763:
to regain adaptive capability. This has led to the suggestion that high mutation rates evolved to allow such mutant spectrum recovery following bottlenecks. Other models attribute high mutation rates to adaptive optimization independent of bottlenecks, or to a mechanistic consequence of rapid replication. Whatever their ultimate origins, high mutation rates serve the purpose of adaptation in multiple circumstances, not only following bottlenecks. A founder virus can introduce a different phenotype for the ensuing evolution. Evolution of viruses in nature and as disease agents can be viewed as succession of mutant spectrum alterations, subjected to expansions and reductions of population size in a continuous interplay of positive and negative selection and random drift. While short-term (for example, intra-host) evolution is observable and measurable, viruses may appear to be relatively static in the long term for decades (as seen with antigenic variants of FMDV ) or longer. Intra-host evolution is generally more rapid than inter-host evolution, as documented with viruses and other biological systems. Apparent invariance may be the result of selection for long-term survival of populations that have previously frenziedly tested evolutionary outcomes in short-term processes.
546:
different mutations to the same genome molecule) can be established. Each of these procedures implies some limitations: biological cloning can bias the representation in favor of infectious genomes, while molecular cloning can introduce non-infectious (defective) genomes in the analysis. Whole genome quasispecies description is still technically challenging due to the artifactual introduction of mutations. Most current deep sequencing platforms yield sequences of short reads for a given amplicon (sequence under analysis); minority mutations in an amplicon cannot be reliably linked to mutations in a different amplicon of the same genome; at most, statistical inferences on linkage can be proposed. Despite these limitations, control experiments and improvements of bioinformatic procedures support that the majority of sequence heterogeneity analyzed in viral populations indeed reflects differences in the natural template populations. If mutation linkage can be solved on a routine basis, a new wave of molecular information relevant to epistatic interactions will enter the picture.
710:
of the population. Complementation was described between two truncated FMDV genomic forms. The genomes with internal deletions became detectable upon high multiplicity passage of a clonal population of standard FMDV, a virus with a monopartite single stranded RNA genome. Infectivity was generated by complementation of the two truncated forms, in absence of standard, full length FMDV genomes. For complementation to be effective, prior exploration of sequence space through point mutations was a requirement. The system underwent a remarkable evolutionary transition akin to genome segmentation. Drastic genetic lesions in viral genomes are difficult to observe unless a mechanism such as complementation comes into the rescue of the deviant genomes. Additional examples of complementation among RNA viruses have been reported. Complementation is a means to maintain defective genomes at detectable frequencies in viral populations.
550:
uncertainties or bona fide fluctuations of genome frequencies. It is not justified to accept a rough similarity because even a single mutation in a given sequence context may affect biological properties. In the words of John
Holland and colleagues: "It is important to remember that every quasispecies genome swarm in an infected individual is unique and "new" in the sense that no identical population of genomes has ever existed before and none such will ever exist again". On top of the fleeting nature of any mutant distribution, the standard methods available for quasispecies characterization provide genomic sequences of a minority of the population (estimated in 10 to 10 for molecular cloning-Sanger sequencing, and in 10 to 10 for deep sequencing). We can only have an approximate representation of viral populations and their dynamics, as evidenced by many experimental studies.
928:
error threshold relationship. Both the error threshold and lethal mutagenesis are highly fitness landscape-dependent, but both can occur in complex fitness landscapes as those pertinent to viral populations. The term lethal mutagenesis was coined by
Lawerence Loeb and colleagues, and it is now widely used to describe the antiviral activity of base and nucleoside analogues that increase the viral mutation rate. Although several models have been proposed to account for virus extinction by excess mutations, an extension of the violation of the error threshold stands as a likely mechanism. Interestingly, some antiviral agents licensed for human use, initially thought to act only as inhibitors of viral replication, may actually exert their antviral activity against some RNA viruses at least partially by lethal mutagenesis. This is the case of
601:(very strong selection for a trait), an individual (or a limited number of individuals) that encodes signatures prone to be selected, may approach dominance while becoming the founder of a mutant cloud (because formation of a cloud is inherent to replication). Conditions for dominance (in this case in response to selection) are that the genome senses the selective sweep and that its replication in the new selective environment is permitted. In other cases, a collection of mutants is selected. This was illustrated with a FMDV quasispecies that was reconstructed in the laboratory with multiple antigenic variants (each at low frequency) that belonged to two different categories, and shared resistance to the same
980:
746:, or fitness decrease by the irreversible incorporation of mutations in asexual organisms in absence of compensatory mechanisms. The serial bottleneck transfers unveiled the presence rare mutations, not seen in standard laboratory or natural viral populations. In absence of forced bottleneck events, such rare mutations would be lost by negative selection because of their fitness cost. The investigation of how FMDV clones debilitated by Müller’s ratchet regained replicative fitness revealed several alternative molecular pathways for fitness recovery. The implications of this observation went largely unnoticed until recent results with
161:
1056:
951:, etc.) represent a natural counterpart of the principle utilized by lethal mutagenesis. Applicability to pathogenic cellular elements is a real possibility, and lethal mutagenesis to control tumor cells is an active field of investigation. Thus, the recognition of quasispecies dynamics has suggested some fundamental guidelines for disease prevention and control that are gradually permeating clinical practice. This is in line with the recognized need to apply Darwinian principles to the control of infectious disease.
