1182:
settlement cue. For example, the settlement of zoospores from the green alga Ulva intestinalis onto the biofilms of specific bacteria is mediated by their attraction to the quorum-sensing molecule, acyl-homoserine lactone, secreted by the bacteria. Classic examples of marine host–microbe developmental dependence include the observation that algal cultures grown in isolation exhibited abnormal morphologies and the subsequent discovery of morphogenesis-inducing compounds, such as thallusin, secreted by epiphytic bacterial symbionts. Bacteria are also known to influence the growth of marine plants, macroalgae, and phytoplankton by secreting phytohormones such as indole acetic acid and cytokinin-type hormones. In the marine choanoflagellate
Salpingoeca rosetta, both multicellularity and reproduction are triggered by specific bacterial cues, offering a view into the origins of bacterial control over animal development (reviewed by Woznica and King. The benefit to the bacteria, in return, is that they receive physical space to colonize at particular points in the water column typically accessible only to planktonic microbes. Perhaps the best-studied example of intimate host–microbe interactions controlling animal development is the Hawaiian bobtail squid Euprymna scolopes. It lives in a mutualistic symbiosis with the bioluminescent bacteria Aliivibrio fischeri. The bacteria are fed a solution of sugars and amino acids by the host and, in return, provide bioluminescence for countershading and predator avoidance. This mutualism with microbes provides a selective advantage for the squid in predator–prey interactions. Another invertebrate example can be found in tubeworms, in which Hydroides elegans metamorphosis is mediated by a bacterial inducer and mitogen-activated protein kinase (MAPK) signaling in biofilms.
1298:
by sequestering nutrients in the form of polyphosphate granules in the tissue of their host and nitrogen cycling, e.g., through nitrification, denitrification, and ammonia oxidation.]. Many macroalgal-associated bacteria are specifically adapted to degrade complex algal polysaccharides (e.g., fucoidan, porphyran, and laminarin ) and modify both the quality and quantity of organic carbon supplied to the ecosystem. The sulfur-oxidizing gill endosymbionts of lucinid clams contribute to primary productivity through chemosynthesis and facilitate the growth of seagrasses (important foundation species) by lowering sulfide concentrations in tropical sediments. Gammaproteobacterial symbionts of lucinid clams and stilbonematid nematodes were also recently shown to be capable of nitrogen fixation (bacterial symbiont genomes encode and express nitrogenase genes, highlighting the role of symbiotic microbes in nutrient cycling in shallow marine systems.
1455:
and have a relatively simple body plan that commonly associates with bacteria, archaea, algal protists, fungi, and viruses. Sponge microbiomes are composed of specialists and generalists, and complexity of their microbiome appears to be shaped by host phylogeny. Studies have shown that the sponge microbiome contributes to nitrogen cycling in the oceans, especially through the oxidation of ammonia by archaea and bacteria. Most recently, microbial symbionts of tropical sponges were shown to produce and store polyphosphate granules, perhaps enabling the host to survive periods of phosphate depletion in oligotrophic marine environments. The microbiomes of some sponge species do appear to change in community structure in response to changing environmental conditions, including temperature and ocean acidification, as well as synergistic impacts.
1609:
1360:
1521:, and cellular debris derived from the linings of the airways which, when released into the relatively cooler outdoor air, condense to form a visible mass of vapor, which can be collected. There are various methods for collecting exhaled breath samples, one of the most recent is through the use of aerial drones. This method provides a safer, quieter, and less invasive alternative and often a cost-effective option for monitoring fauna and flora. Once obtained, the blow samples are taken to the laboratory and we proceed with the amplification and sequencing of the respiratory tract microbiota. The use of aerial drones has been more successful with large cetaceans due to slow swim speeds and larger blow sizes.
1351:
bacterial endosymbionts that reside upon coordinated use of sulfur present in the environment. This system has benefited from some of the most sophisticated 'omics and visualization tools. For example, multi-labeled probing has improved visualization of the microbiome and transcriptomics and proteomics have been applied to examine host–microbiome interactions, including energy transfer between the host and microbes and recognition of the consortia by the worm's innate immune system. The major strength of this system is that it does offer the ability to study host–microbiome interactions with a low diversity microbial consortium, and it also offers a number of host and microbial genomic resources
1311:
1567:
1269:
bacteriostatic antibiotics but also compounds like halogenated furanones, cyclic dipeptides, and acyl-homoserine lactone mimics that disrupt bacterial quorum sensing and inhibit biofilm formation. The bacteria likely are able to utilize the carbon-rich exudates from their hosts. For example, in the case of giant kelp, the alga emits approximately 20% of primary production as dissolved organic carbon. Whereas these prior examples illustrate how the microbiomes can protect hosts from surface colonization, a similar phenomenon has also been observed internally in the shipworm Bankia setacea, in which symbionts produce a boronated
1581:
717:
926:
786:, which themselves can vary throughout the ontogeny of the host and as a result of environmental perturbations. Rather than host-associated microbes functioning independently, complex multi-assemblage microbiomes have major impact on the fitness and function of their hosts. Studying these complex interactions and biological outcomes is difficult, but to understand the origin and evolution of organisms and populations and the structure and function of communities and ecosystems, the understanding of symbioses in host–microbiome systems needs advancing.
894:, while the coral provides the algae with a protected environment and limiting compounds (e.g., nitrogen species) needed for photosynthesis. However, this is a classic example of a mutualistic symbiosis that is sensitive to environmental disturbances, which can disrupt the fragile interactions between host and microbe. When reefs become warm and eutrophic, mutualistic Symbiodiniaceae may induce cellular damage to the host and/or sequester more resources for their own growth, thereby injuring and parasitizing their hosts.
1459:
1595:
1482:
732:
656:
105:
1138:
790:
biodiversity (i.e., the richness of species and their interactions) pervasively influences the functioning of Earth's ecosystems, including ecosystem productivity. However, this research has focused almost exclusively on macroorganisms. Because microbial symbionts are integral parts of most living organisms, the understanding of how microbial symbionts contribute to host performance and adaptability needs broadening.
6201:
6140:
4999:
4936:
3722:
1748:
1670:
799:
1045:
1191:
1302:
framework from the fields of microbiology, evolutionary biology, community ecology, and oceanography. Individual taxa within the microbiome may help hosts withstand a wide range of environmental conditions, including those predicted under scenarios of climate change. Next, we explore two different avenues of how interdisciplinary collaborations could advance this line of research.
19:
1494:
surrounding environment. Knowing the microbiome of the skin of marine mammals under ''normal'' conditions has allowed us to understand how these communities are different from the free microbial communities found in the sea and how they can change according to abiotic and biotic variations, and also ''
1489:
The access of microbial samples from the gut out of marine mammals is limited because most species are rare, endangered, and deep divers. There are different techniques for sampling the cetacean's gut microbiome. The most common is collecting fecal samples from the environment and taking a probe from
1386:
Studies have also suggested that resident bacteria, archaea, and fungi additionally contribute to nutrient and organic matter cycling within the coral, with viruses also possibly playing a role in structuring the composition of these members, thus providing one of the first glimpses at a multi-domain
1454:
are common members of the ocean's diverse benthic habitats and their abundance and ability to filter large volumes of seawater have led to the awareness that these organisms play critical roles in influencing benthic and pelagic processes in the ocean. They are one of the oldest lineages of animals,
1378:
are one of the more common examples of an animal host whose symbiosis with microalgae can turn to dysbiosis, and is visibly detected as bleaching. Coral microbiomes have been examined in a variety of studies, which demonstrate how variations in the ocean environment, most notably temperature, light,
1297:
organisms. Along with the coral–Symbiodiniaceae mutualism, this sponge-bacterial symbiosis helps explain Darwin's paradox, i.e., how highly productive coral reef ecosystems exist within otherwise oligotrophic tropical seas. Some sponge symbionts play a significant role in the marine phosphorus cycle
1181:
Extending beyond nutritional symbioses, microbial symbionts can alter the reproduction, development, and growth of their hosts. Specific bacterial strains in marine biofilms often directly control the recruitment of planktonic larvae and propagules, either by inhibiting settlement or by serving as a
49:. The potential for microbiomes to influence the health, physiology, behavior, and ecology of marine animals could alter current understandings of how marine animals adapt to change, and especially the growing climate-related and anthropogenic-induced changes already impacting the ocean environment.
1504:
are in danger because they are affected by multiple stress factors which make them more vulnerable to various diseases. These animals have been noted to show high susceptibility to airway infections, but very little is known about their respiratory microbiome. Therefore, the sampling of the exhaled
1336:
is one of the best studied symbiotic relationships in the sea and is a choice system for general symbiosis research. This relationship has provided insight into fundamental processes in animal-microbial symbioses, and especially biochemical interactions and signaling between the host and bacterium.
27:
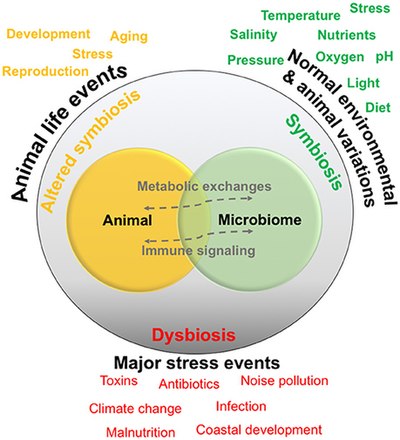
Relationships are generally thought to exist in a symbiotic state, and are normally exposed to environmental and animal-specific factors that may cause natural variations. Some events may change the relationship into a functioning but altered symbiotic state, whereas extreme stress events may cause
789:
There are many outstanding questions in ecology and evolution that could be addressed by expanding the phylogenetic and ecological breadth of host-associated microbiome studies, including all possible interactions throughout the microbiome. There is strong empirical evidence and new consensus that
1350:
is another relatively well-studied marine host to microbes. These three centimetre long worms reside within shallow marine sediments of the
Mediterranean Sea. The worms do not contain a mouth or a digestive or excretory system, but are instead nourished with the help of a suite of extracellular
1288:
Host-associated microbiomes also influence biogeochemical cycling within ecosystems with cascading effects on biodiversity and ecosystem processes. For example, microbial symbionts comprise up to 40% of the biomass of their sponge hosts. Through a process termed the "sponge-loop," they convert
1268:
of marine macroalgae excrete a diverse chemical arsenal capable of selectively shaping further bacterial colonization and deterring the settlement of biofouling marine invertebrates such as bryozoans. As in corals, these diverse, microbially secreted compounds include not only bactericidal and
966:
group) These illustrate the incorporation of various new biochemical functions, such as photosynthesis, nitrogen fixation and recycling, and methanogenesis, into protist hosts by endosymbionts. Endosymbiosis in protists is widespread and represents an important source of innovation. Previously
1301:
These examples demonstrate the importance of microbial symbioses for the functioning of ocean ecosystems. Understanding symbioses with this same level of detail in the context of complex communities (i.e., whole microbiomes) remains ripe for exploration and, indeed, requires a more integrated
1263:
and regulate microbiome assembly and maintenance in many marine organisms, including sponges, macroalgae, and corals. For example, tropical corals harbor diverse bacteria in their surface mucus layer that produce quorum-sensing inhibitors and other antibacterial compounds as a defense against
1493:
The outermost epidermal layer, i.e. the skin, is the first barrier that protects the individual from the outside world and the epidermal microbiome on it is considered an indicator not only of the health of the animal but is also considered an ecological indicator that shows the state of the
782: have revealed some of the profoundly important symbiotic roles microbes play in the lives of their hosts. These studies, however, have tended to focus on a small number of specific microbial taxa. In contrast, most hosts retain groups of many hundreds of different microbes (i.e., a
833:
nutritional symbioses with microbes. There are many examples of marine nutritional mutualisms in which microbes enable hosts to utilize resources or substrates otherwise unavailable to the host alone. Such symbioses have been described in detail in reduced and anoxic sediments (e.g.,
4645:
Kleiner, M., Wentrup, C., Lott, C., Teeling, H., Wetzel, S., Young, J., Chang, Y.J., Shah, M., VerBerkmoes, N.C., Zarzycki, J. and Fuchs, G. (2012) "Metaproteomics of a gutless marine worm and its symbiotic microbial community reveal unusual pathways for carbon and energy use".
36:
All animals on Earth form associations with microorganisms, including protists, bacteria, archaea, fungi, and viruses. In the ocean, animal–microbial relationships were historically explored in single host–symbiont systems. However, new explorations into the diversity of
5040:
Thomas, T., Moitinho-Silva, L., Lurgi, M., Björk, J.R., Easson, C., Astudillo-García, C., Olson, J.B., Erwin, P.M., López-Legentil, S., Luter, H. and Chaves-Fonnegra, A. (2016) "Diversity, structure and convergent evolution of the global sponge microbiome".
4671:
Wippler, J., Kleiner, M., Lott, C., Gruhl, A., Abraham, P.E., Giannone, R.J., Young, J.C., Hettich, R.L. and
Dubilier, N. (2016) "Transcriptomic and proteomic insights into innate immunity and adaptations to a symbiotic lifestyle in the gutless marine worm
5199:
Morrow, K.M., Bourne, D.G., Humphrey, C., Botté, E.S., Laffy, P., Zaneveld, J., Uthicke, S., Fabricius, K.E. and
Webster, N.S. (2015) "Natural volcanic CO 2 seeps reveal future trajectories for host–microbial associations in corals and sponges".
1396:
is emerging as a central member of the coral's microbiome, with flexibility in its lifestyle. Given the recent mass bleaching occurring on reefs, corals will likely continue to be a useful and popular system for symbiosis and dysbiosis research.
4564:
Dubilier, N., Mülders, C., Ferdelman, T., de Beer, D., Pernthaler, A., Klein, M., Wagner, M., Erséus, C., Thiermann, F., Krieger, J. and Giere, O. (2001) "Endosymbiotic sulphate-reducing and sulphide-oxidizing bacteria in an oligochaete worm".
4593:
Woyke, T., Teeling, H., Ivanova, N.N., Huntemann, M., Richter, M., Gloeckner, F.O., Boffelli, D., Anderson, I.J., Barry, K.W., Shapiro, H.J. and Szeto, E. (2006) "Symbiosis insights through metagenomic analysis of a microbial consortium".
