa and has HUH endonuclease activity), and an eight-domain PiF1 family helicase (SuperFamily1) which are universally conserved in Helitrons. The zinc finger-like-motifs have been associated with DNA binding. The ~400-aa Hel domain is classified as a 5’ to 3’ DNA Hel which is involved in the breaking and joining of single-stranded DNA and are characterized by both the presence of the HUH motif (two
411:
reptiles, fish, invertebrates, and insect viruses. The
Helitrons present in these species have a patchy distribution and are closely related (80–98% sequence identity), despite the deep divergence times among hosts. In contrast to genes, Helitrons that have horizontally transferred into new host genomes can amplify, in some cases reaching up to several hundred copies and representing a substantial fraction of the genome. As Helitrons are known to frequently capture and amplify gene fragments, HT of this unique group of DNA transposons could lead to horizontal gene transfer and incur dramatic shifts in the trajectory of genome evolution.
223:
260:
106:. They described the structure and coding potential of canonical Helitrons and proposed the rolling-circle mechanism of transposition as well as the possibility that some of the encoded genes captured from the host are now used for replication. Their survey of the genome of these organisms showed that Helitron activity could contribute to a significant fraction (~ 2%) of the plant and invertebrate genomes where they were found, but the extent of their distribution elsewhere was not clear.
189:
simultaneously while in the sequential model they occur in a stepwise fashion. The concerted model does not require a circular intermediate although they could occur if a step fails or is bypassed during transposition. The sequential model differs in that a circular intermediate is a required step of transposition and because, until very recently, circular intermediates were not known for
Helitrons, the concerted model was adapted to explain transposition.
337:
transposition despite a reduced number of copies. The RTS indicates to the Rep-Hel protein the end of the helitron and thus the end of transposition. The whole of this information lies in the hairpin structure formed by the palindromic sequence of DNA in the 3' end. Such a small structure is likely to be modified over time, enabling to by-pass the helitron's end during its transposition and to capture neighbouring gene sequence.
333:; Transposable element capture which is based on the integration of TEs via transposition into other TEs, also called TE nesting. Despite all these proposed models, there is a lack of examples to limit the mechanism of gene capture to a single model. Further research is needed to understand the molecular mechanism behind gene capture and how it favors the survival of Helitrons.
321:
through homologous recombination, and is often accompanied by insertions of 100–4000 bp long “filler DNA” copied from diverse genomic or extra-chromosomal DNA regions into DSB. This model predicts that 2 to 8 bp regions of microhomology exist between the regions that flank the DSB in the
Helitron and that flank the original host sequence captured by the Helitron.
134:
is negligible and limited to remnants of old transposons, with the exception of bat genomes, which are populated by numerous young elements. However, many years after the description autonomous
Helitrons, no mechanistic studies have been published and therefore the rolling-circle mechanism of transposition remains a well-supported but not yet tested hypothesis.
285:
Stepwise capture would result in
Helitrons that contain gene fragments from different locations. The sequential capture model may explain Helitrons carrying multiple gene fragments observed in other organisms. There are three major models proposed in order to explain the mechanism of gene capture at the DNA level in Helitrons.
450:. This can be done by computational identification of complete young Helitrons. In a near future, detailed computer-assisted sequence studies allow investigators to understand the evolutionary history of Helitrons, together with their mechanism of gene capture and their overall significance for gene evolution.