444:
731:
234:
650:
in which an effective competition among variants is established, for example within replication complexes. This important concept was first derived theoretically, and then approached experimentally with several viruses. In an early study, Juan Carlos de la Torre and John
Holland described suppression of high fitness VSV by mutant spectra of inferior fitness. Suppressive effects have since been documented with standard and mutagenized viral populations. Some examples are:
575:(IFN) or to respond to IFN, virulence or particle stability, among other phenotypic traits. Mutant spectra can also mediate cyclical adaptation to different cell types. A mutant spectrum defines a consensus but the consensus is an abstraction; it may not be represented in the population. Many events in viral pathogenesis and evolution are due to mutant spectrum modifications or interactions which cannot be properly interpreted solely on the basis of consensus sequences.
288:, among other studies with animal and plant viruses in the middle of the 20th century. When put in the context of present-day knowledge, we realize that these observations on phenotypic changes were the tip of the iceberg of an extremely complex reality of viral populations. High mutation rates and population heterogeneity characterize RNA viruses, with consequences for viral pathogenesis and the control of viral disease. Detailed studies on quasispecies dynamics
614:
dominant variants were surrounded by a cloud of mutants of the other antigenic variant category. Conversely, passages in the presence of the antibody led to selection of variants with altered receptor recognition, surrounded by a cloud of antigenic variants that maintained receptor recognition. The results underlined the role of mutant clouds in selective events, and unveiled a new mechanism of antigenic flexibility.
702:
capacity and pathogenic potential of RNA viruses. In the first mutant studied, amino acid substitution G46S in the PV polymerase resulted in about four-fold increase in template-copying fidelity. This modification reduced PV adaptability and infective potential in vivo. The mutant in isolation did not replicate efficiently in the brain of susceptible mice, but it did when its mutant spectrum was broadened by
755:
interesting to investigate whether focused adaptation of other viruses to a specific environment may also entail a broadening of diversity, with many phenotypic variants attaining similar fitness levels. If generalized, this broadening of phenotypic space would provide a new interpretation of the molecular basis of adaptation, and explain why adaptation to alternative environments may not lead to
529:
equated with the effective population size in general genetics), and harboring multiple mutations per genome. The scenarios suggested by current experimental data defy our imagination. The relative frequency of individual mutations fluctuates in an unceasing exploration of sequence space, with phenotypic changes (not only genotypic changes) being far more frequent than previously thought. The
320:. These objectives approximate quasispecies to the real case of RNA viruses, which are compelled to deal with dramatic variations in population size and environment. Research on quasispecies has proceeded through several theoretical and experimental avenues that include continuing studies on evolutionary optimization and the origin of life,
344:(a class of proteins with conformation-dependent pathogenic potential; in this case the quasispecies is defined by a distribution of conformations). New inputs into experimental quasispecies research have come from deep sequencing to probe viral and cellular populations, recognition of interactions within mutant spectra, models of viral
636:. Thus, memory is a history-dependent, collective property of the quasispecies that confers a selective advantage to respond to environmental changes previously experienced by the same evolutionary lineage. It can be manifested only if the mutant spectrum maintains its completeness, since memory is lost when the population undergoes a
739:
have an important participation in shaping evolutionary lineages for all kinds of organisms, and also for viruses. They occur frequently not only upon host-to host transmission but also inside infected hosts, and they can perturb positive and negative selection events in processes that are difficult to identify and characterize.
537:(FMDV) such a design led to a remarkable phenotypic diversification into subpopulations of colonizers and competitors, that modulated virulence of the mutant ensemble. In HCV such a design unveiled continuous mutation waves and a more accurate understanding of the types of fitness landscapes occupied by high fitness viruses.