4900:
Hughes, T.P., Kerry, J.T., Álvarez-Noriega, M., Álvarez-Romero, J.G., Anderson, K.D., Baird, A.H., Babcock, R.C., Beger, M., Bellwood, D.R., Berkelmans, R. and Bridge, T.C. (2017) "Global warming and recurrent mass bleaching of corals".
4397:
Webster, N.S., Negri, A.P., Botté, E.S., Laffy, P.W., Flores, F., Noonan, S., Schmidt, C. and
Uthicke, S. (2016) "Host-associated coral reef microbes respond to the cumulative pressures of ocean warming and ocean acidification".
1273:
antibiotic thought to keep the wood-digesting cecum clear of bacterial foulants. By producing antimicrobial compounds, these microbes are able to defend their niche space to prevent other organisms from crowding them out.
4619:
Schimak, M.P., Kleiner, M., Wetzel, S., Liebeke, M., Dubilier, N. and Fuchs, B.M. (2016) "MiL-FISH: Multilabeled oligonucleotides for fluorescence in situ hybridization improve visualization of bacterial cells".
4701:
Ruehland, C., Blazejak, A., Lott, C., Loy, A., Erséus, C. and
Dubilier, N. (2008) "Multiple bacterial symbionts in two species of co‐occurring gutless oligochaete worms from Mediterranean sea grass sediments".
4874:
Neave, M.J., Michell, C.T., Apprill, A. and
Voolstra, C.R. (2017) "Endozoicomonas genomes reveal functional adaptation and plasticity in bacterial strains symbiotically associated with diverse marine hosts".
4926:
1875:"Phylogenetic characterization and in situ localization of the bacterial symbiont of shipworms (Teredinidae: Bivalvia) by using 16S rRNA sequence analysis and oligodeoxynucleotide probe hybridization"
1930:
Ruehland C, Blazejak A, Lott C, Loy A, Erséus C, Dubilier N (2008). "Multiple bacterial symbionts in two species of co-occurring gutless oligochaete worms from
Mediterranean sea grass sediments".
1429:
sequences, since sequences produced by the
Illumina platform are of insufficient length (approximately 250 base pairs) for the design of primers and probes. In 2019, Goldsmith et al demonstrated
1016:
and spirilloxanthin. The endosymbionts are photosynthetically active; hence, this symbiosis represents an evolutionary transition of an aerobic organism to an anaerobic one while incorporating
5120:
Zhang, F., Blasiak, L.C., Karolin, J.O., Powell, R.J., Geddes, C.D. and Hill, R.T. (2015) "Phosphorus sequestration in the form of polyphosphate by microbial symbionts in marine sponges".
1323:
The microbiomes of diverse marine animals are currently under study, from simplistic organisms including sponges and ctenophores to more complex organisms such as sea squirts and sharks.
5690:
Johnson WR, Torralba M, Fair PA, Bossart GD, Nelson KE, Morris PJ (December 2009). "Novel diversity of bacterial communities associated with bottlenose dolphin upper respiratory tracts".
6222:
5286:
Acevedo-Whitehouse K, Rocha-Gosselin A, Gendron D (April 2010). "A novel non-invasive tool for disease surveillance of free-ranging whales and its relevance to conservation programs".
5170:
Simister, R., Taylor, M.W., Tsai, P., Fan, L., Bruxner, T.J., Crowe, M.L. and
Webster, N. (2012) "Thermal stress responses in the bacterial biosphere of the Great Barrier Reef sponge,
4260:
4004:
De Goeij JM, Van Oevelen D, Vermeij MJ, Osinga R, Middelburg JJ, De Goeij AF, et al. (2013). "Surviving in a Marine Desert: The Sponge Loop Retains Resources within Coral Reefs".
5849:
Lima N, Rogers T, Acevedo-Whitehouse K, Brown MV (February 2012). "Temporal stability and species specificity in bacteria associated with the bottlenose dolphins respiratory system".
4372:
Peixoto, R.S., Rosado, P.M., Leite, D.C.D.A., Rosado, A.S. and Bourne, D.G. (2017) "Beneficial microorganisms for corals (BMC): proposed mechanisms for coral health and resilience".
902:, are important in fostering coral recovery in the wake of disturbance. Epulopiscium bacteria in the guts of surgeonfishes produce enzymes that allow their hosts to digest complex
5502:"Microbial diversity and structure in the gastrointestinal tracts of two stranded short-finned pilot whales (Globicephala macrorhynchus) and a pygmy sperm whale (Kogia breviceps)"
3359:
Amin SA, Hmelo LR, Van Tol HM, Durham BP, Carlson LT, Heal KR, et al. (2015). "Interaction and signalling between a cosmopolitan phytoplankton and associated bacteria".
766:, i.e., persistent interactions between host and microbe in which none of the partners gets harmed and at least one of them benefits, are ubiquitous from shallow reefs to
680:
4795:
Anthony, K.R., Kline, D.I., Diaz-Pulido, G., Dove, S. and Hoegh-Guldberg, O.(2008) "Ocean acidification causes bleaching and productivity loss in coral reef builders".
1490:
the center that is non-contaminated. Besides there are studies from rectal swabs and rare studies from stranded dead or living animals direct from the intestine.
5225:
Ribes, M., Calvo, E., Movilla, J., Logares, R., Coma, R. and Pelejero, C. (2016) "Restructuring of the sponge microbiome favors tolerance to ocean acidification
808:
Black circle: macronucleus, white big circle: food vacuoles, green circles: phototrophs, brown circles: chemoautotrophs, yellow ovals: heterotrophic prokaryotes
5095:
Radax, R., Hoffmann, F., Rapp, H.T., Leininger, S. and Schleper, C. (2012) "Ammonia‐oxidizing archaea as main drivers of nitrification in cold‐water sponges".
1608:
1249:
image of the surface and scales of the fish, with arrows pointing to bacterial-sized cells and larger cells (which are not noted) are presumably phytoplankton.
918:
are critical to the functioning of Indo-Pacific coral reefs, as they are among the only fishes capable of consuming large macroalgae that bloom in the wake of
64:
effects on biodiversity and ecosystem processes. The microbiomes of diverse marine animals are currently under study, from simplistic organisms including
2827:
Hoey AS, Bellwood DR (2009). "Limited Functional Redundancy in a High Diversity System: Single Species Dominates Key Ecological Process on Coral Reefs".
5794:"Respiratory Microbiome of Endangered Southern Resident Killer Whales and Microbiota of Surrounding Sea Surface Microlayer in the Eastern North Pacific"
935:
depends on symbiotic bacteria living under its cuticle as its source of food. The bacteria are responsible for the bright white appearance of the worms.
5066:
Bayer, K., Schmitt, S. and Hentschel, U. (2008) "Physiology, phylogeny and in situ evidence for bacterial and archaeal nitrifiers in the marine sponge
2668:
Quigley KM, Bay LK, Willis BL (2018). "Leveraging new knowledge of Symbiodinium community regulation in corals for conservation and reef restoration".
4820:
Bourne, D.G., Morrow, K.M. and Webster, N.S. (2016) "Insights into the coral microbiome: underpinning the health and resilience of reef ecosystems".
5557:
Apprill A, Mooney TA, Lyman E, Stimpert AK, Rappé MS (April 2011). "Humpback whales harbour a combination of specific and variable skin bacteria".
5253:
Lesser, M.P., Fiore, C., Slattery, M. and Zaneveld, J. (2016) "Climate change stressors destabilize the microbiome of the Caribbean barrel sponge,
4845:
Neave, M.J., Apprill, A., Ferrier-Pagès, C. and Voolstra, C.R. (2016) "Diversity and function of prevalent symbiotic marine bacteria in the genus
1566:
4208:
Pfister CA, Altabet MA, Weigel BL (2019). "Kelp beds and their local effects on seawater chemistry, productivity, and microbial communities".
829:
and create the structural habitats and nutrient resources that are the foundation of their respective ecosystems. All of these taxa engage in
3139:"Revisiting the STEC Testing Approach: Using espK and espV to Make Enterohemorrhagic Escherichia coli (EHEC) Detection More Reliable in Beef"
2617:
Croft MT, Lawrence AD, Raux-Deery E, Warren MJ, Smith AG (2005). "Algae acquire vitamin B12 through a symbiotic relationship with bacteria".
1168:
1004:
3190:"Acyl-homoserine lactones modulate the settlement rate of zoospores of the marine alga Ulva intestinalis via a novel chemokinetic mechanism"
1371:. OTUs from next-generation sequencing are displayed if the OTU contained more than two sequences in the unrarefied OTU table (3626 OTUs).
4485:
Givens, C.E., Ransom, B., Bano, N. and Hollibaugh, J.T. (2015) "Comparison of the gut microbiomes of 12 bony fish and 3 shark species".
967:
unrecognized metabolic innovations of marine microbial symbioses that are ecologically important are discovered regularly. For example,
3035:
Gast RJ, Sanders RW, Caron DA (2009). "Ecological strategies of protists and their symbiotic relationships with prokaryotic microbes".
1255:
breaching and (H) a SEM image of a humpback's skin surface associated bacteria, with arrows indicating two different cell morphologies.
4456:
Blasiak, L.C., Zinder, S.H., Buckley, D.H. and Hill, R.T. (2014) "Bacterial diversity associated with the tunic of the model chordate
2349:
Duffy JE, Godwin CM, Cardinale BJ (2017). "Biodiversity effects in the wild are common and as strong as key drivers of productivity".
4927:
USGS scientists publish long-read microbiome sequences from temperate coral, providing community resource for probe and primer design
6159:"Rhodoliths holobionts in a changing ocean: host-microbes interactions mediate coralline algae resilience under ocean acidification"
4778:
Dubinsky, Z. and Jokiel, P.L. (1994) "Ratio of energy and nutrient fluxes regulates symbiosis between zooxanthellae and corals".
3229:
Provasoli L, Pintner IJ (1980). "Bacteria Induced Polymorphism in an Axenic Laboratory Strain of Ulva Lactuca (Chlorophyceae)1".
5357:
Suzuki A, Ueda K, Segawa T, Suzuki M (June 2019). "Fecal microbiota of captive Antillean manatee Trichechus manatus manatus".
6247:
1437:, while also producing longer sequences useful to the research community for probe and primer design (see diagram on right).
687:
1560:), and associated bacteria and viruses. Co-evolutionary patterns exist for coral microbial communities and coral phylogeny.
5651:"Interannual comparison of core taxa and community composition of the blow microbiota from East Australian humpback whales"
3272:
Matsuo Y, Imagawa H, Nishizawa M, Shizuri Y (2005). "Isolation of an Algal Morphogenesis Inducer from a Marine Bacterium".
287:
5394:"Microbiome Composition and Function in Aquatic Vertebrates: Small Organisms Making Big Impacts on Aquatic Animal Health"
393:
1651:
Apprill, A. (2017) "Marine animal microbiomes: toward understanding host–microbiome interactions in a changing ocean".
1580:
906:, enabling the host fish to feed on tough, leathery red and brown macroalgae. This trophic innovation has facilitated
6232:
1505:
breath or "blow" of the cetaceans can provide an assessment of the state of health. Blow is composed of a mixture of
1359:
1161:
388:
4510:
McFall-Ngai, M.J. (2000) "Negotiations between animals and bacteria: the 'diplomacy'of the squid-vibrio symbiosis".
2519:"The importance of sponges and mangroves in supporting fish communities on degraded coral reefs in Caribbean Panama"
1204:
292:
3896:"Comparing and Evaluating Metagenome Assembly Tools from a Microbiologist's Perspective - Not Only Size Matters!"
45:
hosts is moving the field into studies that address interactions between the animal host and a more multi-member
5943:
Knowlton N, Rohwer F (October 2003). "Multispecies microbial mutualisms on coral reefs: the host as a habitat".
3844:"Patterns and controls of reef-scale production of dissolved organic carbon by giant kelp M acrocystis pyrifera"
1615:
3622:"Characterization of thegacA-dependent surface and coral mucus colonization by an opportunistic coral pathogen
473:
1310:
2921:"Caribbean Spiny Lobster Fishery is Underpinned by Trophic Subsidies from Chemosynthetic Primary Production"
56:
that do not live in close relationship with a microbial partner. Host-associated microbiomes also influence
1417:
to study microbial community interactions associated with symbiotic state. However, the ability to develop
1246:
1154:
1114:
572:
57:
2570:"Host-Microbe Coevolution: Applying Evidence from Model Systems to Complex Marine Invertebrate Holobionts"
4116:"Evolution of a Vegetarian Vibrio: Metabolic Specialization of Vibrio breoganii to Macroalgal Substrates"
1283:
1119:
226:
6047:"Microbes in the coral holobiont: partners through evolution, development, and ecological interactions"
4730:"Season, but not symbiont state, drives microbiome structure in the temperate coral Astrangia poculata"
1434:
1418:
1368:
1290:
767:
701:
199:
4293:
3072:"Recruitment in the sea: Bacterial genes required for inducing larval settlement in a polychaete worm"
1409:, widely documented along the eastern coast of the United States. The coral can live with and without
1379:
and inorganic nutrients, affect the abundance and performance of the microalgal symbionts, as well as
798:
3955:"Boronated tartrolon antibiotic produced by symbiotic cellulose-degrading bacteria in shipworm gills"
1099:
1009:
4324:
Petersen JM, Kemper A, Gruber-Vodicka H, Cardini U, Van Der Geest M, Kleiner M, et al. (2017).
4259:
Van Der Heide T, Govers LL, De Fouw J, Olff H, Van Der Geest M, Van Katwijk MM, et al. (2012).
5602:"Extensive Core Microbiome in Drone-Captured Whale Blow Supports a Framework for Health Monitoring"
1770:"Coral-associated micro-organisms and their roles in promoting coral health and thwarting diseases"
739:
639:
634:
342:
4539:
McFall-Ngai, M. (2014) "Divining the essence of symbiosis: insights from the squid-vibrio model".
2986:
Seah BK, Antony CP, Huettel B, Zarzycki J, Schada von Borzyskowski L, Erb TJ, et al. (2019).