419:
Two different scenarios describe the most likely fate of a host gene captured by
Helitrons: 1. The captured gene would be destroyed by multiple mutations if it did not provide any selective advantage to the transposons. 2. It would be kept as a gene related to the original host gene if its capture is
403:
Genome-wide analyses showed that the bulk of Helitrons tend to be quite recent. The young age of Helitron families is of course biased by the genomes that have been examined carefully, which are predominately plant and insect where the unconstrained DNA half-life (the average amount of time when half
263:
Donor sequence (black) and target sequence (blue) ; helitron divided into three parts (LTS in blue, coding sequence in grey and RTS in purple). a) tyrosine of the Rep-Hel protein cleaves 5’ end of the LTS in the donor sequence ; b) using helicase activity from 5’ to 3’, Rep-Hel rolls to the
391:
The sensitivity of the structure based approach (correctly identified/(correctly identified + false negatives)) is 93%, and the specificity (correctly identified/(correctly identified + false positives)) is 99%. There are several reasons why all other techniques for Helitron discovery have been less
302:
or capture of flanking sequence in the target site. This failure to recognize the termination signal for Helitron transposition may result in the DNA flanking the 3' end of the Helitron being transferred along with the Helitron to the donor site as well (gene capture). This may be how Helitrons have
293:
Also known as "transduction" or "read-through" model 1 (RTM1). Transposition initiates at the 5′ end and gene capture occurs if the 3′ termination signal is missed. A cryptic downstream palindrome could furnish a new terminator if the normal terminator was bypassed and all intervening sequence would
133:
In recent years, Helitrons have been identified in all eukaryotic kingdoms but their genomic copy numbers are highly variable, even among closely related species. They make up 1–5% of the genomic DNA in different fruit flies, 0–3% in mammals, >0.5% in the frog. In most mammals Helitron's presence
387:
The atypical structure, lack of target site modification, and sequence heterogeneity of Helitrons have made automated identification of Helitrons difficult. For genome-wide analysis there are two approaches that have been applied to find canonical Helitrons: De novo repeat identification approaches
311:
Also known as "read-through" model 2 (RTM2). In this model, transposition initiates at the 5′ end of a Helitron and if the 3′ end of that Helitron is missing, so transposition is terminated at the next 3′ end of a Helitron in the correct orientation, gene capture would occur. The result is that all
208:
Helitron could be either autonomous or non-autonomous. One transposase molecule cleaves at the donor (by the first tyrosine (Y1) residue of the Rep protein) and target sites (by the second tyrosine (Y2) residue) and binds to the resulting 5' ends. The free 3' OH in the target DNA attacks the DNA–Y1
231:
In 2016, one of the first mechanistic studies of helitron transposition was published in order to shed light on the different steps of transposition. Based on a consensus sequence, it reconstructed the likely ancestor of the Helibat family of helitrons present in the genome of the little brown bat
213:
for DNA synthesis by host DNA polymerase and replication proceeds to displace one strand of the helitron. If the palindrome and 3' end of the element are recognized correctly, cleavage occurs after the CTRR sequence and the one Helitron strand is transferred to the donor site where DNA replication
336:
Evidence supporting the "read-through" models seems to lie in the relative lack of importance of the 3' RTS when compared to the 5' LTS: deletion of the LTS leads to a severe reduction in the efficiency of helitron transposition, whereas the complete deletion of the RTS still leads to significant
254:
This model is supported by the fact that the deletion of one of the two tyrosines (Y727) of the Rep domain thought to be involved in cleavage of the strands doesn't actually affect the efficiency of helitron transposition. Only one of the tyrosines would be required, in order to ensure a two-step
428:
genomes are present in the genomes in multiple highly diverged families. Considering the young age of these families and the extent of protein conservation, it is highly unlikely that the divergence observed is resulted from mutations accumulated by the transposons integrated in the host genome,
410:
The impact of horizontal transfer (HT) of transposable elements may be significant due to their mutagenic potential, inherent mobility, and abundance. Researchers found evidence for the repeated HT of four different families of Helitrons in an unprecedented array of organisms, including mammals,
320:
In this model, portions of genes or non-coding regions can accidentally serve as templates during repair of double stranded breaks (DSBs) occurring in Helitrons during their transposition. Low-fidelity repair of DSB by Non-Homologous End Joining is more frequent in plants and mammals than repair
284:
within the host DNA carried by Helitrons suggested a DNA based mechanism of acquisition. Helitron gene capture was proposed to occur in a stepwise or sequential manner, i.e., gene capture occurs during one transposition and capture of a second gene occurs during a subsequent transposition event.
162:
Most Helitrons are non-autonomous elements and share common termini and other structural hallmarks with autonomous Helitrons, but they do not encode any complete set of proteins encoded by the autonomous elements. The main enzymatic hallmarks of Helitrons are the rolling-circle (RC) replication
150:
Helitrons are structurally asymmetric and are the only class of eukaryotic DNA transposons that do not generate duplications of target sites during transposition. Canonical Helitrons typically begin with a 5′ T (C/T) and terminate with the nucleotides CTRR (most frequently CTAG, but occasionally
441:
Although it is generally accepted that Helitrons are RC transposons and through numerous investigations, the role of Helitron transposition in gene duplication and shaping the genetic architecture has been proven, but neither the various mechanisms by which this occurs nor the frequency is well
250:
During transposition of the helitron, a circular intermediate is formed which was isolated in the cells transfected with the plasmid. It is formed by the joining of the terminal ends and suggests a rolling-circle model of transposition during which the cleavage of both the donor and the target
226:
a) Plasmid containing the helitron: the antibiotic resistance gene (kanamycin) is inserted between the left and right terminal sequences (LTS and RTS respectively) ; b) Circular intermediate of transposition: the terminal sequences are joined together (grey arrow indicates promoter of the
188:
Helitrons are proposed to transpose by a mechanism similar to rolling-circle replication via a single-stranded DNA intermediate. Two models are proposed for the transposition mechanism: the concerted and the sequential. In the concerted model, the donor strand cleavage and ligation occurs
297:
Indeed, in the one-ended-type fusions, the inserted fragment of donor DNA is flanked at one end (constant end) by IRR and at the other end by the CTTG or GTTC sequence present in the donor (variable end) in a way that usually results in multiple tandem insertions of the donor
86:, a small flowering plant. Despite these discoveries, the classification of Helitrons was unknown until 2001 when the discovery of protein coding-elements which were predicted to be the autonomous partners. Kapitonov and Jurka investigated the coding capacity of Helitrons in
442:
understood. At this point, it is even unclear whether the 3' terminus in a Helitron transposon initiates or terminates the Helitron replicative transposition. An important step towards investigating this mechanism would be the isolation of autonomous Helitrons active
264:
3’ end of the RTS ; c) cleavage of the 3’ end after detection of the RTS ; d) joining of the end sequences and formation of circle intermediate ; e) cleavage of the target strand and integration of the helitron after passive resolution
171:
residues that are separated by several amino acids). The PiF1 family of helicases (Hel) has 5′ to 3′ unwinding activity which for many rolling-circle entities this activity is host encoded. Plant Helitrons also encode an open reading frame with
353:. In some cases it has been seen that a Helitron insertion has provided regulatory motifs necessary for transcription initiation. Investigators presented evidence that Helitrons have contributed putative promoters, exons, splice sites,
81:
Helitrons were the first group of TEs to be discovered by computational analysis of whole genome sequences. The first Helitrons described were called Aie, AthE1, Atrep and Basho which are Non-autonomous Helitrons found in the genome of
365:
sites. Helitrons also can alter the length and sequence of both 5′ UTRs and 3′ UTRs of the coding transcripts. Another way Helitrons can control gene expression is through contributing to novel splice variants by promoting alternative
294:
be captured. In this regard, Helitrons can be viewed as an exon shuffling machines. As a random sequence provides the novel termination signal, this model does not require a high density of Helitrons in the genome.
109:
In 2003, a group of investigators studied the structure of proteins related to Helitrons and the different coding domains within them by looking for Helitron-like elements in vertebrates, specifically zebra fish,
370:
and by providing cryptic splice sites. A number of spontaneous mutations have been reported in plants that are caused by intronic Helitron insertions that result in the generation of chimeric transcript species.
176:
to single-stranded DNA-binding proteins (RPA). Typically, the RPA proteins in Helitrons are 150 – 500-aa long and are encoded by several exons. In all Helitrons, the Rep domain precedes the Hel domain.
1074:
Grabundzija, Ivana; Messing, Simon A.; Thomas, Jainy; Cosby, Rachel L.; Bilic, Ilija; Miskey, Csaba; Gogol-Döring, Andreas; Kapitonov, Vladimir; Diem, Tanja; Dalda, Anna; Jurka, Jerzy (2016-03-02).
192:
In either case, using reconstituted Helraiser transposons to study Helitron transposition, it was shown that the donor site must be double-stranded and that single-stranded donors will not suffice.
118:. The Rep/Helicase proteins were predicted to be 500 to 700 amino acids longer because of a C-terminal fusion of a domain with homology to apurinic-apyrimidinic (AP) endonuclease. Previous
247:
acting as the helitron donor. An antibiotic resistance gene was included between the two terminal sequences of the helitron to enable isolation of the cells where transposition occurred.
361:
binding sites to transcripts otherwise conserved across mammals. Helitrons drive the expression and provides de novo regulatory elements such as CAAT-box, GCbox, octamer motif, and
209:
bond and forms a bond with the donor strand resulting in strand transfer. Replication at the cleaved donor site initiates at the free 3' OH where the donor strand serves as a
1556:
126:
is nested within the Chicken Repeat 1 (CR1) clade of non-long terminal repeat (non-LTR) retrotransposons. This relationship suggested that AP endonuclease originated from a
329:
There are also other gene capture mechanism models proposed for Helitrons: Site-specific recombination model which is based on the shared features between Helitrons and
155:(16 to 20 nucleotides) hairpin about 11 bp from the 3′ end. They integrate between an AT host dinucleotide. Some families of Helitrons also carry tandem repeats, like
1613:
886:
Kosek, Dalibor; Grabundzija, Ivana; Lei, Haotian; Bilic, Ilija; Wang, Huaibin; Jin, Yukun; Peaslee, Graham F.; Hickman, Alison B.; Dyda, Fred (August 2021).
1638:
1588:
130:
insertion either nearby or within a Helitron. These investigators were not able to identify the ends of the Rep/Helicase/Endonuclease unit of Helitrons.