316:. These conditions are common in initial theoretical formulations of complex phenomena because they confer mathematical tractability. Since then, several extensions of the theory to non-equilibrium conditions with stochastic components have been developed, with the aim of finding general solutions for multi-peak
559:
conveying the information recapitulated in a mutant spectrum, blurs and enfeebles biological interpretations. Experimental results have demonstrated that minority genomes from a mutant spectrum (that cannot be identified by examining the consensus sequence) can include mutations that confer resistance to
831:
of antiviral designs. The aim of vaccination is to evoke a protective response that either prevents virus replication or disease. The aim of an antiviral pharmacological intervention is to inhibit virus replication to provide the immune system with an opportunity to clear the virus. Expressed simply,
762:
Deprivation of an individual virus from possible suppression, complementation or cooperation, may represent a liberation to initiate a new evolutionary process, or a condemnation to extinction. If liberated from suppression, the isolated genome must replicate and be able to reconstruct a mutant cloud
962:
In theory, if the mutation rate was sufficiently high, the viral population would not be able to maintain the genotype with the highest fitness, and therefore the ability of the population to adapt to its environment would be compromised. A practical application of this dynamic is in antiviral drugs
927:
is the process of virus extinction at the error rate at which a virus can no longer maintain its genetic information. Application of lethal mutagenesis as an antiviral strategy deserves attention in the context of the present article because its origins lie in quasispecies theory, in the form of the
709:
Complementation (often occurring when a functional protein encoded by a set of genomes is used by another set of genomes whose encoded protein is not functional) may underlie some collective responses of quasispecies such as fitness of individuals isolated from a population being inferior to fitness
649:
Individual genomes surrounded by a cloud of related mutants can be either suppressed to be kept at low frequency, or helped to be maintained in the population. The two alternative fates are dependent on several factors, one being the surrounding mutant spectrum in those steps of the infectious cycle
549:
There are additional levels of indeterminacy in the sequential analysis of viral populations, in particular those replicating in vivo. Components of the mutant spectrum represented at a given time in the sample taken for sequencing may differ from those in the next time point, due either to sampling
545:
The nucleotide sequence of an individual genome from a population (no matter which the degree of population complexity might be), can be determined either following a biological or molecular cloning event or by deep sequencing of entire viral genomes, in a manner that mutation linkage (assignment of
507:
systems may be overwhelmed by the high infectious dose, but also because the mutant repertoire that engages in adaptive explorations is larger. Part of the variants of a mutant spectrum, either in isolation or in consortium with others, may perform better than other members of the same population in
390:
have established some general observations on the mechanisms of mutant generation, and implications of quasispecies dynamics. In RNA virus genetics when we speak of "a mutant" the entity we handle is a cloud of mutants in which the specific mutation to which we direct our attention is present in all
164:
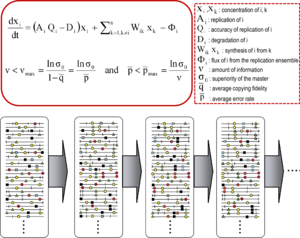
The equations are the mathematical expression of the major concepts implied by quasispecies theory. The first equation describes the change of concentration of molecule i as a function of replication parameters, and its production from other molecules of the same ensemble. The second equation is the
738:
A means to interrupt the participation of individual genomes in interactions with their mutant spectrum is for the quasispecies swarm to undergo drastic reductions in population size that isolate one or few individual genomes from their surroundings. Such reductions are termed bottlenecks, and they
640:
event that excludes minorities. A relevant example of the consequences of memory occurs in antiviral pharmacology with the administration for a second time of the same or a related antiviral agent (capable of evoking shared resistance mutations) used in a previous treatment. The second intervention
631:
that mobilizes and expands minority components in response to stimuli previously faced by the system. In the experiments designed to identify memory in viral quasispecies, members of the mutant spectrum increased in frequency as a consequence of their replication during a selection event that drove
520:
cells, to withstand neutralizing antibodies, etc.). Adaptability of RNA viruses is linked to parameters that facilitate exploration of sequence space: genome size (1.8 to 33 Kb), population size (variable but that can attain an impressive 10 individual genomes in an infected host at a given time),
245:
populations whose replication had been initiated by a single virus particle. Individual genomes differed from the consensus sequence in an average of one to two mutations per individual genome. Fitness of biological clones was inferior to that of the parental, uncloned population, a difference also
970:
However, these models assume that only the mutations that occur in the fittest sequence are deleterious, and furthermore that they are non-lethal. It has been argued that, if we take into account the deleterious effect of mutations on the population of variants and the fact that many mutations are
818:
In all interactions conductive to disease, the host cells individually and as groups in tissues and organs play decisive roles. The consequences of a viral infection are always host-dependent. However, the virus itself poses a major challenge that a deeper understanding of quasispecies dynamics is
713:
A distinction has been made between complementation and cooperation, in which two different genomes give rise to a new phenotype through the interaction between two variant proteins. An example of cooperation was characterized during studies with measles virus on membrane fusion which is essential
701:
Collective behavior of viruses was documented with mutant RNA viruses resistant to nucleotide analogues. The study of this class of mutants has been instrumental for the understanding of the molecular basis of template copying fidelity, and the consequences of fidelity alterations in the adaptive
558:
The points summarized in previous sections fully justifies addressing analytical tools towards the mutant spectrum rather than ignoring it or considering its presence a side issue. Use of consensus sequences to describe the genome of a virus isolate, despite being warranted by the difficulties of
503:, each population is made of an array of variant genomes, with a total number which is commensurate with the virus population size. To infect a plant, animal or cell culture with 10 infectious units can have very different consequences than to infect with 10 infectious units, not only because the
382:
genomes, do not prevent the formation of mutant spectra, although their amplitude may be lower than for other RNA viruses, at least in populations close to a clonal (single genome) origin. Quasispecies dynamics will operate in any viral or cellular system in which due to high mutation rates (as a
271:
High mutation rates and quasispecies were verified for other RNA viruses based on dissection of viral populations by molecular or biological cloning, and sequence analysis of individual clones. John
Holland and colleagues were the first to recognize that a rapidly evolving RNA world inserted in a
754:
conferred the capacity to infect several human cell lines in addition to the expected fitness increase for multiplication in BHK-21 cells. Thus, several lines of evidence suggest that fitness gain in a specific environment may paradoxically broaden the phenotypic potential of a virus. It will be
641:
may face inhibitor-resistant memory genomes from the earlier treatment, thus contributing to virus escape. This is an aspect that has not received adequate attention in the planning of antiviral interventions for patients who fail a first treatment and have to be subjected to a second treatment.