2107:"Marine Animal Microbiomes: Toward Understanding Host–Microbiome Interactions in a Changing Ocean"
1190:
1594:
5443:
Bik EM, Costello EK, Switzer AD, Callahan BJ, Holmes SP, Wells RS, et al. (February 2016).
2149:
McFall-Ngai M, Hadfield MG, Bosch TC, Carey HV, Domazet-Lošo T, Douglas AE, et al. (2013).
6157:
Cavalcanti GS, Shukla P, Morris M, Ribeiro B, Foley M, Doane MP, et al. (September 2018).
5892:
Geoghegan JL, Pirotta V, Harvey E, Smith A, Buchmann JP, Ostrowski M, et al. (June 2018).
3412:"Effects of epibiotic bacteria on leaf growth and epiphytes of the seagrass Posidonia oceanica"
1327:
1200:
567:
2401:
1425:
to more specifically target key microbial groups has been hindered by the lack of full length
18:
5986:
Pollock FJ, McMinds R, Smith S, Bourne DG, Willis BL, Medina M, et al. (November 2018).
5735:"The use of Unmanned Aerial Vehicles (UAVs) to sample the blow microbiome of small cetaceans"
5392:
Sehnal L, Brammer-Robbins E, Wormington AM, Blaha L, Bisesi J, Larkin I, et al. (2021).
5324:
Pirotta V, Smith A, Ostrowski M, Russell D, Jonsen ID, Grech A, et al. (December 2017).
4114:
Corzett CH, Elsherbini J, Chien DM, Hehemann JH, Henschel A, Preheim SP, et al. (2018).
3953:
Elshahawi SI, Trindade-Silva AE, Hanora A, Han AW, Flores MS, Vizzoni V, et al. (2013).
1225:
1059:
1013:
919:
907:
577:
378:
221:
38:
4423:
Daniels, C. and Breitbart, M. (2012) "Bacterial communities associated with the ctenophores
6274:
5999:
5858:
5805:
5792:
Raverty SA, Rhodes LD, Zabek E, Eshghi A, Cameron CE, Hanson MB, et al. (March 2017).
5746:
5699:
5566:
5456:
4272:
4217:
4068:
4013:
3907:
3855:
3795:
3748:
3637:
3566:
3502:"The Importance of Microbes in Animal Development: Lessons from the Squid-Vibrio Symbiosis"
3423:
3368:
3331:
3238:
3083:
2932:
2836:
2781:
2724:
2677:
2626:
2471:
2416:
2358:
2221:
2162:
1976:
Nyholm SV, McFall-Ngai M (2004). "The winnowing: Establishing the squid–vibrio symbiosis".
1939:
1886:
1485:
Relative abundance of bacterial classes identified in sampled whale blow, air and seawater.
1388:
1089:
855:
826:
751:
673:
660:
538:
398:
383:
240:
42:
4326:"Chemosynthetic symbionts of marine invertebrate animals are capable of nitrogen fixation"
3842:
Reed DC, Carlson CA, Halewood ER, Nelson JC, Harrer SL, Rassweiler A, et al. (2015).
3739:
Steinberg PD, De Nys R (2002). "Chemical Mediation of Colonization of Seaweed Surfaces1".
2056:"A Developing Symbiosis: Enabling Cross-Talk Between Ecologists and Microbiome Scientists"
939:
Along with more standard examples of nutritional symbioses in animals, recent advances in
8:
3453:"Lessons from simple marine models on the bacterial regulation of eukaryotic development"
2400:
Cardinale BJ, Duffy JE, Gonzalez A, Hooper DU, Perrings C, Venail P, et al. (2012).
1476:
1346:
1332:
1124:
974:
931:
830:
716:
594:
457:
245:
190:
180:
53:
6224:
The marine microbiome: an untapped source of biodiversity and biotechnological potential
6003:
5862:
5809:
5750:
5703:
5649:
Vendl C, Ferrari BC, Thomas T, Slavich E, Zhang E, Nelson T, et al. (August 2019).
5570:
5460:
4833:
4276:
4221:
4072:
4017:
3911:
3859:
3799:
3752:
3641:
3570:
3517:
3427:
3372:
3335:
3242:
3087:
2936:
2840:
2785:
2728:
2681:
2630:
2475:
2420:
2362:
2225:
2166:
1943:
1890:
6185:
6158:
6124:
6097:
6073:
6046:
6022:
5987:
5968:
5920:
5893:
5826:
5793:
5769:
5734:
5626:
5601:
5539:
5526:
5501:
5477:
5444:
5420:
5393:
5303:
4983:
4954:
4756:
4729:
4350:
4325:
4306:
4241:
4142:
4115:
4091:
4056:
4037:
3981:
3954:
3930:
3895:
3873:
3821:
3764:
3706:
3679:
3602:
3589:
3550:
3526:
3501:
3477:
3452:
3392:
3297:
3254:
3250:
3165:
3138:
3119:
3106:
3071:
3012:
2987:
2968:
2896:
2871:
2852:
2804:
2769:
2745:
2712:
2693:
2650:
2594:
2569:
2545:
2518:
2494:
2459:
2440:
2382:
2326:
2301:
2282:
2185:
2150:
2128:
2082:
2055:
2001:
1850:
1825:
1794:
1769:
1732:
1705:
1586:
1446:
1401:
1265:
1242:
1142:
1094:
1074:
925:
814:
587:
447:
251:
235:
61:
5733:
Centelleghe C, Carraro L, Gonzalvo J, Rosso M, Esposti E, Gili C, et al. (2020).
4527:
2300:
Cavicchioli R, Ripple WJ, Timmis KN, Azam F, Bakken LR, Baylis M, et al. (2019).
2257:
Azam F, Worden AZ (2004). "OCEANOGRAPHY: Microbes, Molecules, and Marine Ecosystems".
1907:
1874:
6253:
6243:
6228:
6190:
6129:
6078:
6027:
5960:
5925:
5874:
5870:
5831:
5774:
5715:
5711:
5672:
5631:
5582:
5578:
5543:
5531:
5482:
5425:
5374:
5299:
5108:
5083:
4988:
4761:
4715:
4444:
4355:
4298:
4245:
4233:
4147:
4096:
4029:
3986:
3935:
3813:
3760:
3711:
3655:
3594:
3531:
3482:
3384:
3289:
3211:
3206:
3189:
3170:
3111:
3052:
3017:
2960:
2901:
2809:
2770:"Genomic diversification of giant enteric symbionts reflects host dietary lifestyles"
2750:
2642:
2599:
2550:
2499:
2432:
2374:
2331:
2274:
2239:
2234:
2209:
2190:
2087:
2036:
1993:
1955:
1951:
1912:
1855:
1799:
1737:
1430:
996:
940:
357:
151:
5972:
5307:
4310:
3877:
3825:
3768:
3606:
3301:
3258:
3123:
2972:
2856:
2697:
2458:
Gould AL, Zhang V, Lamberti L, Jones EW, Obadia B, Korasidis N, et al. (2018).
2286:
2005:
1898:
6180:
6170:
6119:
6109:
6068:
6058:
6017:
6007:
5952:
5915:
5905:
5866:
5821:
5813:
5764:
5754:
5707:
5662:
5621:
5613:
5600:
Apprill A, Miller CA, Moore MJ, Durban JW, Fearnbach H, Barrett-Lennard LG (2017).
5574:
5521:
5513:
5472:
5464:
5415:
5405:
5366:
5337:
5295:
5266:
5236:
5209:
5183:
5154:
5129:
5104:
5079:
5050:
5024:
4978:
4970:
4910:
4884:
4858:
4829:
4804:
4751:
4741:
4711:
4685:
4655:
4629:
4603:
4574:
4548:
4523:
4494:
4469:
4440:
4407:
4381:
4345:
4337:
4288:
4280:
4225:
4188:
4178:
4137:
4127:
4086:
4076:
4041:
4021:
3976:
3966:
3925:
3915:
3863:
3803:
3756:
3701:
3691:
3645:
3584:
3574:
3521:
3513:
3472:
3464:
3431:
3396:
3376:
3339:
3281:
3246:
3201:
3160:
3150:
3101:
3091:
3044:
3007:
2999:
2950:
2940:
2891:
2883:
2844:
2799:
2789:
2740:
2732:
2685:
2654:
2634:
2589:
2581:
2540:
2530:
2489:
2479:
2444:
2424:
2386:
2366:
2321:
2313:
2266:
2229:
2180:
2170:
2132:
2118:
2077:
2067:
2028:
1985:
1947:
1902:
1894:
1845:
1837:
1789:
1781:
1727:
1717:
1660:
1600:
1530:
743:
depends on intracellular methanotrophic bacteria in its gills as a source of carbon
507:
452:
426:
317:
213:
185:
114:
6114:
5445:"Marine mammals harbor unique microbiotas shaped by and yet distinct from the sea"
3137:
Delannoy S, Chaves BD, Ison SA, Webb HE, Beutin L, Delaval J, et al. (2016).
1704:
Wilkins LG, Leray M, o'Dea A, Yuen B, Peixoto RS, Pereira TJ, et al. (2019).
1433:
was capable of reproducing the biologically-relevant diversity detected by deeper
5759:
4552:
3920:
3808:
3783:
1722:
1572:
1458:
1422:
1364:
1209:
1109:
1104:
1079:
944:
847:
722:
437:
421:
352:
146:
5270:
4974:
4341:
4167:"Utilization of kelp-derived carbon sources by kelp surface-associated bacteria"
2988:"Sulfur-Oxidizing Symbionts without Canonical Genes for Autotrophic CO2Fixation"
1535:
Reef-building corals are holobionts that include the coral itself (a eukaryotic
6205:
6144:
6012:
5817:
5028:
5003:
4940:
3726:
2768:
Ngugi DK, Miyake S, Cahill M, Vinu M, Hackmann TJ, Blom J, et al. (2017).
1752:
1706:"Host-associated microbiomes drive structure and function of marine ecosystems"
1674:
1547:
1544:
1510:
1414:
1392:
1252:
1231:
1069:
1054:
903:
879:
878:, one of the most well-known examples of a mutualistic symbiosis, in which the
866:(for example, half of more than 300 surveyed species were unable to synthesize
562:
442:
347:
6257:
6175:
5617:
5410:
4862:
4746:
4689:
3696:
3468:
3048:
2945:
2920:
2848:
2736:
2317:
6268:
6063:
5342:
5325:
4385:
3155:
2123:
2106:
2072:
2019:
Hammer TJ, Sanders JG, Fierer N (2019). "Not all animals need a microbiome".
1664:
1551:
1506:
1481:
1410:
1380:
1294:
851:
839:
759:
614:
609:
367:
332:
5667:
5650:
5517:
5370:
5240:
5187:
5158:
5133:
4808:
4659:
4284:
4081:
4025:
3971:
3650:
3621:
3579:
3285:
2794:
2484:
2270:
2210:"Marine sponges and their microbial symbionts: Love and other relationships"
2175:
2032:
1235:
cells (probed yellow using in situ hybridization) within the tentacles of a
6194:
6133:
6082:
6031:
5964:
5929:
5878:
5835:
5778:
5719:
5676:
5635:
5586:
5535:
5486:
5429:
5378:
5213:
4992:
4765:
4473:
4359:
4302:
4237:
4151:
4100:
4033:
3990:
3939:
3817:
3715:
3659:
3598:
3535:
3486:
3388:
3293:
3215:
3174:
3115:
3056:
3021:
2964:
2905:
2887:
2813:
2754:
2646:
2603:
2554:
2503:
2436:
2378:
2335:
2278:
2243:
2194:
2091:
2040:
1997:
1959:
1859:
1803:
1785:
1741:
1556:
1536:
1084:
983:
948:
943:
technology have led to the discovery of many endosymbiotic associations in
875:
843:
731:
599:
582:
502:
468:
156:
127:
3003:
2585:
1916:
1841:
4633:
1406:
1341:
1036:
1008:. The ciliate lives under anaerobic conditions and harbors endosymbiotic
952:
915:
895:
552:
281:
141:
136:
5468:
5054:
4952:
4914:
4607:
4132:
3380:
2955:
2638:
2428:
2370:
6143:
Modified text was copied from this source, which is available under a [
5285:
3680:"The sponge holobiont in a changing ocean: from microbes to ecosystems"
3549:
Shikuma NJ, Antoshechkin I, Medeiros JM, Pilhofer M, Newman DK (2016).
2535:
1989:
1463:
1260:
1218:
1214:
992:
969:
899:
891:
859:
783:
604:
557:
518:
497:
362:
309:
256:
122:
96:
73:
69:
46:
6240:
Marine Microbiome and Biogeochemical Cycles in Marine Productive Areas
6204:
Modified text was copied from this source, which is available under a
5988:"Coral-associated bacteria demonstrate phylosymbiosis and cophylogeny"
5910:
5002:
Modified text was copied from this source, which is available under a
4953:
b Goldsmith D, a Pratte Z, a Kellogg C, e Snader S, h Sharp K (2019).
4939:
Modified text was copied from this source, which is available under a
4888:
4498:
4411:
3868:
3843:
3725:
Modified text was copied from this source, which is available under a
3436:
3411:
3344:
3319:
3096:
2876:
Philosophical Transactions of the Royal Society B: Biological Sciences
2689:
2151:"Animals in a bacterial world, a new imperative for the life sciences"
1751:
Modified text was copied from this source, which is available under a
1673:
Modified text was copied from this source, which is available under a
4578:
4261:"A Three-Stage Symbiosis Forms the Foundation of Seagrass Ecosystems"
4229:
4193:
4183:
4166:
1270:
1017:
995:, which are low-value cellular waste products from their hosts, into
955:
911:
871:
867:
863:
763:
755:
543:
413:
322:
276:
29:
5391:
2302:"Scientists' warning to humanity: Microorganisms and climate change"
754:
that inhabits the world's oceans, it would be challenging to find a
5956:
4323:
3548:
2517:
Seemann J, Yingst A, Stuart-Smith RD, Edgar GJ, Altieri AH (2018).