1648:
1770:
1688:
1678:
1593:
429:
proving that Helitrons work as a powerful tool of evolution. They have recruited host genes, modified them to an extent that is unreachable by the
1598:
69:
of host genomes. They frequently capture diverse host genes, some of which can evolve into novel host genes or become essential for Helitron
180:
The three-dimensional structure of Helitron transposase covalently bound to the left transposon end has been recently determined by cryoEM.
888:"The large bat Helitron DNA transposase forms a compact monomeric assembly that buries and protects its covalently bound 5′-transposon end"
303:
acquired additional coding sequences. Despite this hypothesis, further experiments are necessary to verify the mechanism of transposition.
1324:
379:
349:. They might get inserted within the promoter region of a gene that results in the abolition of measurable transcripts and the observed
1693:
1658:
1608:
1603:
251:
strands do not occur at the same time since a single-stranded circular DNA is first formed with one of the strands of the helitron.
1633:
1583:
70:
1628:
1533:
1653:
65:
where non-autonomous elements frequently outnumber the putative autonomous partner. Helitrons seem to have a major role in the
27:(TEs) so far described. They are the eukaryotic rolling-circle transposable elements which are hypothesized to transpose by a
697:
Poulter, Russell T.m.; Goodwin, Timothy J.d.; Butler, Margaret I. (2003). "Vertebrate Helentrons and Other Novel Helitrons".
1813:
200:
142:
1510:
1497:
1520:
420:
beneficial for the transposon, which is tolerated by the host. Helitrons, as most of other mobile elements in the
151:
variation has been noted) but do not contain terminal inverted repeats. In addition, they frequently have a short
1317:
579:
Surzycki, Stefan A; Belknap, William R. (1999). "Characterization of Repetitive DNA Elements in Arabidopsis".
1551:
1397:
1384:
1302:
1076:"A Helitron transposon reconstructed from bats reveals a novel mechanism of genome shuffling in eukaryotes"
544:
Kapitonov, Vladimir; Jurka, Jerzy (2007). "Helitrons on a Roll: Eukaryotic Rolling-circle Transposons".
1668:
1310:
210:
28:
942:"Helraiser intermediates provide insight into the mechanism of eukaryotic replicative transposition"
222:
1362:
1337:
1792:
1720:
204:
Rolling-Circle Mechanism for Helitron transposition and gene acquisition in the concerted model
103:
49:
259:
24:
1765:
1479:
1341:
1211:
1152:
1087:
1021:
953:
648:
588:
152:
37:
508:
491:
8:
1543:
1414:
32:
1215:
1156:
1091:
1025:
957:
652:
592:
1715:
1283:
1258:
1234:
1199:
1116:
1075:
982:
941:
914:
887:
863:
838:
814:
789:
612:
255:
process: 1) the cleavage of the donor DNA and 2) the integration into the target site.
758:
733:
710:
1288:
1239:
1180:
1175:
1140:
1121:
1103:
1049:
1044:
1010:"Treasures in the Attic: Rolling Circle Transposons Discovered in Eukaryotic Genomes"
1009:
987:
969:
919:
868:
819:
763:
714:
676:
671:
636:
604:
561:
513:
383:
Pipeline for genome-wide identification of candidate Helitrons and their verification
239:
the only group of mammals possessing an important number of helitrons in their genome
173:
616:
1777:
1707:
1345:
1278:
1270:
1229:
1219:
1170:
1160:
1111:
1095:
1039:
1029:
977:
961:
909:
899:
858:
850:
809:
801:
753:
745:
706:
666:
656:
596:
553:
503:
234:
1488:
1460:
1429:
1424:
904:
354:
156:
127:
123:
1760:
1618:
1575:
1403:
1393:
965:
790:"Rolling-Circle Transposons Catalyze Genomic Innovation in a Mammalian Lineage"
557:
1807:
1725:
1465:
1388:
1379:
1371:
1332:
1107:
973:
1224:
839:"Breaking and Joining Single-stranded DNA: The HUH Endonuclease Superfamily"
734:"Evidence That Chicken CR1 Elements Represent a Novel Family of Retroposons"
1737:
1292:
1259:"Pervasive Horizontal Transfer of Rolling-Circle Transposons among Animals"
1243:
1165:
1125:
1053:
1034:
991:
923:
872:
823:
718:
680:
661:
608:
565:
517:
367:
119:
43:
1184:
767:
1732:
1643:
1505:
1274:
1200:"Structure-based Discovery and Description of Plant and Animal Helitrons"
805:
749:
167:
residues separated by a hydrophobic residue) and the Y motif (one or two
102:
studies of repetitive DNA of these organisms, computational analysis and
1099:
854:
53:, and now they have been identified in a diverse range of species, from
1744:
1561:
600:
350:
20:
1139:
Mendiola, M. Victoria; Bernales, Irantzu; De La Cruz, Ferando (1994).