613:
recognition site); in the other category, the substitutions affected the antigenic determinant but not the receptor recognition site. Passages of the virus in absence of the monoclonal antibody resulted in dominance of antigenic variants that maintained the receptor recognition capacity, but the
693:
with members of the mutant spectrum. The position in a fitness landscape influences vulnerability to mutations, as popularized with the terms "advantage of the flattest" or "survival of the flattest", indicating that a variant located at the top of a sharp fitness peak has higher probability to
528:
from an infected host, adaptation to cell culture for studies on experimental evolution, or adaptation to alternative hosts in vivo. The reality is even more complex, given the large population sizes, with an indeterminate proportion of genomes actively replicating at any given time (sometimes
229:
The core quasispecies concepts are described by two fundamental equations: replication with production of error copies, and the error threshold relationship. They capture two major features of RNA viruses at the population level: the presence of a mutant spectrum, and the adverse effect of an
832:
the direct danger for vaccination and treatment is that the virus can escape through selection of mutants resistant to vaccine-triggered defense components or to the externally administered inhibitors. This has led to several proposals to confront viral disease, that can be summarized below.
813:
There is a connection between four parameters that characterize viruses during infection processes: replication rate (the rate at which viral RNA or DNA is synthesized intracellularly for viral progeny production), viral load (the total amount of virus quantified in an infected host or host
1046:(or breadth) of the quasispecies. The mutant viruses extracted from brain tissue were not themselves pathogenic, and the authors speculate that there may be complementation between variant members of the quasispecies that could enable viruses to colonize different host tissues and systems.
916:
recombination is available) or because the multiple mutations inflict a severe fitness cost. Vaccines exposing multiple epitopes and combination therapies follow the same strategy whose aim is to limit possible escape routes to viral quasispecies in the face of the suppressive constraint.
915:
These strategies have as their main objective to avoid selection of treatment-escape mutants by multiple selective constraints that cannot be surmounted by the virus. Control is effective either because exploration of sequence space cannot reach the required multiple mutations (even when
254:
of a population ensemble need not coincide with that of its individual components. The finding that a viral population was essentially a pool of mutants came at a time when mutations in general genetics were considered rare events, and virologists associated a viral genome with a defined
508:
the event of an environmental change. Selective pressures favor replication of some components of a mutant spectrum over others, despite all of them being interconnected by mutation. Differential performance can be at the level of viral genomes (during replication, intracellular
632:
them towards dominance. When the selective constraint was withdrawn, memory genomes remained at levels that were 10- to 100-fold higher than the basal levels attributable solely to their generation by mutation, as documented with independent FMDV genetic markers, and with HIV-1
373:
domain present in replicative cellular DNA polymerases. Also, postreplicative-repair pathways, abundant to correct genetic lesions in replicating cellular DNA, appear as ineffective for double-stranded RNA or RNA-DNA hybrids. The presence of a proofreading-repair activity in
272:
DNA-based biosphere had multiple evolutionary and medical implications. Genome plasticity of RNA viruses had been suspected for many decades. Key early observations were variations in viral traits described by
Findley in the 1930s, the studies of Granoff on transitions of
487:. All modes of molecular variation are compatible, only restricted by the scope of mechanisms accessible to the replicative machinery, and for the need for viral genomes to remain functional. David Evans and colleagues identified many recombination events associated with
173:(p = 1- q; q is the copying fidelity) for maintenance of genetic information. Terms are defined in the box on the right. Below, an evolving mutant spectrum (with mutations represented as symbols on the genomes), with an invariant consensus sequence. Adapted from.
714:
for virus entry into cells. For this virus fusion is mediated by two proteins termed H and F. A truncated H was deficient in cell fusion but the activity was regained when the truncated H was accompanied by two forms of F but not one of the forms individually.
852:. The broad response should minimize selection of escape mutants that may be present as minority components in mutant spectra, as repeatedly documented experimentally. With the current types of available vaccines, those that best comply with the multiple
583:
Mutant spectra are not mere aggregates of mutants acting independently. They are often engaged in collective responses. Two major types are those that depend on the presence of sets of variants, and those that rely on intra-mutant spectrum interactions.
721:
and suppression can emerge from interactions among components of mutant spectra that have their origin in random mutations. Selection acts on whatever sets of mutants can provide a useful trait, to turn random occurrences into biological meaning.
391:(or the great majority of) individual genomes. There is no such a thing as "a" wild type or "a" mutant virus. They are always clouds of mutants. Changes in the relative dominance of components of mutant spectra are particularly severe during
348:
related to disease progression and pathogen transmission, and new teachings from fidelity variants of viruses. Here we summarize the main aspects of quasispecies dynamics, and recent developments relevant to virus evolution and pathogenesis.