1540:
1426:
1316:
887:
822:
327:
104:
4959:
microbiome is reflected across different sequencing methodologies"
5848:
1518:
1501:
1000:
988:
978:
963:
883:
835:
779:
513:
4003:
3952:
3784:"Mini-review: Inhibition of biofouling by marine microorganisms"
3619:
3409:
5732:
5145:
Colman, A.S. (2015) "Sponge symbionts and the marine P cycle".
1451:
818:
775:
489:
65:
4728:
Sharp KH, Pratte ZA, Kerwin AH, Rotjan RD, Stewart FJ (2017).
4258:
4164:
4113:
3320:"Chemical interactions between marine macroalgae and bacteria"
2567:
2516:
1185:
6200:
6156:
6139:
6096:
Ugarelli K, Chakrabarti S, Laas P, Stingl U (December 2017).
6095:
5326:"An economical custom-built drone for assessing whale health"
4998:
4935:
3721:
3678:
Pita L, Rix L, Slaby BM, Franke A, Hentschel U (March 2018).
3555:
elegansis mediated by a bacterial inducer and MAPK signaling"
3271:
2616:
2148:
1747:
1669:
1514:
1375:
1264:
colonization and infection by potential microbial pathogens.
1259:
Some host-associated microbes produce compounds that prevent
959:
870:), and their productivity depends on provisioning from their
771:
77:
5323:
5015:
Bell, J.J. (2008) "The functional roles of marine sponges".
2711:
Baker DM, Freeman CJ, Wong JC, Fogel ML, Knowlton N (2018).
2399:
804:
Possible dynamics in symbiosis with aquatic ciliates as host
5894:"Virological Sampling of Inaccessible Wildlife with Drones"
5891:
2299:
1767:
725:
depend on symbiotic bacteria in their midgut for sustenance
6044:
5689:
5556:
5442:
5281:
5279:
3620:
Krediet CJ, Carpinone EM, Ritchie KB, Teplitski M (2013).
3188:
Wheeler GL, Tait K, Taylor A, Brownlee C, Joint I (2006).
5985:
5648:
3841:
3187:
2985:
2713:"Climate change promotes parasitism in a coral symbiosis"
1703:
1289:
dissolved organic carbon released by reef organisms into
5791:
5599:
5500:
Bai S, Zhang P, Lin M, Lin W, Yang Z, Li S (May 2021).
5276:
4727:
4165:
Bengtsson MM, Sjøtun K, Storesund JE, Øvreås J (2011).
3410:
Celdrán D, Espinosa E, Sánchez-Amat A, Marín A (2012).
2457:
1929:
1774:
Proceedings of the Royal Society B: Biological Sciences
1496:
communities vary between healthy and sick individuals''
793:
6206:
Creative Commons Attribution 4.0 International License
6147:
Creative Commons Attribution 4.0 International License
5356:
5004:
Creative Commons Attribution 4.0 International License
4941:
Creative Commons Attribution 4.0 International License
3781:
3727:
Creative Commons Attribution 4.0 International License
3358:
3317:
3136:
1753:
Creative Commons Attribution 4.0 International License
1675:
Creative Commons Attribution 4.0 International License
3893:
3069:
2767:
2710:
2568:
o'Brien PA, Webster NS, Miller DJ, Bourne DG (2019).
1872:
1768:
Krediet CJ, Ritchie KB, Paul VJ, Teplitski M (2013).
6045:
Thompson JR, Rivera HE, Closek CJ, Medina M (2014).
4207:
2348:
1023:
32:
or a breakdown of the relationship and interactions.
2018:
1363:Phylogenetic tree representing bacterial OTUs from
5259:Journal of Experimental Marine Biology and Ecology
3677:
2918:
2144:
2142:
2053:
1975:
1315:Relationships between corals and their microbial
6266:
6051:Frontiers in Cellular and Infection Microbiology
5726:
3228:
3034:
2667:
758:that does not live in close relationship with a
6238:
6150:
5319:
5317:
5147:Proceedings of the National Academy of Sciences
5122:Proceedings of the National Academy of Sciences
4797:Proceedings of the National Academy of Sciences
4648:Proceedings of the National Academy of Sciences
4061:Proceedings of the National Academy of Sciences
3959:Proceedings of the National Academy of Sciences
3889:
3887:
3738:
3559:Proceedings of the National Academy of Sciences
2869:
2774:Proceedings of the National Academy of Sciences
2464:Proceedings of the National Academy of Sciences
2155:Proceedings of the National Academy of Sciences
2139:
947:(a protist is a general term to refer to a non-
5942:
5885:
5499:
4516:Part A: Molecular & Integrative Physiology
3318:Goecke F, Labes A, Wiese J, Imhoff JF (2010).
2402:"Biodiversity loss and its impact on humanity"
1196:Some marine animals and associated microbiomes
6221:Stal, L. J. and Cretoiu, M. S. (Eds.) (2016)
6089:
6038:
5936:
4946:
3313:
3311:
2207:
1823:
1819:
1817:
1815:
1813:
1763:
1761:
1699:
1697:
1695:
1693:
1691:
1689:
1687:
1685:
1683:
1647:
1645:
1643:
1641:
1639:
1637:
1635:
1633:
1162:
882:alga Symbiodiniaceae supplies the coral with
681:
6145:https://creativecommons.org/licenses/by/4.0/
5785:
5683:
5593:
5314:
4589:
4587:
3884:
2872:"Endosymbiotic associations within protists"
2826:
2460:"Microbiome interactions shape host fitness"
1971:
1969:
6098:"The Seagrass Holobiont and Its Microbiome"
5385:
5350:
3837:
3835:
3673:
3671:
3669:
3499:
3450:
1873:Distel DL, Delong EF, Waterbury JB (1991).
1186:Biofouling and microbial community assembly
981:) nourish their ciliate hosts in the genus
5642:
4294:11370/23625acb-7ec0-4480-98d7-fad737d7d4fe
3782:Dobretsov S, Abed RM, Teplitski M (2013).
3308:
2256:
1810:
1758:
1680:
1630:
1616:Climate change and the rhodolith holobiont
1405:, the northern star coral, is a temperate
1169:
1155:
999:. Another example is the anaerobic marine
688:
674:
24:Marine animal host-microbiome relationship
6184:
6174:
6123:
6113:
6072:
6062:
6021:
6011:
5919:
5909:
5842:
5825:
5768:
5758:
5666:
5625:
5525:
5476:
5419:
5409:
5341:
4982:
4839:
4755:
4745:
4584:
4349:
4292:
4192:
4182:
4141:
4131:
4090:
4080:
4057:"Sponge symbionts and the marine P cycle"
3980:
3970:
3929:
3919:
3894:Vollmers J, Wiegand S, Kaster AK (2017).
3867:
3807:
3705:
3695:
3649:
3588:
3578:
3525:
3476:
3435:
3343:
3205:
3164:
3154:
3105:
3095:
3070:Huang Y, Callahan S, Hadfield MG (2012).
3011:
2954:
2944:
2895:
2803:
2793:
2744:
2593:
2544:
2534:
2493:
2483:
2325:
2233:
2184:
2174:
2122:
2081:
2071:
1966:
1906:
1849:
1793:
1731:
1721:
1277:
817:, such as many types of corals, deep-sea
52:In the oceans, it is challenging to find
5493:
5436:
3832:
3666:
3551:"Stepwise metamorphosis of the tubeworm
1480:
1457:
1358:
1309:
1189:
924:
797:
17:
4512:Comparative Biochemistry and Physiology
2919:Higgs ND, Newton J, Attrill MJ (2016).
2104:
6267:
4851:Applied Microbiology and Biotechnology
4622:Applied and Environmental Microbiology
4054:
2054:Tipton L, Darcy JL, Hynson NA (2019).
1879:Applied and Environmental Microbiology
1413:(algal symbionts), making it an ideal
6215:
5017:Estuarine, Coastal and Shelf Science
1524:
846:) and hydrothermal vents (e.g., the
794:Foundations of productive ecosystems
4834:10.1146/annurev-micro-102215-095440
3518:10.1146/annurev-micro-091313-103654
1462:Collecting a sample of blow from a
1221:nucleus (N) within the light organ.
13:
5851:Environmental Microbiology Reports
5692:Environmental Microbiology Reports
5559:Environmental Microbiology Reports
5229:Environmental Microbiology Reports
3251:10.1111/j.1529-8817.1980.tb03019.x
72:to more complex organisms such as
14:
6286:
1330:and the bioluminescent bacterium
1024:Reproduction and host development
874:bacteria. Reefs often consist of
6199:
6138:
5979:
5871:10.1111/j.1758-2229.2011.00306.x
5712:10.1111/j.1758-2229.2009.00080.x
5579:10.1111/j.1758-2229.2010.00213.x
5300:10.1111/j.1469-1795.2009.00326.x
5109:10.1111/j.1462-2920.2011.02661.x
5084:10.1111/j.1462-2920.2008.01582.x
4997:
4934:
4716:10.1111/j.1462-2920.2008.01728.x
4445:10.1111/j.1574-6941.2012.01409.x
3761:10.1046/j.1529-8817.2002.02042.x
3720:
3207:10.1111/j.1365-3040.2005.01440.x
2235:10.1111/j.1462-2920.2011.02460.x
1952:10.1111/j.1462-2920.2008.01728.x
1746:
1668:
1607:
1593:
1579:
1565:
1136:
1043:
1031:Part of a series of overviews on
730:
715:
655:
654:
103:
5550:
5247:
5219:
5193:
5164:
5139:
5114:
5089:
5060:
5034:
5009:
4931:United States Geological Survey
4920:
4894:
4868:
4814:
4789:
4772:
4721:
4695:
4665:
4639:
4613:
4558:
4533:
4504:
4479:
4450:
4417:
4391:
4366:
4317:
4252:
4201:
4158:
4107:
4048:
3997:
3946:
3775:
3732:
3613:
3542:
3493:
3457:Current Opinion in Microbiology
3444:
3403:
3352:
3265:
3222:
3181:
3130:
3063:
3028:
2979:
2912:
2870:Nowack EC, Melkonian M (2010).
2863:
2820:
2761:
2704:
2661:
2610:
2561:
2510:
2451:
2393:
2342:
2293:
2250:
2201:
2098:
1899:10.1128/AEM.57.8.2376-2382.1991
1466:using a helicopter drone
4955:"Stability of temperate coral
4487:Marine Ecology Progress Series
3416:Marine Ecology Progress Series
3324:Marine Ecology Progress Series
2670:Marine Ecology Progress Series
2208:Webster NS, Taylor MW (2012).
2047:
2012:
1923:
1866:
1229:and (D) a microscopy image of
1:
6242:. Frontiers Media S.A. 2020.
6115:10.3390/microorganisms5040081
4822:Annual Review of Microbiology
4528:10.1016/S1095-6433(00)00233-6
3506:Annual Review of Microbiology
1824:Webster NS, Thomas T (2016).
1623:
1387:marine animal symbiosis. The
1326:The relationship between the
1217:(MV) and in proximity to the
1213:cells associating with dense
922:and suppress coral recovery.
83:
5760:10.1371/journal.pone.0235537
4553:10.1371/journal.pbio.1001783
3921:10.1371/journal.pone.0169662
3809:10.1080/08927014.2013.776042
1723:10.1371/journal.pbio.3000533
1470:
1383:and physiology of the host.
1223:(C) the reef-building coral
573:Microbial population biology
7:
5330:Frontiers in Marine Science
5271:10.1016/j.jembe.2015.11.004
4975:10.3934/microbiol.2019.1.62
4342:10.1038/nmicrobiol.2016.195
3194:Plant, Cell and Environment
2306:Nature Reviews Microbiology
2111:Frontiers in Marine Science
1978:Nature Reviews Microbiology
1653:Frontiers in Marine Science
1305:
1284:Marine biogeochemical cycle
768:deep-sea hydrothermal vents
10:
6291:
6013:10.1038/s41467-018-07275-x
5818:10.1038/s41598-017-00457-5
5176:Environmental Microbiology
5097:Environmental Microbiology
5072:Environmental Microbiology
5029:10.1016/j.ecss.2008.05.002
4704:Environmental microbiology
3848:Limnology and Oceanography
3451:Woznica A, King N (2018).
2214:Environmental Microbiology
1932:Environmental Microbiology
1528:
1474:
1444:
1440:
1435:next-generation sequencing
1369:next-generation sequencing
1291:particulate organic carbon
1281:
977:found in association with
709:Deep-sea hydrothermal vent
702:Marine microbial symbiosis
699:
200:Marine microbial symbiosis
6176:10.1186/s12864-018-5064-4
5655:FEMS Microbiology Ecology
5618:10.1128/mSystems.00119-17
5411:10.3389/fmicb.2021.567408
5398:Frontiers in Microbiology
5359:FEMS Microbiology Letters
4863:10.1007/s00253-016-7777-0
4747:10.1186/s40168-017-0329-8
4690:10.1186/s12864-016-3293-y
4433:FEMS Microbiology Ecology
4374:Frontiers in Microbiology
4171:Aquatic Microbial Ecology
3697:10.1186/s40168-018-0428-1
3630:FEMS Microbiology Ecology
3469:10.1016/j.mib.2017.12.013
3143:Frontiers in Microbiology
3049:10.1016/j.tim.2009.09.001
2946:10.1016/j.cub.2016.10.034
2849:10.1007/s10021-009-9291-z
2737:10.1038/s41396-018-0046-8
2318:10.1038/s41579-019-0222-5
2060:Frontiers in Microbiology
2021:FEMS Microbiology Letters
1354:
1010:purple nonsulfur bacteria
41:associating with diverse
6064:10.3389/fcimb.2014.00176
5343:10.3389/fmars.2017.00425
4386:10.3389/fmicb.2017.00341
3156:10.3389/fmicb.2016.00001
2124:10.3389/fmars.2017.00222
2073:10.3389/fmicb.2019.00292
1665:10.3389/fmars.2017.00222
1293:that can be consumed by
821:, and hydrothermal vent
740:Bathymodiolus childressi
640:Earth Microbiome Project
635:Human Microbiome Project
394:Accessible carbohydrates
5945:The American Naturalist
5518:10.1111/1749-4877.12502
5241:10.1111/1758-2229.12430
5188:10.1111/1462-2920.12010
5159:10.1073/pnas.1502763112
5134:10.1073/pnas.1423768112
4809:10.1073/pnas.0804478105
4660:10.1073/pnas.1121198109
4285:10.1126/science.1219973
4120:Journal of Bacteriology
4082:10.1073/pnas.1502763112
4026:10.1126/science.1241981
3972:10.1073/pnas.1213892110
3651:10.1111/1574-6941.12064
3580:10.1073/pnas.1603142113
3500:McFall-Ngai MJ (2014).