1663:
1354:
430:
330:
164:
146:
Structure and coding capacity of canonical animal and plant Helitrons
66:
1141:"Differential Roles of the Transposon Termini in IS91 Transposition"
1787:
1673:
1437:
1333:
443:
362:
358:
346:
168:
54:
1683:
940:
Grabundzija, Ivana; Hickman, Alison B.; Dyda, Fred (2018-03-29).
447:
299:
281:
244:
159:
and minisatellites which are generally highly mutable sequences.
62:
58:
492:"Helitrons, the Eukaryotic Rolling-circle Transposable Elements"
1782:
395:
1528:
1073:
277:
183:
1138:
345:
Helitrons, like all other TEs, are potential insertional
885:
1257:
Thomas, Jainy; Schaack, Sarah; Pritham, Ellen (2010).
939:
1256:
696:
35:
intermediate. They were first discovered in plants (
61:. Helitrons make up a substantial fraction of many
433:process, and multiplied them in the host genomes.
1805:
1007:
1204:Proceedings of the National Academy of Sciences
1145:Proceedings of the National Academy of Sciences
1014:Proceedings of the National Academy of Sciences
641:Proceedings of the National Academy of Sciences
578:
306:
1008:Feschotte, Ce´dric; Wessler, Susan R. (2001).
634:
543:
1318:
1197:
489:
374:
271:
1003:
1001:
630:
628:
626:
539:
537:
535:
533:
531:
529:
527:
396:Vertical inheritance and horizontal transfer
340:
414:
243:This active transposon was inserted into a
1325:
1311:
731:
637:"Rolling-circle Transposons in Eukaryotes"
635:Kapitonov, Vladimir; Jurka, Jerzy (2001).
485:
483:
315:
184:Mechanisms of rolling-circle transposition
1282:
1233:
1223:
1198:Yang, Lixing; Bennetzen, Jeffrey (2009).
1174:
1164:
1115:
1043:
1033:
998:
981:
913:
903:
862:
813:
783:
781:
779:
777:
757:
692:
690:
670:
660:
623:
524:
507:
481:
479:
477:
475:
473:
471:
469:
467:
465:
463:
1132:
836:
378:
258:
221:
199:
141:
1250:
837:Chandler, Michael; et al. (2013).
830:
572:
217:
1806:
787:
774:
732:Silva, Rosane; Burch, John B. (1989).
687:
490:Thomas, Jainy; Pritham, Ellen (2014).
460:
195:
1306:
1069:
1067:
1065:
1063:
935:
933:
509:10.1128/microbiolspec.mdna3-0049-2014
1191:
725:
788:Thomas, Jainy; et al. (2014).
288:
13:
1394:Short tandem repeat/Microsatellite
1060:
930:
312:intervening sequence is captured.
14:
1825:
19:are one of the three groups of
1398:Trinucleotide repeat disorders
879:
738:Molecular and Cellular Biology
581:Journal of Molecular Evolution
1:
1385:Variable number tandem repeat
711:10.1016/s0378-1119(03)00679-6
453:
1263:Genome Biology and Evolution
905:10.1016/j.molcel.2021.07.028
794:Genome Biology and Evolution
307:Chimeric transposition model
137:
7:
1814:DNA mobile genetic elements
843:Nature Reviews Microbiology
276:The presence of contiguous
214:resolves the heteroduplex.
10:
1830:
966:10.1038/s41467-018-03688-w
375:Genome-wide identification
272:Mechanisms of gene capture
76:
29:rolling circle replication
1753:
1706:
1574:
1542:
1519:
1496:
1487:
1478:
1453:
1413:
1370:
1361:
1352:
558:10.1016/j.tig.2007.08.004
436:
341:Impact on gene expression
324:
415:Evolutionary implication
122:studies showed that the
1225:10.1073/pnas.0905563106
316:Filler DNA (FDNA) model
1793:Protein tandem repeats
1721:Tandemly arrayed genes
1166:10.1073/pnas.91.5.1922
1035:10.1073/pnas.171326198
662:10.1073/pnas.151269298
384:
265:
228:
205:
147:
104:Monte Carlo simulation
96:Caenorhabditis elegans
50:Caenorhabditis elegans
47:) and in the nematode
1080:Nature Communications
946:Nature Communications
496:Microbiology Spectrum
382:
262:
225:
203:
145:
25:transposable elements
1766:Pathogenicity island
898:(20): 4271–4286.e4.
750:10.1128/mcb.9.8.3563
408:Horizontal Transfer:
218:The sequential model
153:palindromic sequence
116:Sphoeroides nephelus
84:Arabidopsis thaliana
38:Arabidopsis thaliana
1216:2009PNAS..10612832Y
1210:(31): 12832–12837.