267:
of individual genomes to accept an undetermined proportion of the newly arising mutations, despite fitness costs. The error rate estimated for bacteriophage Qβ has been confirmed, and is comparable to values calculated for other RNA viruses.
814:
compartment), genetic heterogeneity, and replicative fitness (the yield of infectious particles that can contribute to the next generation). They can influence disease progression, and any of them can be targeted for disease control.
402:
procedures have been developed to unveil the relationships among different but closely related genome types that may suggest some hierarchical order of mutation acquisition or identification of transmission clusters (examples are
66:
will outcompete a quasispecies located at a higher but narrower fitness peak in which the surrounding mutants are unfit. This phenomenon has been called 'the quasispecies effect' or, more recently, the 'survival of the flattest'.
2159:
Domingo E, Martínez-Salas E, Sobrino F, de la Torre JC, Portela A, Ortín J, et al. (January 1985). "The quasispecies (extremely heterogeneous) nature of viral RNA genome populations: biological relevance--a review".
495:. High mutation and recombination rates have led to the conceptual distinction between mechanistically unavoidable and evolutionarily relevant variation, in connection with the issue of clonal versus non-clonal nature of
626:
behavior of a viral quasispecies, suggested by the presence of core information (considered the one that defines viral identity) despite variation of constitutive elements (the mutant spectrum). A well-known example is
217:
relationship, which marks the maximum mutation rate at which the master (or dominant) sequence can stabilize the mutant ensemble. Violation of the error threshold results in loss of dominance of the master sequence and
117:, though the concepts can apply to other biological entities such as reverse transcribing DNA viruses like hepatitis B. In such scenarios, complex distributions of closely related variant genomes are subjected to
971:
lethal, then the error threshold disappears, i.e. the fittest sequence always maintains itself. Empirical data on the effect of mutations in viruses is rare, but appears to correspond with this scenario.
810:
The larger the actively replicating (effective) population size and the replication rate, the most effective is exploration of sequence space for phenotypic expansions that favor survival and persistence.
1011:
to preserve diversity in a population. At least in simulations, a slower replicator can be shown to be able to outcompete a faster one in cases where it is more robust and the mutation rate is high.
4925:
Perales C, Mateo R, Mateu MG, Domingo E (June 2007). "Insights into RNA virus mutant spectrum and lethal mutagenesis events: replicative interference and complementation by multiple point mutants".
201:
information, as an essential step in the origin of life. The theory portrayed early replicon populations as organized mutant spectra dominated by a master sequence, the one endowed with the highest
734:
Illustration of bottleneck of different severity, defined by the different circles inserted in the entire population (large rectangle) and the outer rectangles. Symbols represent mutant classes.
622:
Quasispecies memory is a type of molecular memory dependent on the recent history of the evolutionary lineage and the integrity of the mutant spectrum. The search for memory was prompted by the
6972:
2294:
Meyerhans A, Cheynier R, Albert J, Seth M, Kwok S, Sninsky J, et al. (September 1989). "Temporal fluctuations in HIV quasispecies in vivo are not reflected by sequential HIV isolations".
742:
Drastic bottleneck events have been reproduced with laboratory populations of viruses in the form of plaque-to-plaque transfers. This design served to verify experimentally the operation of
4296:
García-Arriaza J, Ojosnegros S, Dávila M, Domingo E, Escarmís C (July 2006). "Dynamics of mutation and recombination in a replicating population of complementing, defective viral genomes".
872:> single peptide antigen. The scarcity of effective synthetic vaccines for RNA viral pathogens despite huge scientific and economic efforts is a reflection of the underlying problems.
521:
replication rate, mutation rate, fecundity (yield of viral particles per cell), and number of mutations required for a phenotypic change (surprisingly low for several relevant traits).
4188:
Gallego I, Gregori J, Soria ME, García-Crespo C, García-Álvarez M, Gómez-González A, et al. (October 2018). "Resistance of high fitness hepatitis C virus to lethal mutagenesis".
669:
959:
This may be defined as "The inability of a genetic element to be maintained in a population as the fidelity of its replication machinery decreases beyond a certain threshold value".
6957:
3210:
Figlerowicz M, Alejska M, Kurzynska-Kokorniak A, Figlerowicz M (2003-09-23). "Genetic
Variability: The Key Problem in the Prevention and Therapy of RNA-Based Virus Infections".
362:
129:. Therefore, the evolutionary trajectory of the viral infection cannot be predicted solely from the characteristics of the fittest sequence. High mutation rates also place an
1000:(with mutational neighbours substantially less fit). This has been called 'survival of the flattest' - referring to the fitness profiles of the two strategies respectively.
939:
Defense mechanisms based on genome modification of invading genetic parasites such as editing cellular activities that are recruited as part of the innate immune response (
524:
Mutant spectrum dynamics has been depicted in different ways, and we have chosen one that encompasses frequent events in natural populations and research designs, such as
2027:
Batschelet E, Domingo E, Weissmann C (January 1976). "The proportion of revertant and mutant phage in a growing population, as a function of mutation and growth rate".