3286:10.1126/science.1105486
2795:10.1073/pnas.1703070114
2485:10.1073/pnas.1809349115
2271:10.1126/science.1093892
2176:10.1073/pnas.1218525110
1826:"The Sponge Hologenome"
840:stilbonematid nematodes
60:within ecosystems with
5214:10.1038/ismej.2014.188
5172:Rhopaloeides odorabile
4474:10.1038/ismej.2013.156
3037:Trends in Microbiology
2888:10.1098/rstb.2009.0188
1786:10.1098/rspb.2012.2328
1486:
1467:
1372:
1328:Hawaiian bobtail squid
1320:
1278:Biogeochemical cycling
1256:
1201:Hawaiian bobtail squid
1143:Marine life portal
936:
898:, which seek homes on
811:
568:Biological dark matter
58:biogeochemical cycling
33:
5992:Nature Communications
5668:10.1093/femsec/fiz102
5449:Nature Communications
5371:10.1093/femsle/fnz134
5043:Nature Communications
3004:10.1128/mBio.01112-19
2586:10.1128/mBio.02241-18
2470:(51): E11951–E11960.
2033:10.1093/femsle/fnz117
1842:10.1128/mBio.00135-16
1484:
1461:
1362:
1313:
1226:Stylophora pistillata
1193:
1115:Biogeochemical cycles
1014:bacteriochlorophyll a
1005:Strombidium purpureum
928:
920:ecosystem disturbance
908:niche diversification
801:
737:The deepwater mussel
578:Microbial cooperation
39:marine microorganisms
21:
4634:10.1128/AEM.02776-15
3741:Journal of Phycology
3231:Journal of Phycology
1389:gammaproteobacterium
973:Kentron (a clade of
856:foundational species
827:primary productivity
752:biological diversity
539:Biomass partitioning
474:hologenome evolution
399:Flora (microbiology)
54:eukaryotic organisms
6004:2018NatCo...9.4921P
5951:(4 Suppl): S51–62.
5863:2012EnvMR...4...89L
5810:2017NatSR...7..394R
5751:2020PLoSO..1535537C
5704:2009EnvMR...1..555J
5571:2011EnvMR...3..223A
5506:Integrative Zoology
5469:10.1038/ncomms10516
5461:2016NatCo...710516B
5288:Animal Conservation
5055:10.1038/ncomms11870
4915:10.1038/nature21707
4803:(45): 17442–17446.
4674:Olavius algarvensis
4654:(19): E1173–E1182.
4608:10.1038/nature05192
4330:Nature Microbiology
4277:2012Sci...336.1432V
4271:(6087): 1432–1434.
4222:2019Ecol..100E2798P
4133:10.1128/JB.00020-18
4073:2015PNAS..112.4191C
4018:2013Sci...342..108D
3912:2017PLoSO..1269662V
3860:2015LimOc..60.1996R
3800:2013Biofo..29..423D
3753:2002JPcgy..38..621S
3642:2013FEMME..84..290K
3571:2016PNAS..11310097S
3565:(36): 10097–10102.
3428:2012MEPS..456...21C
3381:10.1038/nature14488
3373:2015Natur.522...98A
3336:2010MEPS..409..267G
3243:1980JPcgy..16..196P
3088:2012NatSR...2E.228H
2937:2016CBio...26.3393H
2841:2009Ecosy..12.1316H
2786:2017PNAS..114E7592N
2780:(36): E7592–E7601.
2729:2018ISMEJ..12..921B
2682:2018MEPS..600..245Q
2639:10.1038/nature04056
2631:2005Natur.438...90C
2476:2018PNAS..11511951G
2429:10.1038/nature11148
2421:2012Natur.486...59C
2371:10.1038/nature23886
2363:2017Natur.549..261D
2265:(5664): 1622–1624.
2226:2012EnvMi..14..335W
2167:2013PNAS..110.3229M
1944:2008EnvMi..10.3404R
1891:1991ApEnM..57.2376D
1477:Cetacean microbiome
1347:Olavius algarvensis
1340:The gutless marine
1333:Aliivibrio fischeri
975:Gammaproteobacteria
932:Olavius algarvensis
815:Ecosystem engineers
756:eukaryotic organism
595:Metatranscriptomics
389:Initial acquisition
384:Microbial community
91:Part of a series on
6216:Further references
5798:Scientific Reports
5068:Aplysina aerophoba
4957:Astrangia poculata
4877:Scientific Reports
4458:Ciona intestinalis
4400:Scientific reports
4055:Colman AS (2015).
3076:Scientific Reports
2536:10.7717/peerj.4455
2105:Apprill A (2017).
1990:10.1038/nrmicro957
1587:Seagrass holobiont
1487:
1468:
1447:Sponge microbiomes
1402:Astrangia poculata
1373:
1321:
1266:Epiphytic bacteria
1257:
1243:Atlantic killifish
1100:Primary production
1012:that contain both
937:
854:). Moreover, many
812:
173:Marine microbiomes
34:
6249:978-2-88963-276-3
5911:10.3390/v10060300
5255:Xestospongia muta
5182:(12): 3232–3246.
5153:(14): 4191–4192.
5128:(14): 4381–4386.
5078:(11): 2942–2955.
4963:AIMS Microbiology
4909:(7645): 373–377.
4889:10.1038/srep40579
4857:(19): 8315–8324.
4710:(12): 3404–3416.
4602:(7114): 950–955.
4573:(6835): 298–302.
4499:10.3354/meps11034
4425:Mnemiopsis leidyi
4412:10.1038/srep19324
4067:(14): 4191–4192.
4012:(6154): 108–110.
3869:10.1002/lno.10154
3626:marcescensPDL100"
3437:10.3354/meps09672
3345:10.3354/meps08607
3097:10.1038/srep00228
2931:(24): 3393–3398.
2882:(1541): 699–712.
2690:10.3354/meps12652
2357:(7671): 261–264.
1938:(12): 3404–3416.
1525:Marine holobionts
1431:Sanger sequencing
1179:
1178:
941:genome sequencing
929:The gutless worm
910:among coral reef
809:
760:microbial partner
698:
697:
288:Built environment
270:Other microbiomes
214:Human microbiomes
115:Plant microbiomes
6282:
6261:
6209:
6203:
6198:
6188:
6178:
6154:
6148:
6142:
6137:
6127:
6117:
6093:
6087:
6086:
6076:
6066:
6042:
6036:
6035:
6025:
6015:
5983:
5977:
5976:
5940:
5934:
5933:
5923:
5913:
5889:
5883:
5882:
5846:
5840:
5839:
5829:
5789:
5783:
5782:
5772:
5762:
5730:
5724:
5723:
5687:
5681:
5680:
5670:
5646:
5640:
5639:
5629:
5597:
5591:
5590:
5554:
5548:
5547:
5529:
5497:
5491:
5490:
5480:
5440:
5434:
5433:
5423:
5413:
5389:
5383:
5382:
5354:
5348:
5347:
5345:
5321:
5312:
5311:
5283:
5274:
5251:
5245:
5223:
5217:
5202:The ISME Journal
5197:
5191:
5168:
5162:
5143:
5137:
5118:
5112:
5093:
5087:
5064:
5058:
5038:
5032:
5013:
5007:
5001:
4996:
4986:
4950:
4944:
4938:
4933:, 6 March 2019.
4924:
4918:
4898:
4892:
4872:
4866:
4843:
4837:
4818:
4812:
4793:
4787:
4776:
4770:
4769:
4759:
4749:
4725:
4719:
4699:
4693:
4669:
4663:
4643:
4637:
4617:
4611:
4591:
4582:
4579:10.1038/35077067
4562:
4556:
4537:
4531:
4508:
4502:
4483:
4477:
4462:The ISME Journal
4454:
4448:
4421:
4415:
4395:
4389:
4370:
4364:
4363:
4353:
4321:
4315:
4314:
4296:
4256:
4250:
4249:
4230:10.1002/ecy.2798
4205:
4199:
4198:
4196:
4186:
4184:10.3354/ame01477
4162:
4156:
4155:
4145:
4135:
4111:
4105:
4104:
4094:
4084:
4052:
4046:
4045:
4001:
3995:
3994:
3984:
3974:
3965:(4): E295–E304.
3950:
3944:
3943:
3933:
3923:
3891:
3882:
3881:
3871:
3854:(6): 1996–2008.
3839:
3830:
3829:
3811:
3779:
3773:
3772:
3736:
3730:
3724:
3719:
3709:
3699:
3675:
3664:
3663:
3653:
3617:
3611:
3610:
3592:
3582:
3546:
3540:
3539:
3529:
3497:
3491:
3490:
3480:
3448:
3442:
3441:
3439:
3407:
3401:
3400:
3367:(7554): 98–101.
3356:
3350:
3349:
3347:
3315:
3306:
3305:
3269:
3263:
3262:
3226:
3220:
3219:
3209:
3185:
3179:
3178:
3168:
3158:
3134:
3128:
3127:
3109:
3099:
3067:
3061:
3060:
3032:
3026:
3025:
3015:
2983:
2977:
2976:
2958:
2948:
2916:
2910:
2909:
2899:
2867:
2861:
2860:
2835:(8): 1316–1328.
2824:
2818:
2817:
2807:
2797:
2765:
2759:
2758:
2748:
2717:The ISME Journal
2708:
2702:
2701:
2665:
2659:
2658:
2614:
2608:
2607:
2597:
2565:
2559:
2558:
2548:
2538:
2514:
2508:
2507:
2497:
2487:
2455:
2449:
2448:
2406:
2397:
2391:
2390:
2346:
2340:
2339:
2329:
2297:
2291:
2290:
2254:
2248:
2247:
2237:
2205:
2199:
2198:
2188:
2178:
2161:(9): 3229–3236.
2146:
2137:
2136:
2126:
2102:
2096:
2095:
2085:
2075:
2051:
2045:
2044:
2016:
2010:
2009:
1973:
1964:
1963:
1927:
1921:
1920:
1910:
1885:(8): 2376–2382.
1870:
1864:
1863:
1853:
1836:(2): e00135-16.
1821:
1808:
1807:
1797:
1765:
1756:
1750:
1745:
1735:
1725:
1716:(11): e3000533.
1701:
1678:
1672:
1649:
1611:
1601:Sponge holobiont
1597:
1583:
1569:
1531:Marine holobiont
1511:organic material
1171:
1164:
1157:
1141:
1140:
1139:
1047:
1046:
1028:
1027:
852:deep-sea mussels
825:, contribute to
807:
750:Within the vast
734:
723:Giant tube worms
719:
690:
683:
676:
663:
658:
657:
427:Marine holobiont
227:Fecal transplant
107:
88:
87:
6290:
6289:
6285:
6284:
6283:
6281:
6280:
6279:
6265:
6264:
6250:
6218:
6213:
6212:
6155:
6151:
6094:
6090:
6043:
6039:
5984:
5980:
5941:
5937:
5890:
5886:
5847:
5843:
5790:
5786:
5745:(7): e0235537.
5731:
5727:
5688:
5684:
5647:
5643:
5598:
5594:
5555:
5551:
5498:
5494:
5441:
5437:
5390:
5386:
5355:
5351:
5322:
5315:
5284:
5277:
5252:
5248:
5224:
5220:
5198:
5194:
5169:
5165:
5144:
5140:
5119:
5115:
5094:
5090:
5065:
5061:
5039:
5035:
5014:
5010:
4951:
4947:
4925:
4921:
4899:
4895:
4873:
4869:
4844:
4840:
4819:
4815:
4794:
4790:
4780:Pacific Science
4777:
4773:
4726:
4722:
4700:
4696:
4670:
4666:
4644:
4640:
4618:
4614:
4592:
4585:
4563:
4559:
4547:(2): e1001783.
4538:
4534:
4509:
4505:
4484:
4480:
4455:
4451:
4422:
4418:
4396:
4392:
4371:
4367:
4322:
4318:
4257:
4253:
4206:
4202:
4163:
4159:
4112:
4108:
4053:
4049:
4002:
3998:
3951:
3947:
3906:(1): e0169662.
3892:
3885:
3840:
3833:
3780:
3776:
3737:
3733:
3676:
3667:
3618:
3614:
3547:
3543:
3498:
3494:
3449:
3445:
3408:
3404:
3357:
3353:
3316:
3309:
3270:
3266:
3227:
3223:
3186:
3182:
3135:
3131:
3068:
3064:
3043:(12): 563–569.
3033:
3029:
2984:
2980:
2925:Current Biology
2917:
2913:
2868:
2864:
2825:
2821:
2766:
2762:
2709:
2705:
2666:
2662:
2625:(7064): 90–93.
2615:
2611:
2566:
2562:
2515:
2511:
2456:
2452:
2415:(7401): 59–67.
2404:
2398:
2394:
2347:
2343:
2298:
2294:
2255:
2251:
2206:
2202:
2147:
2140:
2103:
2099:
2052:
2048:
2017:
2013:
1974:
1967:
1928:
1924:
1871:
1867:
1822:
1811:
1766:
1759:
1702:
1681:
1650:
1631:
1626:
1619:
1618:
1612:
1603:
1598:
1589:
1584:
1575:
1573:Coral holobiont
1570:
1548:dinoflagellates
1533:
1527:
1479:
1473:
1449:
1443:
1365:clone libraries
1357:
1319:
1308:
1286:
1280:
1250:
1240:
1222:
1210:Vibrio fischeri
1198:
1188:
1175:
1137:
1135:
1044:
1026:
945:marine protists
904:polysaccharides
848:giant tube worm
810:
806:
796:
748:
747:
746:
745:
744:
735:
727:
726:
720:
711:
710:
704:
694:
653:
646:
645:
644:
629:
621:
620:
619:
548:
533:
525:
524:
523:
510:
492:
482:
481:
480:
464:
431:
422:Plant holobiont
416:
406:
405:
404:
403:
374:
312:
302:
301:
300:
284:
271:
263:
262:
261:
248:
231:
216:
206:
205:
204:
195:
175:
165:
164:
163:
152:soil microbiome
147:root microbiome
132:
117:
86:
26:
12:
11:
5:
6288:
6278:
6277:
6263:
6262:
6248:
6236:
6217:
6214:
6211:
6210:
6149:
6102:Microorganisms
6088:
6037:
5978:
5957:10.1086/378684
5935:
5884:
5841:
5784:
5725:
5682:
5641:
5592:
5565:(2): 223–232.