1157:1994PNAS...91.1922M
1100:10.1038/ncomms10716
1092:2016NatCo...710716G
1026:2001PNAS...98.8923F
958:2018NatCo...9.1278G
855:10.1038/nrmicro3067
653:2001PNAS...98.8714K
593:1999JMolE..48..684S
196:The concerted model
114:and a puffer fish,
33:single-stranded DNA
1716:Gene amplification
1275:10.1093/gbe/evq050
806:10.1093/gbe/evu204
601:10.1007/pl00006512
546:Trends in Genetics
385:
266:
229:
206:
148:
1801:
1800:
1702:
1701:
1570:
1569:
1474:
1473:
1363:Repeated sequence
1338:repeated sequence
1020:(16): 8923–8924.
800:(10): 2595–2610.
647:(15): 8714–8719.
1821:
1778:Low copy repeats
1771:Symbiosis island
1708:Gene duplication
1494:
1493:
1485:
1484:
1368:
1367:
1346:gene duplication
1327:
1320:
1313:
1304:
1303:
1297:
1296:
1286:
1254:
1248:
1247:
1237:
1227:
1195:
1189:
1188:
1178:
1168:
1151:(5): 1922–1926.
1136:
1130:
1129:
1119:
1071:
1058:
1057:
1047:
1037:
1005:
996:
995:
985:
937:
928:
927:
917:
907:
883:
877:
876:
866:
834:
828:
827:
817:
785:
772:
771:
761:
744:(8): 3563–3566.
729:
723:
722:
694:
685:
684:
674:
664:
632:
621:
620:
576:
570:
569:
541:
522:
521:
511:
487:
289:End bypass model
235:Myotis Lucifugus
31:mechanism via a
1829:
1828:
1824:
1823:
1822:
1820:
1819:
1818:
1804:
1803:
1802:
1797:
1749:
1698:
1566:
1538:
1515:
1489:Retrotransposon
1470:
1461:Inverted repeat
1449:
1434:DNA transposon
1430:Retrotransposon
1425:Gene conversion
1416:
1409:
1406:
1357:
1348:
1331:
1301:
1300:
1255:
1251:
1196:
1192:
1137:
1133:
1072:
1061:
1006:
999:
938:
931:
884:
880:
835:
831:
786:
775:
730:
726:
695:
688:
633:
624:
577:
573:
552:(10): 521–529.
542:
525:
488:
461:
456:
439:
417:
398:
377:
355:polyadenylation
343:
327:
318:
309:
291:
274:
268:
256:
220:
198:
186:
157:microsatellites
140:
128:retrotransposon
124:AP endonuclease
79:
12:
11:
5:
1827:
1817:
1816:
1799:
1798:
1796:
1795:
1790:
1785:
1780:
1775:
1774:
1773:
1768:
1761:Genomic island
1757:
1755:
1751:
1750:
1748:
1747:
1742:
1741:
1740:
1730:
1729:
1728:
1718:
1712:
1710:
1704:
1703:
1700:
1699:
1697:
1696:
1691:
1686:
1681:
1676:
1671:
1666:
1661:
1656:
1651:
1646:
1641:
1636:
1631:
1626:
1621:
1616:
1611:
1606:
1601:
1596:
1591:
1586:
1580:
1578:
1576:DNA transposon
1572:
1571:
1568:
1567:
1565:
1564:
1559:
1554:
1548:
1546:
1540:
1539:
1537:
1536:
1531:
1525:
1523:
1517:
1516:
1514:
1513:
1508:
1502:
1500:
1491:
1482:
1476:
1475:
1472:
1471:
1469:
1468:
1463:
1457:
1455:
1451:
1450:
1448:
1447:
1446:
1445:
1440:
1432:
1427:
1421:
1419:
1411:
1410:
1408:
1407:
1404:Macrosatellite
1401:
1391:
1382:
1376:
1374:
1372:Tandem repeats
1365:
1359:
1358:
1353:
1350:
1349:
1330:
1329:
1322:
1315:
1307:
1299:
1298:
1249:
1190:
1131:
1059:
997:
929:
892:Molecular Cell
878:
849:(8): 525–538.
829:
773:
724:
686:
622:
587:(6): 684–691.
571:
523:
502:(4): 893–926.