1790:
Fornés J, Tomás Lázaro J, Alarcón T, Elena SF, Sardanyés J (January 2019). "Viral replication modes in single-peak fitness landscapes: A dynamical systems analysis".
6013:
Escarmís C, Dávila M, Domingo E (January 1999). "Multiple molecular pathways for fitness recovery of an RNA virus debilitated by operation of Muller's ratchet".
500:
6962:
2557:
Bernad A, Blanco L, Lázaro JM, Martín G, Salas M (October 1989). "A conserved 3'----5' exonuclease active site in prokaryotic and eukaryotic DNA polymerases".
471:. Accumulating laboratory and clinical evidence renders untenable that minority components of mutant spectra should be dismissed on the grounds of their being
321:
263:. The cloud nature of Qβ was understood as a consequence of its high mutation rate, calculated in 10 mutations introduced per nucleotide copied, together with
6091:"Evolution of the capsid protein genes of foot-and-mouth disease virus: antigenic variation without accumulation of amino acid substitutions over six decades"
1018:
evolved or is intrinsic to genetic systems is unconfirmed, because the basic mechanism behind robustness would depend upon the peculiarities of each system.
891:
Splitting a treatment into two steps: first an induction regimen, and a second maintenance regimen. Drugs administered in the two steps should be different.
499:(microbial evolution in general). Only a minority of the nascent variation during replication can be successfully propagated. Within limits that are set by
789:
Virus attenuation and virulence is dependent on viral genetic traits. Variant forms of a given virus may display increased virulence or atypical disease.
5978:
Escarmís C, Dávila M, Charpentier N, Bracho A, Moya A, Domingo E (November 1996). "Genetic lesions associated with Muller's ratchet in an RNA virus".
4606:
Farber DL, Netea MG, Radbruch A, Rajewsky K, Zinkernagel RM (February 2016). "Immunological memory: lessons from the past and a look to the future".
205:(replicative capacity) in the distribution. It introduced the notion of a mutant ensemble as a unit of selection, thus emphasizing the relevance of
5477:
Aaskov J, Buzacott K, Thu HM, Lowry K, Holmes EC (January 2006). "Long-term transmission of defective RNA viruses in humans and Aedes mosquitoes".
3697:"Recombination in enteroviruses is a biphasic replicative process involving the generation of greater-than genome length 'imprecise' intermediates"
2935:"Deep-Sequence Identification and Role in Virus Replication of a JC Virus Quasispecies in Patients with Progressive Multifocal Leukoencephalopathy"
160:
6394:
Perales C, Gallego I, de Ávila AI, Soria ME, Gregori J, Quer J, Domingo E (July 2019). "The increasing impact of lethal mutagenesis of viruses".
324:
and replicator networks, the error threshold in variable fitness landscapes, consideration of chemical mutagenesis and proofreading mechanisms,
2519:
Steinhauer DA, Domingo E, Holland JJ (December 1992). "Lack of evidence for proofreading mechanisms associated with an RNA virus polymerase".
884:(use of a single antiviral agent) is to be avoided. The following recommendations have been made and in some cases successfully implemented:
2984:"An unusually high substitution rate in transplant-associated BK polyomavirus in vivo is further concentrated in HLA-C-bound viral peptides"
779:) and antiviral inhibitors used in therapy. It is an open question if vaccination can promote long-term evolution of antigenic determinants.
771:
Soon after quasispecies was evidenced for viruses, some medical implications were made explicit. Several specific and general points below.
750:(HCV) have also suggested the accessibility of multiple pathways for fitness gain. Also, extensive passage of a biological clone of FMDV in
475:. They can participate in selective processes and cannot be excluded from interpretations of virus behavior. Variation universally involves
459:
The crux of the matter regarding quasispecies implications is that at any given time, the viral population includes a reservoir not only of
1042:
sequences. Pathogenicity could then be restored by mutagen application. This was interpreted to mean lower mutation rates had reduced the
447:
Upon isolation from an infected host (middle boxes), a virus sample may be adapted to cultured cells and subjected to large population or
533:
design that consists of passaging viral populations for long time periods (many sequential infections) is often extremely revealing. In
6967:
4336:"Correlation between amount of virus with altered nucleotide sequence and the monkey test for acceptability of oral poliovirus vaccine"
4077:"Internal Disequilibria and Phenotypic Diversification during Replication of Hepatitis C Virus in a Noncoevolving Cellular Environment"
3561:"In vivo evolution of viral virulence: switching of deformed wing virus between hosts results in virulence changes and sequence shifts"
2383:
Eigen M, Schuster P (November 1977). "The hypercycle. A principle of natural self-organization. Part A: Emergence of the hypercycle".
4960:
Crowder S, Kirkegaard K (July 2005). "Trans-dominant inhibition of RNA viral replication can slow growth of drug-resistant viruses".