5549:
5512:(3): 324–335.
5492:
5435:
5384:
5349:
5313:
5294:(2): 217–225.
5275:
5246:
5235:(4): 536–544.
5218:
5208:(4): 894–908.
5192:
5163:
5138:
5113:
5103:(4): 909_923.
5088:
5059:
5033:
5023:(3): 341–353.
5008:
4945:
4919:
4893:
4867:
4847:Endozoicomonas
4838:
4813:
4788:
4771:
4720:
4694:
4664:
4638:
4612:
4583:
4557:
4532:
4522:(4): 471–480.
4503:
4478:
4468:(2): 309–320.
4449:
4416:
4390:
4365:
4316:
4251:
4216:(10): e02798.
4200:
4177:(2): 191–199.
4157:
4106:
4047:
3996:
3945:
3883:
3831:
3794:(4): 423–441.
3774:
3747:(4): 621–629.
3731:
3665:
3636:(2): 290–301.
3612:
3541:
3492:
3443:
3402:
3351:
3307:
3280:(5715): 1598.
3264:
3237:(2): 196–201.
3221:
3200:(4): 608–618.
3180:
3129:
3062:
3027:
2978:
2911:
2862:
2819:
2760:
2723:(3): 921–930.
2703:
2660:
2609:
2560:
2509:
2450:
2392:
2341:
2312:(9): 569–586.
2292:
2249:
2220:(2): 335–346.
2200:
2138:
2097:
2046:
2011:
1984:(8): 632–642.
1965:
1922:
1865:
1809:
1757:
1679:
1628:
1627:
1625:
1622:
1621:
1620:
1614:
1613:
1606:
1604:
1599:
1592:
1590:
1585:
1578:
1576:
1571:
1564:
1545:photosynthetic
1529:Main article:
1526:
1523:
1507:microorganisms
1475:Main article:
1472:
1469:
1445:Main article:
1442:
1439:
1415:model organism
1393:Endozoicomonas
1356:
1353:
1314:
1307:
1304:
1279:
1276:
1253:humpback whale
1232:Endozoicomonas
1194:
1187:
1184:
1177:
1176:
1174:
1173:
1166:
1159:
1151:
1148:
1147:
1146:
1145:
1130:
1129:
1128:
1127:
1122:
1117:
1112:
1107:
1102:
1097:
1092:
1087:
1082:
1077:
1072:
1067:
1062:
1060:Microorganisms
1057:
1049:
1048:
1040:
1039:
1033:
1032:
1025:
1022:
951:collection of
880:dinoflagellate
842:, and gutless
802:
795:
792:
736:
729:
728:
721:
714:
713:
712:
708:
707:
706:
705:
696:
695:
693:
692:
685:
678:
670:
667:
666:
665:
664:
648:
647:
643:
642:
637:
631:
630:
627:
626:
623:
622:
618:
617:
612:
607:
602:
597:
592:
591:
590:
580:
575:
570:
565:
563:Quorum sensing
560:
555:
549:
547:
546:
541:
535:
534:
531:
530:
527:
526:
522:
521:
516:
511:
505:
500:
494:
493:
488:
487:
484:
483:
479:
478:
477:
476:
465:
463:
462:
461:
460:
455:
450:
445:
440:
432:
430:
429:
424:
418:
417:
412:
411:
408:
407:
402:
401:
396:
391:
386:
381:
375:
373:
372:
371:
370:
365:
360:
355:
350:
339:
338:
337:
336:
335:
330:
325:
314:
313:
308:
307:
304:
303:
299:
298:
290:
285:
279:
273:
272:
269:
268:
265:
264:
260:
259:
254:
249:
243:
238:
236:Gut–brain axis
232:
230:
229:
224:
218:
217:
212:
211:
208:
207:
203:
202:
196:
194:
193:
188:
183:
177:
176:
171:
170:
167:
166:
162:
161:
160:
159:
154:
149:
144:
133:
131:
130:
125:
119:
118:
113:
112:
109:
108:
100:
99:
93:
92:
85:
82:
22:
9:
6:
4:
3:
2:
6287:
6276:
6273:
6272:
6270:
6259:
6255:
6251:
6245:
6241:
6237:
6234:
6233:9783319330006
6230:
6226:
6225:
6220:
6219:
6207:
6202:
6196:
6192:
6187:
6182:
6177:
6172:
6168:
6164:
6160:
6153:
6146:
6141:
6135:
6131:
6126:
6121:
6116:
6111:
6107:
6103:
6099:
6092:
6084:
6080:
6075:
6070:
6065:
6060:
6056:
6052:
6048:
6041:
6033:
6029:
6024:
6019:
6014:
6009:
6005:
6001:
5997:
5993:
5989:
5982:
5974:
5970:
5966:
5962:
5958:
5954:
5950:
5946:
5939:
5931:
5927:
5922:
5917:
5912:
5907:
5903:
5899:
5895:
5888:
5880:
5876:
5872:
5868:
5864:
5860:
5856:
5852:
5845:
5837:
5833:
5828:
5823:
5819:
5815:
5811:
5807:
5803:
5799:
5795:
5788:
5780:
5776:
5771:
5766:
5761:
5756:
5752:
5748:
5744:
5740:
5736:
5729:
5721:
5717:
5713:
5709:
5705:
5701:
5698:(6): 555–62.
5697:
5693:
5686:
5678:
5674:
5669:
5664:
5660:
5656:
5652:
5645:
5637:
5633:
5628:
5623:
5619:
5615:
5611:
5607:
5603:
5596:
5588:
5584:
5580:
5576:
5572:
5568:
5564:
5560:
5553:
5545:
5541:
5537:
5533:
5528:
5523:
5519:
5515:
5511:
5507:
5503:
5496:
5488:
5484:
5479:
5474:
5470:
5466:
5462:
5458:
5454:
5450:
5446:
5439:
5431:
5427:
5422:
5417:
5412:
5407:
5403:
5399:
5395:
5388:
5380:
5376:
5372:
5368:
5364:
5360:
5353:
5344:
5339:
5335:
5331:
5327:
5320:
5318:
5309:
5305:
5301:
5297:
5293:
5289:
5282:
5280:
5272:
5268:
5264:
5260:
5256:
5250:
5244:
5242:
5238:
5234:
5228:
5222:
5215:
5211:
5207:
5203:
5196:
5189:
5185:
5181:
5177:
5173:
5167:
5160:
5156:
5152:
5148:
5142:
5135:
5131:
5127:
5123:
5117:
5110:
5106:
5102:
5098:
5092:
5085:
5081:
5077:
5073:
5069:
5063:
5056:
5052:
5048:
5044:
5037:
5030:
5026:
5022:
5018:
5012:
5005:
5000:
4994:
4990:
4985:
4980:
4976:
4972:
4968:
4964:
4960:
4958:
4949:
4942:
4937:
4932:
4928:
4923:
4916:
4912:
4908:
4904:
4897:
4890:
4886:
4882:
4878:
4871:
4864:
4860:
4856:
4852:
4848:
4842:
4835:
4831:
4827:
4823:
4817:
4810:
4806:
4802:
4798:
4792:
4786:(3): 313–324.
4785:
4781:
4775:
4767:
4763:
4758:
4753:
4748:
4743:
4739:
4735:
4731:
4724:
4717:
4713:
4709:
4705:
4698:
4691:
4687:
4683:
4679:
4675:
4668:
4661:
4657:
4653:
4649:
4642:
4635:
4631:
4627:
4623:
4616:
4609:
4605:
4601:
4597:
4590:
4588:
4580:
4576:
4572:
4568:
4561:
4554:
4550:
4546:
4542:
4536:
4529:
4525:
4521:
4517:
4513:
4507:
4500:
4496:
4492:
4488:
4482:
4475:
4471:
4467:
4463:
4459:
4453:
4446:
4442:
4439:(1): 90–101.
4438:
4434:
4430:
4426:
4420:
4413:
4409:
4405:
4401:
4394:
4387:
4383:
4379:
4375:
4369:
4361:
4357:
4352:
4347:
4343:
4339:
4335:
4331:
4327:
4320:
4312:
4308:
4304:
4300:
4295:
4290:
4286:
4282:
4278:
4274:
4270:
4266:
4262:
4255:
4247:
4243:
4239:
4235:
4231:
4227:
4223:
4219:
4215:
4211:
4204:
4195:
4190:
4185:
4180:
4176:
4172:
4168:
4161:
4153:
4149:
4144:
4139:
4134:
4129:
4125:
4121:
4117:
4110:
4102:
4098:
4093:
4088:
4083:
4078:
4074:
4070:
4066:
4062:
4058:
4051:
4043:
4039:
4035:
4031:
4027:
4023:
4019:
4015:
4011:
4007:
4000:
3992:
3988:
3983:
3978:
3973:
3968:
3964:
3960:
3956:
3949:
3941:
3937:
3932:
3927:
3922:
3917:
3913:
3909:
3905:
3901:
3897:
3890:
3888:
3879:
3875:
3870:
3865:
3861:
3857:
3853:
3849:
3845:
3838:
3836:
3827:
3823:
3819:
3815:
3810:
3805:
3801:
3797:
3793:
3789:
3785:
3778:
3770:
3766:
3762:
3758:
3754:
3750:
3746:
3742:
3735:
3728:
3723:
3717:
3713:
3708:
3703:
3698:
3693:
3689:
3685:
3681:
3674:
3672:
3670:
3661:
3657:
3652:
3647:
3643:
3639:
3635:
3631:
3627:
3625:
3616:
3608:
3604:
3600:
3596:
3591:
3586:
3581:
3576:
3572:
3568:
3564:
3560:
3556:
3554:
3545:
3537:
3533:
3528:
3523:
3519:
3515:
3511:
3507:
3503:
3496:
3488:
3484:
3479:
3474:
3470:
3466:
3462:
3458:
3454:
3447:
3438:
3433:
3429:
3425:
3421:
3417:
3413:
3406:
3398:
3394:
3390:
3386:
3382:
3378:
3374:
3370:
3366:
3362:
3355:
3346:
3341:
3337:
3333:
3329:
3325:
3321:
3314:
3312:
3303:
3299:
3295:
3291:
3287:
3283:
3279:
3275:
3268:
3260:
3256:
3252:
3248:
3244:
3240:
3236:
3232:
3225:
3217:
3213:
3208:
3203:
3199:
3195:
3191:
3184:
3176:
3172:
3167:
3162:
3157:
3152:
3148:
3144:
3140:
3133:
3125:
3121:
3117:
3113:
3108:
3103:
3098:
3093:
3089:
3085:
3081:
3077:
3073:
3066:
3058:
3054:
3050:
3046:
3042:
3038:
3031:
3023:
3019:
3014:
3009:
3005:
3001:
2997:
2993:
2989:
2982:
2974:
2970:
2966:
2962:
2957:
2952:
2947:
2942:
2938:
2934:
2930:
2926:
2922:
2915:
2907:
2903:
2898:
2893:
2889:
2885:
2881:
2877:
2873:
2866:
2858:
2854:
2850:
2846:
2842:
2838:
2834:
2830:
2823:
2815:
2811:
2806:
2801:
2796:
2791:
2787:
2783:
2779:
2775:
2771:
2764:
2756:
2752:
2747:
2742:
2738:
2734:
2730:
2726:
2722:
2718:
2714:
2707:
2699:
2695:
2691:
2687:
2683:
2679:
2675:
2671:
2664:
2656:
2652:
2648:
2644:
2640:
2636:
2632:
2628:
2624:
2620:
2613:
2605:
2601:
2596:
2591:
2587:
2583:
2579:
2575:
2571:
2564:
2556:
2552:
2547:
2542:
2537:
2532:
2528:
2524:
2520:
2513:
2505:
2501:
2496:
2491:
2486:
2481:
2477:
2473:
2469:
2465:
2461:
2454:
2446:
2442:
2438:
2434:
2430:
2426:
2422:
2418:
2414:
2410:
2403:
2396:
2388:
2384:
2380:
2376:
2372:
2368:
2364:
2360:
2356:
2352:
2345:
2337:
2333:
2328:
2323:
2319:
2315:
2311:
2307:
2303:
2296:
2288:
2284:
2280:
2276:
2272:
2268:
2264:
2260:
2253:
2245:
2241:
2236:
2231:
2227:
2223:
2219:
2215:
2211:
2204:
2196:
2192:
2187:
2182:
2177:
2172:
2168:
2164:
2160:
2156:
2152:
2145:
2143:
2134:
2130:
2125:
2120:
2116:
2112:
2108:
2101:
2093:
2089:
2084:
2079:
2074:
2069:
2065:
2061:
2057:
2050:
2042:
2038:
2034:
2030:
2026:
2022:
2015:
2007:
2003:
1999:
1995:
1991:
1987:
1983:
1979:
1972:
1970:
1961:
1957:
1953:
1949:
1945:
1941:
1937:
1933:
1926:
1918:
1914:
1909:
1904:
1900:
1896:
1892:
1888:
1884:
1880:
1876:
1869:
1861:
1857:
1852:
1847:
1843:
1839:
1835:
1831:
1827:
1820:
1818:
1816:
1814:
1805:
1801:
1796:
1791:
1787:
1783:
1779:
1775:
1771:
1764:
1762:
1754:
1749:
1743:
1739:
1734:
1729:
1724:
1719:
1715:
1711:
1707:
1700:
1698:
1696:
1694:
1692:
1690:
1688:
1686:
1684:
1676:
1671:
1666:
1662:
1658:
1654:
1648:
1646:
1644:
1642:
1640:
1638:
1636:
1634:
1629:
1617:
1610:
1605:
1602:
1596:
1591:
1588:
1582:
1577:
1574:
1568:
1563:
1562:
1561:
1559:
1558:
1553:
1552:zooxanthellae
1549:
1546:
1542:
1539:within class
1538:
1532:
1522:
1520:
1516:
1512:
1508:
1503:
1499:
1497:
1491:
1483:
1478:
1465:
1460:
1456:
1453:
1448:
1438:
1436:
1432:
1428:
1424:
1420:
1416:
1412:
1411:zooxanthellae
1408:
1404:
1403:
1398:
1395:
1394:
1390:
1384:
1382:
1381:calcification
1377:
1370:
1366:
1361:
1352:
1349:
1348:
1343:
1338:
1335:
1334:
1329:
1324:
1318:
1312:
1303:
1299:
1296:
1295:heterotrophic
1292:
1285:
1275:
1272:
1267:
1262:
1254:
1248:
1244:
1238:
1237:S. pistillata