458:
457:
455:
452:
438:
435:
416:
413:
397:
394:
376:
373:
342:
339:
326:
323:
317:
314:
308:
305:
290:
287:
273:
270:
219:
216:
197:
194:
185:
182:
139:
136:
78:
75:
9:
6:
4:
3:
2:
1826:
1815:
1812:
1811:
1809:
1794:
1791:
1789:
1786:
1784:
1781:
1779:
1776:
1772:
1769:
1767:
1764:
1763:
1762:
1759:
1758:
1756:
1752:
1746:
1743:
1739:
1736:
1735:
1734:
1731:
1727:
1726:Ribosomal DNA
1724:
1723:
1722:
1719:
1717:
1714:
1713:
1711:
1709:
1705:
1695:
1692:
1690:
1687:
1685:
1682:
1680:
1677:
1675:
1672:
1670:
1667:
1665:
1662:
1660:
1657:
1655:
1652:
1650:
1647:
1645:
1642:
1640:
1637:
1635:
1632:
1630:
1627:
1625:
1622:
1620:
1617:
1615:
1612:
1610:
1607:
1605:
1602:
1600:
1597:
1595:
1592:
1590:
1587:
1585:
1582:
1581:
1579:
1577:
1573:
1563:
1560:
1558:
1555:
1553:
1550:
1549:
1547:
1545:
1541:
1535:
1532:
1530:
1527:
1526:
1524:
1522:
1518:
1512:
1509:
1507:
1504:
1503:
1501:
1499:
1495:
1492:
1490:
1486:
1483:
1481:
1477:
1467:
1466:Direct repeat
1464:
1462:
1459:
1458:
1456:
1452:
1444:
1441:
1439:
1436:
1435:
1433:
1431:
1428:
1426:
1423:
1422:
1420:
1418:
1412:
1405:
1402:
1399:
1395:
1392:
1390:
1389:Minisatellite
1386:
1383:
1381:
1380:Satellite DNA
1378:
1377:
1375:
1373:
1369:
1366:
1364:
1360:
1356:
1351:
1347:
1343:
1339:
1335:
1328:
1323:
1321:
1316:
1314:
1309:
1308:
1305:
1294:
1290:
1285:
1280:
1276:
1272:
1268:
1264:
1260:
1253:
1245:
1241:
1236:
1231:
1226:
1221:
1217:
1213:
1209:
1205:
1201:
1194:
1186:
1182:
1177:
1172:
1167:
1162:
1158:
1154:
1150:
1146:
1142:
1135:
1127:
1123:
1118:
1113:
1109:
1105:
1101:
1097:
1093:
1089:
1085:
1081:
1077:
1070:
1068:
1066:
1064:
1055:
1051:
1046:
1041:
1036:
1031:
1027:
1023:
1019:
1015:
1011:
1004:
1002:
993:
989:
984:
979:
975:
971:
967:
963:
959:
955:
951:
947:
943:
936:
934:
925:
921:
916:
911:
906:
901:
897:
893:
889:
882:
874:
870:
865:
860:
856:
852:
848:
844:
840:
833:
825:
821:
816:
811:
807:
803:
799:
795:
791:
784:
782:
780:
778:
769:
765:
760:
755:
751:
747:
743:
739:
735:
728:
720:
716:
712:
708:
704:
700:
693:
691:
682:
678:
673:
668:
663:
658:
654:
650:
646:
642:
638:
631:
629:
627:
618:
614:
610:
606:
602:
598:
594:
590:
586:
582:
575:
567:
563:
559:
555:
551:
547:
540:
538:
536:
534:
532:
530:
528:
519:
515:
510:
505:
501:
497:
493:
486:
484:
482:
480:
478:
476:
474:
472:
470:
468:
466:
464:
459:
451:
449:
445:
434:
432:
427:
423:
412:
409:
405:
402:
393:
389:
381:
372:
369:
364:
360:
356:
352:
348:
338:
334:
332:
322:
313:
304:
301:
295:
286:
283:
279:
269:
261:
257:
252:
248:
246:
242:
238:
236:
224:
215:
212:
202:
193:
190:
181:
178:
175:
170:
166:
160:
158:
154:
144:
135:
131:
129:
125:
121:
117:
113:
107:
105:
101:
97:
93:
89:
85:
74:
72:
71:transposition
68:
64:
60:
56:
52:
51:
46:
45:
40:
39:
34:
30:
26:
22:
18:
1738:Gene cluster
1623:
1506:Alu sequence
1442:
1415:Interspersed
1266:
1262:
1252:
1207:
1203:
1193:
1148:
1144:
1134:
1086:(1): 10716.
1083:
1079:
1017:
1013:
949:
945:
895:
891:
881:
846:
842:
832:
797:
793:
741:
737:
727:
702:
698:
644:
640:
584:
580:
574:
549:
545:
499:
495:
440:
425:
421:
418:
407:
406:
401:Inheritance:
400:
399:
390:
386:
344:
335:
328:
319:
310:
296:
292:
275:
267:
253:
249:
240:
233:
230:
207:
191:
187:
179:
161:
149:
132:
120:phylogenetic
115:
111:
108:
99:
95:
92:Oryza sativa
91:
87:
83:
80:
48:
44:Oryza sativa
42:
36:
16:
15:
1733:Gene family
1644:Tc1/mariner
1599:EnSpm/CACTA
1269:: 656–664.