775:
High mutation rates and population heterogeneity endow viruses with the potential to escape immune pressures (including those due to
4477:"Influence of the mutant spectrum in viral evolution: focused selection of antigenic variants in a reconstructed viral quasispecies"
101:
The quasispecies model is most applicable when the genome size is limited and the mutation rate is high, and so is most relevant to
4837:"Suppression of lymphocytic choriomeningitis virus--induced growth hormone deficiency syndrome by disease-negative virus variants"
491:
replication, and only a few recombinants made their way towards continued replication. Recombination can mediate adaptability and
685:
Opposite to suppression is maintenance of a mutant either by a favorable position in a fitness landscape or by interactions of
996:
to generate a broad quasispecies with members of approximately equal fitness than to have a sharply defined 'most fit' single
807:
processes in which some viral subpopulations are replaced by others to persist or to invade new cell types, tissues or organs.
6977:
4429:
Perales C (October 2018). "Quasispecies dynamics and clinical significance of hepatitis C virus (HCV) antiviral resistance".
3988:
2495:
1551:
Molecular characterization of full genome hepatitis b virus sequences from an urban hospital cohort in
Pretoria, South Africa
936:(1-β-D-ribofuranosyl-1-H-1,2,4-triazole-3-carboxamide) that are currently being intensively investigated as lethal mutagens.
59:
3038:"Differential Shape of Geminivirus Mutant Spectra Across Cultivated and Wild Hosts With Invariant Viral Consensus Sequences"
3036:
Sánchez-Campos S, Domínguez-Huerta G, Díaz-Martínez L, Tomás DM, Navas-Castillo J, Moriones E, Grande-Pérez A (2018-07-02).
1067:
6294:
Perales C, Ortega-Prieto AM, Beach NM, Sheldon J, Menéndez-Arias L, Domingo E (2017). "Quasispecies and Drug
Resistance".
4788:"Distinct repertoire of antigenic variants of foot-and-mouth disease virus in the presence or absence of immune selection"
1899:
Domingo E, Sabo D, Taniguchi T, Weissmann C (April 1978). "Nucleotide sequence heterogeneity of an RNA phage population".
4698:
Arias A, Ruiz-Jarabo CM, Escarmís C, Domingo E (May 2004). "Fitness increase of memory genomes in a viral quasispecies".
897:
Use of innate immune response-stimulating drugs (for example, inhibitors of enzymes involved in pyrimidine biosynthesis).
293:
144:
The relevance of quasispecies in virology has been the subject of extended debate. However, standard clonal analyses and
148:
methodologies have confirmed the presence of myriads of mutant genomes in viral populations, and their participation in
6823:
Wagner GP, Krall P (November 1993). "What is the difference between models of error thresholds and Muller's ratchet?".
3608:
Baccam P, Thompson RJ, Fedrigo O, Carpenter S, Cornette JL (January 2001). "PAQ: Partition
Analysis of Quasispecies".
308:
The first mathematical formulation of quasispecies was deterministic; it assumed steady state mutant distributions in
6987:
6311:
6212:
5658:
5335:
5176:
4750:
4405:
4253:
2858:
2443:
2221:
2011:
1868:
378:
increases their copying accuracy in about 15-fold. This and other repair activities, that may act on standard RNA or
5645:. Current Topics in Microbiology and Immunology. Vol. 392. Springer International Publishing. pp. 219–29.
5322:. Current Topics in Microbiology and Immunology. Vol. 392. Springer International Publishing. pp. 303–22.
5163:. Current Topics in Microbiology and Immunology. Vol. 392. Springer International Publishing. pp. 161–79.
2845:. Current Topics in Microbiology and Immunology. Vol. 392. Springer International Publishing. pp. 141–59.
2430:. Current Topics in Microbiology and Immunology. Vol. 392. Springer International Publishing. pp. 121–39.
1855:. Current Topics in Microbiology and Immunology. Vol. 392. Springer International Publishing. pp. 61–120.
383:
result of low fidelity nucleic acid polymerases or environmental alterations) mutant spectra are rapidly generated.
5159:
Tejero H, Montero F, Nuño JC (2016). "Theories of Lethal
Mutagenesis: From Error Catastrophe to Lethal Defection".
3800:"Reproductive clonality of pathogens: a perspective on pathogenic viruses, bacteria, fungi, and parasitic protozoa"
2116:
Holland J, Spindler K, Horodyski F, Grabau E, Nichol S, VandePol S (March 1982). "Rapid evolution of RNA genomes".
4651:"Memory in retroviral quasispecies: experimental evidence and theoretical model for human immunodeficiency virus"
800:
than most genomes in the same population, with implications for the emergence and re-emergence of viral disease.
5103:"The fittest versus the flattest: experimental confirmation of the quasispecies effect with subviral pathogens"
2267:
1755:
Swetina J, Schuster P (December 1982). "Self-replication with errors. A model for polynucleotide replication".
993:
6050:"Expansion of host-cell tropism of foot-and-mouth disease virus despite replication in a constant environment"
4524:
237:
Flow of conceptual derivations of quasispecies theory for viral populations, and some biological consequences.
4238:. Current Topics in Microbiology and Immunology. Vol. 176. Berlin, Heidelberg: Springer. pp. 1–20.