1234:
1233:
1228:
1227:
1220:
1216:
1212:
1211:
1206:
1202:
1197:
1192:
1183:
1172:
1167:
1165:
1160:
1158:
1153:
1152:
1150:
1149:
1144:
1134:
1133:
1132:
1131:
1126:
1123:
1121:
1118:
1116:
1113:
1111:
1108:
1106:
1103:
1101:
1098:
1096:
1093:
1091:
1090:Invertebrates
1088:
1086:
1083:
1081:
1078:
1076:
1073:
1071:
1068:
1066:
1063:
1061:
1058:
1056:
1053:
1052:
1051:
1050:
1042:
1041:
1038:
1035:
1034:
1030:
1029:
1021:
1019:
1015:
1011:
1007:
1006:
1002:
998:
994:
990:
986:
985:
980:
976:
972:
971:
965:
961:
958:that are not
957:
954:
950:
946:
942:
934:
933:
927:
923:
921:
917:
916:Surgeonfishes
913:
909:
905:
901:
897:
893:
889:
885:
881:
877:
873:
869:
865:
861:
857:
853:
849:
845:
841:
837:
836:lucinid clams
832:
828:
824:
820:
816:
805:
800:
791:
787:
785:
781:
777:
773:
770:. Studies on
769:
765:
761:
757:
753:
742:
741:
733:
724:
718:
703:
691:
686:
684:
679:
677:
672:
671:
669:
668:
662:
652:
651:
650:
649:
641:
638:
636:
633:
632:
625:
624:
616:
615:Symbiogenesis
613:
611:
610:Superorganism
608:
606:
603:
601:
598:
596:
593:
589:
586:
585:
584:
581:
579:
576:
574:
571:
569:
566:
564:
561:
559:
556:
554:
551:
550:
545:
542:
540:
537:
536:
529:
528:
520:
517:
515:
512:
509:
506:
504:
501:
499:
496:
495:
491:
486:
485:
475:
472:
471:
470:
467:
466:
459:
456:
454:
451:
449:
446:
444:
441:
439:
436:
435:
434:
433:
428:
425:
423:
420:
419:
415:
410:
409:
400:
397:
395:
392:
390:
387:
385:
382:
380:
377:
376:
369:
366:
364:
361:
359:
356:
354:
351:
349:
346:
345:
344:
341:
340:
334:
333:rhizobacteria
331:
329:
326:
324:
321:
320:
319:
316:
315:
311:
306:
305:
297:
295:
291:
289:
286:
283:
280:
278:
275:
274:
267:
266:
258:
255:
253:
250:
247:
244:
242:
239:
237:
234:
233:
228:
225:
223:
220:
219:
215:
210:
209:
201:
198:
197:
192:
189:
187:
184:
182:
179:
178:
174:
169:
168:
158:
155:
153:
150:
148:
145:
143:
140:
139:
138:
135:
134:
129:
126:
124:
121:
120:
116:
111:
110:
106:
102:
101:
98:
95:
94:
90:
89:
81:
79:
75:
71:
67:
63:
59:
55:
50:
48:
44:
43:marine animal
40:
31:
25:
20:
16:
6239:
6223:
6166:
6163:BMC Genomics
6162:
6152:
6105:
6101:
6091:
6054:
6050:
6040:
5995:
5991:
5981:
5948:
5944:
5938:
5901:
5897:
5887:
5857:(1): 89–96.
5854:
5850:
5844:
5801:
5797:
5787:
5742:
5738:
5728:
5695:
5691:
5685:
5658:
5654:
5644:
5609:
5605:
5595:
5562:
5558:
5552:
5509:
5505:
5495:
5452:
5448:
5438:
5401:
5397:
5387:
5362:
5358:
5352:
5333:
5329:
5291:
5287:
5262:
5258:
5254:
5249:
5232:
5230:
5226:
5221:
5205:
5201:
5195:
5179:
5175:
5171:
5166:
5150:
5146:
5141:
5125:
5121:
5116:
5100:
5096:
5091:
5075:
5071:
5067:
5062:
5046:
5042:
5036:
5020:
5016:
5011:
4969:(1): 62–76.
4966:
4962:
4956:
4948:
4930:
4922:
4906:
4902:
4896:
4880:
4876:
4870:
4854:
4850:
4846:
4841:
4825:
4821:
4816:
4800:
4796:
4791:
4783:
4779:
4774:
4737:
4733:
4723:
4707:
4703:
4697:
4681:
4678:BMC Genomics
4677:
4673:
4667:
4651:
4647:
4641:
4628:(1): 62–70.
4625:
4621:
4615:
4599:
4595:
4570:
4566:
4560:
4544:
4541:PLoS Biology
4540:
4535:
4519:
4515:
4511:
4506:
4490:
4486:
4481:
4465:
4461:
4457:
4452:
4436:
4432:
4428:
4424:
4419:
4403:
4399:
4393:
4377:
4373:
4368:
4336:(1): 16195.
4333:
4329:
4319:
4268:
4264:
4254:
4213:
4209:
4203:
4174:
4170:
4160:
4123:
4119:
4109:
4064:
4060:
4050:
4009:
4005:
3999:
3962:
3958:
3948:
3903:
3899:
3851:
3847:
3791:
3787:
3777:
3744:
3740:
3734:
3687:
3683:
3633:
3629:
3623:
3615:
3562:
3558:
3552:
3544:
3509:
3505:
3495:
3460:
3456:
3446:
3419:
3415:
3405:
3364:
3360:
3354:
3327:
3323:
3277:
3273:
3267:
3234:
3230:
3224:
3197:
3193:
3183:
3146:
3142:
3132:
3079:
3075:
3065:
3040:
3036:
3030:
2995:
2991:
2981:
2956:10026.1/9129
2928:
2924:
2914:
2879:
2875:
2865:
2832:
2828:
2822:
2777:
2773:
2763:
2720:
2716:
2706:
2673:
2669:
2663:
2622:
2618:
2612:
2577:
2573:
2563:
2526:
2522:
2512:
2467:
2463:
2453:
2412:
2408:
2395:
2354:
2350:
2344:
2309:
2305:
2295:
2262:
2258:
2252:
2217:
2213:
2203:
2158:
2154:
2114:
2110:
2100:
2063:
2059:
2049:
2024:
2020:
2014:
1981:
1977:
1935:
1931:
1925:
1882:
1878:
1868:
1833:
1829:
1777:
1773:
1713:
1710:PLOS Biology
1709:
1656:
1652:
1557:Symbiodinium
1555:
1537:invertebrate
1534:
1513:, including
1500:
1495:
1492:
1488:
1450:
1400:
1399:
1391:
1385:
1374:
1345:
1339:
1331:
1325:
1322:
1300:
1287:
1258:
1236:
1230:
1224:
1208:
1195:
1180:
1125:Conservation
1120:Human impact
1064:
1003:
987:and recycle
984:Kentrophoros
982:
968:
949:monophyletic
938:
930:
876:stony corals
862:are vitamin
844:oligochaetes
813:
803:
788:
749:
738:
600:Metabolomics
583:Metagenomics
469:Hologenomics
293:
172:
157:spermosphere
128:Phyllosphere
51:
35:
23:
15:
6275:Microbiomes
5998:(1): 4921.
5049:(1): 1-12.
4828:: 317–340.
4493:: 209–223.
4429:Beroe ovata
3512:: 177–194.
3463:: 108–116.
3330:: 267–299.
2676:: 245–253.
1407:stony coral
1342:oligochaete
1110:Carbon pump
1095:Vertebrates
1075:Prokaryotes
1065:Microbiomes
1037:Marine life
953:unicellular
900:coral reefs
896:Reef fishes
892:amino acids
831:mutualistic
553:Gnotobiosis
282:Phycosphere
142:laimosphere
137:Rhizosphere
97:Microbiomes
74:sea squirts
70:ctenophores
6258:1291256407
6227:Springer.
6169:(1): 701.
5904:(6): 300.
5804:(1): 394.
5404:: 567408.
4740:(1): 120.
4734:Microbiome
4684:(1): 942.
3788:Biofouling
3684:Microbiome
2829:Ecosystems
1624:References
1464:blue whale
1282:See also:
1261:biofouling
1245:and (F) a
1219:epithelial
1215:microvilli
1203:and (B) a
1018:organelles
993:propionate
970:Candidatus
962:or in the
956:eukaryotes
912:herbivores
864:auxotrophs
860:macroalgae
858:of marine
784:microbiome
700:See also:
605:Pan-genome
558:Phytobiome
519:Virosphere
414:Holobionts
310:Microbiota
294:Drosophila
257:Necrobiome
222:Human milk
123:Endosphere
84:Background
47:microbiome
6108:(4): 81.
5544:226302293
5455:: 10516.
5265:: 11–18.
4883:: 40579.
4406:: 19324.
4246:195355739
4194:1956/4610
3690:(1): 46.
3553:Hydroides
3422:: 21–27.
2529:: e4455.
1502:Cetaceans
1471:Cetaceans
1317:symbionts
1271:tartrolon
872:epiphytic
868:cobalamin
823:tubeworms
764:symbioses
544:Dysbiosis
458:rhodolith
323:endophyte
277:Mycobiome
241:Placental
62:cascading
30:dysbiosis
6269:Category
6195:30249182
6134:29244764
6083:25621279
6032:30467310
5973:24127308
5965:14583857
5930:29865228
5879:23757234
5836:28341851
5779:32614926
5739:PLOS ONE
5720:23765934
5677:31260051
5636:29034331
5606:mSystems
5587:23761254
5536:33174288
5487:26839246
5430:33776947
5379:31210263
5308:86518859
4993:31384703
4766:28915923
4360:27775707
4311:27806510
4303:22700927
4238:31233610
4152:29632094
4101:25825737
4034:24092742
3991:23288898
3940:28099457
3900:PLOS ONE
3878:85962482
3826:34459128
3818:23574279
3769:83963124
3716:29523192
3660:23278392
3624:Serratia
3607:23501584
3599:27551098
3536:24995875
3487:29331767
3389:26017307
3302:28850526
3294:15761147
3259:85817449
3216:17080611
3175:26834723
3124:14731587
3116:22355742
3057:19828317
3022:31239380
2973:14401680
2965:27939312
2906:20124339
2857:42138428
2814:28835538
2755:29379177
2698:90469901
2647:16267554
2604:30723123
2555:29610704
2504:30510004
2437:22678280
2379:28869964
2336:31213707
2287:10101482
2279:15016987
2244:21443739
2195:23391737
2092:30842763
2041:31132110
2006:21583331
1998:15263898
1960:18764872
1860:27103626
1804:23363627
1780:(1755).
1742:31710600
1541:Anthozoa
1519:proteins
1427:16S rRNA
1306:Examples
1241:(E) the
1105:Food web
1080:Protists
1055:Habitats
979:ciliates
888:glycerol
780:mollusks
661:Category
628:Projects
508:Mangrove
448:seagrass
328:epiphyte
246:Salivary
191:Cetacean
181:Seagrass
6186:6154897
6125:5748590
6074:4286716
6057:: 176.
6023:6250698
6000:Bibcode
5921:6024715
5898:Viruses
5859:Bibcode
5827:5428453
5806:Bibcode
5770:7332044
5747:Bibcode
5700:Bibcode
5627:5634792
5567:Bibcode
5527:9292824
5478:4742810
5457:Bibcode
5421:7995652
5336:: 425.
4984:6646935
4757:5603060
4380:: 341.
4351:6872982
4273:Bibcode
4265:Science
4218:Bibcode
4210:Ecology
4143:6040190
4092:4394276
4069:Bibcode
4042:6720678
4014:Bibcode
4006:Science
3982:3557025
3931:5242441
3908:Bibcode
3856:Bibcode
3796:Bibcode
3749:Bibcode
3707:5845141
3638:Bibcode
3590:5018781
3567:Bibcode
3527:6281398
3478:6051772
3424:Bibcode
3397:4462055
3369:Bibcode
3332:Bibcode
3274:Science
3239:Bibcode
3166:4722105
3107:3260340
3084:Bibcode
3082:: 228.
3013:6593406
2933:Bibcode
2897:2817226
2837:Bibcode
2805:5594648
2782:Bibcode
2746:5864192
2725:Bibcode
2678:Bibcode
2655:4328049
2627:Bibcode
2595:6428750
2546:5878927
2495:6304949
2472:Bibcode
2445:4333166
2417:Bibcode
2387:4459856
2359:Bibcode
2327:7136171
2259:Science
2222:Bibcode
2186:3587249
2163:Bibcode
2133:9729436
2083:6391321
2066:: 292.
1940:Bibcode
1917:1722662
1887:Bibcode
1851:4850255
1795:3574386
1733:6874084
1659:: 222.
1550:called
1452:Sponges
1441:Sponges
1419:primers
1070:Viruses
1001:ciliate
997:biomass
989:acetate
964:Plantae
884:glucose
819:mussels
776:sponges
762:. Such
532:Related
514:Viriome
490:Viromes
368:vaginal
252:Uterine
66:sponges
6256:
6246:
6231:
6193:
6183:
6132:
6122:
6081:
6071:
6030:
6020:
5971:
5963:
5928:
5918:
5877:
5834:
5824:
5777:
5767:
5718:
5675:
5634:
5624:
5585:
5542:
5534:
5524:
5485:
5475:
5428:
5418:
5377:
5365:(11).