952:(1): 1278.
705:: 201–212.
422:A. thaliana
357:sites, and
112:Danio rerio
88:A. thaliana
1745:Pseudogene
1562:retroposon
1480:Transposon
1342:transposon
454:References
426:C. elegans
351:phenotypes
21:eukaryotic
1664:P element
1614:Harbinger
1355:Repeatome
1108:2041-1723
974:2041-1723
431:Mendelian
331:Integrons
165:histidine
138:Structure
100:in silico
67:evolution
17:Helitrons
1808:Category
1788:Telomere
1754:See also
1694:Zisupton
1674:Polinton
1669:PiggyBac
1624:Helitron
1443:Helitron
1438:Polinton
1334:Genetics
1293:20693155
1244:19622734
1126:26931494
1054:11481459
992:29599430
924:34403695
873:23832240
824:25223768
719:12957391
681:11447285
617:19472500
609:10229572
566:17850916
518:26350323
444:in vitro
368:splicing
363:TATA box
359:microRNA
347:mutagens
174:homology
169:tyrosine
55:protists
23:class 2
1684:Transib
1659:Novosib
1639:Kolobok
1609:Ginger2
1604:Ginger1
1589:Crypton
1284:2997563
1235:2722332
1212:Bibcode
1185:8127907
1153:Bibcode
1117:4778049
1088:Bibcode
1022:Bibcode
983:5876387
954:Bibcode
915:9364955
864:6493337
815:4224331
768:2477689
649:Bibcode
589:Bibcode
448:in vivo
300:plasmid
282:introns
245:plasmid
77:History
63:genomes
59:mammals
1783:CRISPR
1649:Merlin
1634:ISL2EU
1584:Academ
1417:repeat
1291:
1281:
1242:
1232:
1183:
1173:
1124:
1114:
1106:
1052:
1042:
990:
980:
972:
922:
912:
871:
861:
822:
812:
766:
759:362407
756:
717:
679:
669:
615:
607:
564:
516:
437:Future
325:Others
211:primer
98:using
94:, and
1689:Zator
1629:IS3EU
1534:LINE2
1529:LINE1
1521:LINEs
1498:SINEs
1454:Other
1176:43276
1045:55346
672:37501
613:S2CID
278:exons
227:gene)
1679:Sola
1654:MuDR
1594:Dada
1557:MER4
1552:HERV
1544:LTRs
1289:PMID
1240:PMID
1181:PMID
1122:PMID
1104:ISSN
1050:PMID
988:PMID
970:ISSN
920:PMID
869:PMID
820:PMID
764:PMID
715:PMID
699:Gene
677:PMID
605:PMID
562:PMID
514:PMID
446:and
424:and
280:and
41:and
1619:hAT
1511:MIR
1279:PMC
1271:doi
1230:PMC
1220:doi
1208:106
1171:PMC
1161:doi
1112:PMC
1096:doi
1040:PMC
1030:doi
978:PMC
962:doi
910:PMC
900:doi
859:PMC
851:doi
810:PMC
802:doi
754:PMC
746:doi
707:doi
703:313
667:PMC
657:doi
597:doi
554:doi
504:doi
57:to
1810::
1344:,
1340:,
1336::
1287:.
1277:.
1265:.
1261:.
1238:.
1228:.
1218:.
1206:.
1202:.
1179:.
1169:.
1159:.
1149:91
1147:.
1143:.
1120:.
1110:.
1102:.
1094:.
1082:.
1078:.
1062:^
1048:.
1038:.
1028:.
1018:98
1016:.
1012:.
1000:^
986:.
976:.
968:.
960:.
948:.
944:.
932:^
918:.
908:.
896:81
894:.
890:.
867:.
857:.
847:11
845:.
841:.
818:.
808:.
796:.
792:.
776:^
762:.
752:.
740:.
736:.
713:.
701:.
689:^
675:.
665:.
655:.
645:98
643:.
639:.
625:^
611:.
603:.
595:.
585:48
583:.
560:.
550:23
548:.
526:^
512:.
498:.
494:.
462:^
237:),
90:,
73:.
1400:)
1396:(
1387:/
1326:e
1319:t
1312:v
1295:.
1273::
1267:2
1246:.
1222::
1214::
1187:.
1163::
1155::
1128:.
1098::
1090::
1084:7
1056:.
1032::
1024::
994:.
964::
956::
950:9
926:.
902::
875:.
853::
826:.
804::
798:6
770:.
748::
742:9
721:.
709::
683:.
659::
651::
619:.
599::
591::
568:.
556::
520:.
506::
500:3
241:.
232:(
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.