5306:
4991:
4981:
4903:Nature
4764:
4754:
4596:Nature
4567:Nature
4358:
4348:
4309:
4301:
4244:
4236:
4150:
4140:
4126:(15).
4099:
4089:
4040:
4032:
3989:
3979:
3938:
3928:
3876:
3824:
3816:
3767:
3714:
3704:
3658:
3605:
3597:
3587:
3534:
3524:
3485:
3475:
3395:
3387:
3361:Nature
3300:
3292:
3257:
3214:
3173:
3163:
3122:
3114:
3104:
3055:
3020:
3010:
2971:
2963:
2904:
2894:
2855:
2812:
2802:
2753:
2743:
2696:
2653:
2645:
2619:Nature
2602:
2592:
2553:
2543:
2502:
2492:
2443:
2435:
2409:Nature
2385:
2377:
2351:Nature
2334:
2324:
2285:
2277:
2242:
2193:
2183:
2131:
2090:
2080:
2039:
2027:(10).
2004:
1996:
1958:
1915:
1908:183578
1905:
1858:
1848:
1802:
1792:
1740:
1730:
1515:lipids
1423:probes
1376:Corals
1355:Corals
1251:(G) a
890:, and
778:, and
772:corals
659:
453:sponge
379:Marine
78:sharks
5969:S2CID
5661:(8).
5612:(5).
5540:S2CID
5304:S2CID
4307:S2CID
4242:S2CID
4038:S2CID
3874:S2CID
3822:S2CID
3765:S2CID
3603:S2CID
3393:S2CID
3298:S2CID
3255:S2CID
3149:: 1.
3120:S2CID
2998:(3).
2969:S2CID
2853:S2CID
2694:S2CID
2651:S2CID
2580:(1).
2523:PeerJ
2441:S2CID
2405:(PDF)
2383:S2CID
2283:S2CID
2129:S2CID
2002:S2CID
1344:worm
1239:host.
1085:Fungi
960:fungi
588:viral
503:Human
438:coral
343:Human
318:Plant
186:Coral
6254:OCLC
6244:ISBN
6229:ISBN
6191:PMID
6130:PMID
6079:PMID
6028:PMID
5961:PMID
5926:PMID
5875:PMID
5832:PMID
5775:PMID
5716:PMID
5673:PMID
5632:PMID
5583:PMID
5532:PMID
5483:PMID
5426:PMID
5375:PMID
4989:PMID
4762:PMID
4427:and
4356:PMID
4299:PMID
4234:PMID
4148:PMID
4097:PMID
4030:PMID
3987:PMID
3936:PMID
3814:PMID
3712:PMID
3656:PMID
3595:PMID
3532:PMID
3483:PMID
3385:PMID
3290:PMID
3212:PMID
3171:PMID
3112:PMID
3053:PMID
3018:PMID
2992:mBio
2961:PMID
2902:PMID
2810:PMID
2751:PMID
2643:PMID
2600:PMID
2574:mBio
2551:PMID
2500:PMID
2433:PMID
2375:PMID
2332:PMID
2275:PMID
2240:PMID
2191:PMID
2088:PMID
2037:PMID
1994:PMID
1956:PMID
1913:PMID
1856:PMID
1830:mBio
1800:PMID
1738:PMID
1509:and
1421:and
1367:and
1199:(A)
991:and
443:crab
363:skin
358:oral
353:lung
76:and
68:and
6181:PMC
6171:doi
6120:PMC
6110:doi
6069:PMC
6059:doi
6018:PMC
6008:doi
5953:doi
5949:162
5916:PMC
5906:doi
5867:doi
5822:PMC
5814:doi
5765:PMC
5755:doi
5708:doi
5663:doi
5622:PMC
5614:doi
5575:doi
5522:PMC
5514:doi
5473:PMC
5465:doi
5416:PMC
5406:doi
5367:doi
5363:366
5338:doi
5296:doi
5267:doi
5263:475
5257:".
5237:doi
5210:doi
5184:doi
5155:doi
5151:112
5130:doi
5126:112
5105:doi
5080:doi
5070:".
5051:doi
5025:doi
4979:PMC
4971:doi
4911:doi
4907:543
4885:doi
4859:doi
4855:100
4849:".
4830:doi
4805:doi
4801:105
4752:PMC
4742:doi
4712:doi
4686:doi
4676:".
4656:doi
4652:109
4630:doi
4604:doi
4600:443
4575:doi
4571:411
4549:doi
4524:doi
4520:126
4495:doi
4491:518
4470:doi
4460:".
4441:doi
4431:".
4408:doi
4382:doi
4346:PMC
4338:doi
4289:hdl
4281:doi
4269:336
4226:doi
4214:100
4189:hdl
4179:doi
4138:PMC
4128:doi
4124:200
4087:PMC
4077:doi
4065:112
4022:doi
4010:342
3977:PMC
3967:doi
3963:110
3926:PMC
3916:doi
3864:doi
3804:doi
3757:doi
3702:PMC
3692:doi
3646:doi
3585:PMC
3575:doi
3563:113
3522:PMC
3514:doi
3473:PMC
3465:doi
3432:doi
3420:456
3377:doi
3365:522
3340:doi
3328:409
3282:doi
3278:307
3247:doi
3202:doi
3161:PMC
3151:doi
3102:PMC
3092:doi
3045:doi
3008:PMC
3000:doi
2951:hdl
2941:doi
2892:PMC
2884:doi
2880:365
2845:doi
2800:PMC
2790:doi
2778:114
2741:PMC
2733:doi
2686:doi
2674:600
2635:doi
2623:438
2590:PMC
2582:doi
2541:PMC
2531:doi
2490:PMC
2480:doi
2468:115
2425:doi
2413:486
2367:doi
2355:549
2322:PMC
2314:doi
2267:doi
2263:303
2230:doi
2181:PMC
2171:doi
2159:110
2119:doi
2078:PMC
2068:doi
2029:doi
2025:366
1986:doi
1948:doi
1903:PMC
1895:doi
1846:PMC
1838:doi
1790:PMC
1782:doi
1778:280
1728:PMC
1718:doi
1661:doi
1543:),
1247:SEM
1207:of
1205:TEM
850:or
498:Bat
348:gut
296:gut
6271::
6252:.
6189:.
6179:.
6167:19
6165:.
6161:.
6128:.
6118:.
6104:.
6100:.
6077:.
6067:.
6053:.
6049:.
6026:.
6016:.
6006:.
5994:.
5990:.
5967:.
5959:.
5947:.
5924:.
5914:.
5902:10
5900:.
5896:.
5873:.
5865:.
5853:.
5830:.
5820:.
5812:.
5800:.
5796:.
5773:.
5763:.
5753:.
5743:15
5741:.
5737:.
5714:.
5706:.
5694:.
5671:.
5659:95
5657:.
5653:.
5630:.
5620:.
5608:.
5604:.
5581:.
5573:.
5561:.
5538:.
5530:.
5520:.
5510:16
5508:.
5504:.
5481:.
5471:.
5463:.
5451:.
5447:.
5424:.
5414:.
5402:12
5400:.
5396:.
5373:.
5361:.
5332:.
5328:.
5316:^
5302:.
5292:13
5290:.
5278:^
5261:,
5231:,
5227:.
5204:,
5180:14
5178:,
5174:.
5149:,
5124:,
5101:14
5099:,
5076:10
5074:,
5045:,
5021:79
5019:,
4987:.
4977:.
4965:.
4961:.
4929:,
4905:,
4879:,
4853:,
4826:70
4824:,
4799:,
4784:48
4782:,
4760:.
4750:.
4736:.
4732:.
4708:10
4706:,
4682:17
4680:,
4650:,
4626:82
4624:,
4598:,
4586:^
4569:,
4545:12
4543:,
4518:,
4514:,
4489:,
4464:,
4437:82
4435:,
4402:,
4376:,
4354:.
4344:.
4332:.
4328:.
4305:.
4297:.
4287:.
4279:.
4267:.
4263:.
4240:.
4232:.
4224:.
4212:.
4187:.
4175:62
4173:.
4169:.
4146:.
4136:.
4122:.
4118:.
4095:.
4085:.
4075:.
4063:.
4059:.
4036:.
4028:.
4020:.
4008:.
3985:.
3975:.
3961:.
3957:.
3934:.
3924:.
3914:.
3904:12
3902:.
3898:.
3886:^
3872:.
3862:.
3852:60
3850:.
3846:.
3834:^
3820:.
3812:.
3802:.
3792:29
3790:.
3786:.
3763:.
3755:.
3745:38
3743:.
3710:.
3700:.
3686:.
3682:.
3668:^
3654:.
3644:.
3634:84
3632:.
3628:.
3601:.
3593:.
3583:.
3573:.
3561:.
3557:.
3530:.
3520:.
3510:68
3508:.
3504:.
3481:.
3471:.
3461:43
3459:.
3455:.
3430:.
3418:.
3414:.
3391:.
3383:.
3375:.
3363:.
3338:.
3326:.
3322:.
3310:^
3296:.
3288:.
3276:.
3253:.
3245:.
3235:16
3233:.
3210:.
3198:29
3196:.
3192:.
3169:.
3159:.
3145:.
3141:.
3118:.
3110:.
3100:.
3090:.
3078:.
3074:.
3051:.
3041:17
3039:.
3016:.
3006:.
2996:10
2994:.
2990:.
2967:.
2959:.
2949:.
2939:.
2929:26
2927:.
2923:.
2900:.
2890:.
2878:.
2874:.
2851:.
2843:.
2833:12
2831:.
2808:.
2798:.
2788:.
2776:.
2772:.
2749:.
2739:.
2731:.
2721:12
2719:.
2715:.
2692:.
2684:.
2672:.
2649:.
2641:.
2633:.
2621:.
2598:.
2588:.
2578:10
2576:.
2572:.
2549:.
2539:.
2525:.
2521:.
2498:.
2488:.
2478:.
2466:.
2462:.
2439:.
2431:.
2423:.
2411:.
2407:.
2381:.
2373:.
2365:.
2353:.
2330:.
2320:.
2310:17
2308:.
2304:.
2281:.
2273:.
2261:.
2238:.
2228:.
2218:14
2216:.
2212:.
2189:.
2179:.
2169:.
2157:.
2153:.
2141:^
2127:.
2117:.
2113:.
2109:.
2086:.
2076:.
2064:10
2062:.
2058:.
2035:.
2023:.
2000:.
1992:.
1980:.
1968:^
1954:.
1946:.
1936:10
1934:.
1911:.
1901:.
1893:.
1883:57
1881:.
1877:.
1854:.
1844:.
1832:.
1828:.
1812:^
1798:.
1788:.
1776:.
1772:.
1760:^
1736:.
1726:.
1714:17
1712:.
1708:.
1682:^
1667:.
1655:,
1632:^
1517:,
1498:.
1020:.
914:.
886:,
838:,
774:,
80:.
6260:.
6235:.
6208:.
6197:.
6173::
6136:.
6112::
6106:5
6085:.
6061::
6055:4
6034:.
6010::
6002::
5996:9
5975:.
5955::
5932:.
5908::
5881:.
5869::
5861::
5855:4
5838:.
5816::
5808::
5802:7
5781:.
5757::
5749::
5722:.
5710::
5702::
5696:1
5679:.
5665::
5638:.
5616::
5610:2
5589:.
5577::
5569::
5563:3
5546:.
5516::
5489:.
5467::
5459::
5453:7
5432:.
5408::
5381:.
5369::
5346:.
5340::
5334:4
5310:.
5298::
5273:.
5269::
5243:.
5239::
5233:8
5216:.
5212::
5206:9
5190:.
5186::
5161:.
5157::
5136:.
5132::
5111:.
5107::
5086:.
5082::
5057:.
5053::
5047:7
5031:.
5027::
5006:.
4995:.
4973::
4967:5
4943:.
4917:.
4913::
4891:.
4887::
4881:7
4865:.
4861::
4836:.
4832::
4811:.
4807::
4768:.
4744::
4738:5
4718:.
4714::
4692:.
4688::
4662:.
4658::
4636:.
4632::
4610:.
4606::
4581:.
4577::
4555:.
4551::
4530:.
4526::
4501:.
4497::
4476:.
4472::
4466:8
4447:.
4443::
4414:.
4410::
4404:6
4388:.
4384::
4378:8
4362:.
4340::
4334:2
4313:.
4291::
4283::
4275::
4248:.
4228::
4220::
4197:.
4191::
4181::
4154:.
4130::
4103:.
4079::
4071::
4044:.
4024::
4016::
3993:.
3969::
3942:.
3918::
3910::
3880:.
3866::
3858::
3828:.
3806::
3798::
3771:.
3759::
3751::
3729:.
3718:.
3694::
3688:6
3662:.
3648::
3640::
3609:.
3577::
3569::
3538:.
3516::
3489:.
3467::
3440:.
3434::
3426::
3399:.
3379::
3371::
3348:.
3342::
3334::
3304:.
3284::
3261:.
3249::
3241::
3218:.
3204::
3177:.
3153::
3147:7
3126:.
3094::
3086::
3080:2
3059:.
3047::
3024:.
3002::
2975:.
2953::
2943::
2935::
2908:.
2886::
2859:.
2847::
2839::
2816:.
2792::
2784::
2757:.
2735::
2727::
2700:.
2688::
2680::
2657:.
2637::
2629::
2606:.
2584::
2557:.
2533::
2527:6
2506:.
2482::
2474::
2447:.
2427::
2419::
2389:.
2369::
2361::
2338:.
2316::
2289:.
2269::
2246:.
2232::
2224::
2197:.
2173::
2165::
2135:.
2121::
2115:4
2094:.
2070::
2043:.
2031::
2008:.
1988::
1982:2
1962:.
1950::
1942::
1919:.
1897::
1889::
1862:.
1840::
1834:7
1806:.
1784::
1755:.
1744:.
1720::
1677:.
1663::
1657:4
1554:(
1170:e
1163:t
1156:v
689:e
682:t
675:v
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.