948:
387:
2426:
Saakian DB, Hu CK (2016). "Mathematical Models of Quasi-Species Theory and Exact Results for the Dynamics".
5684:"Bottleneck-mediated quasispecies restriction during spread of an RNA virus from inoculation site to brain"
606:
534:
358:
206:
3094:"Cyclical adaptation of measles virus quasispecies to epithelial and lymphocytic cells: To V, or not to V"
698:. Survival of the flattest has been also proposed as an ingredient in some models of the error threshold.
856:
requirement are, in the order of expected efficacy to confer protection against highly variable viruses:
484:
325:
130:
3512:"Within-Host Variations of Human Papillomavirus Reveal APOBEC Signature Mutagenesis in the Viral Genome"
6256:
Williams PD (February 2010). "Darwinian interventions: taming pathogens through evolutionary ecology".
5263:"Quasispecies diversity determines pathogenesis through cooperative interactions in a viral population"
979:
686:
247:
223:
1355:
Elena SF, Agudelo-Romero P, Carrasco P, Codoñer FM, Martín S, Torres-Barceló C, Sanjuán R (May 2008).
1003:
Over the long-term, a flatter fitness profile might better allow a quasispecies to exploit changes in
644:
5641:
Shirogane Y, Watanabe S, Yanagi Y (2016). "Cooperative Interaction Within RNA Virus Mutant Spectra".
2064:"Correlation between mutation rate and genome size in riboviruses: mutation rate of bacteriophage Qβ"
1699:
Eigen M (October 1971). "Selforganization of matter and the evolution of biological macromolecules".
828:
6242:
4772:
1994:
Domingo E, Brun A, Nuñez JI, Cristina J, Briones C, Escarmís C (2006-05-10). "Genomics of Viruses".
2602:"High fidelity of murine hepatitis virus replication is decreased in nsp14 exoribonuclease mutants"
751:
6508:"Perspective: APOBEC mutagenesis in drug resistance and immune escape in HIV and cancer evolution"
3404:
van Boheemen S, Tas A, Anvar SY, van Grootveld R, Albulescu IC, Bauer MP, et al. (May 2017).
911:
Lethal mutagenesis or virus extinction by excess of mutations introduced during viral replication.
6506:
Venkatesan S, Rosenthal R, Kanu N, McGranahan N, Bartek J, Quezada SA, et al. (March 2018).
4525:"Minority report: hidden memory genomes in HIV-1 quasispecies and possible clinical implications"
2751:"Coronaviruses as DNA wannabes: a new model for the regulation of RNA virus replication fidelity"
1298:
623:
333:
277:
2982:
Domingo-Calap P, Schubert B, Joly M, Solis M, Untrau M, Carapito R, et al. (October 2018).
2933:
Takahashi K, Sekizuka T, Fukumoto H, Nakamichi K, Suzuki T, Sato Y, et al. (January 2017).
2251:
2243:
743:
242:
6860:"The distribution of fitness effects caused by single-nucleotide substitutions in an RNA virus"
5589:"Cooperation between distinct viral variants promotes growth of H3N2 influenza in cell culture"
3749:
Xiao Y, Rouzine IM, Bianco S, Acevedo A, Goldstein EF, Farkov M, et al. (September 2017).
3643:
Skums P, Zelikovsky A, Singh R, Gussler W, Dimitrova Z, Knyazev S, et al. (January 2018).
3406:"Quasispecies composition and evolution of a typical Zika virus clinical isolate from Suriname"
782:
Attenuated RNA virus vaccines can revert to virulent forms. RNA viruses released in nature for
530:
366:
190:
6048:
Ruiz-Jarabo CM, Pariente N, Baranowski E, Dávila M, Gómez-Mariano G, Domingo E (August 2004).
5205:"Increased fidelity reduces poliovirus fitness and virulence under selective pressure in mice"
2841:
Wagner N, Atsmon-Raz Y, Ashkenasy G (2016). "Theoretical Models of Generalized Quasispecies".
654:
Suppression of high fitness antigenic variants of FMDV by low fitness antibody-escape mutants.
1015:
1008:
637:
564:
480:
448:
6089:
Martínez MA, Dopazo J, Hernández J, Mateu MG, Sobrino F, Domingo E, Knowles NJ (June 1992).
5782:
Bull RA, Luciani F, McElroy K, Gaudieri S, Pham ST, Chopra A, et al. (September 2011).
2340:
Farci P (November 2011). "New insights into the HCV quasispecies and compartmentalization".
241:
The existence of a mutant spectrum was experimentally evidenced first by clonal analyses of
6871:
6777:
6718:
6460:
6343:
6151:
5897:
5842:
5743:"Virus population bottlenecks during within-host progression and host-to-host transmission"
5695:
5541:
5486:
5431:
5274:
4347:
4234:
Holland JJ, De La Torre JC, Steinhauer DA (1992). "RNA Virus Populations as Quasispecies".
4137:
4020:
3862:
3572:
3417:
2818:
2801:
2662:
2125:
1799:
1708:
1502:
1310:
1246:
1179:
1162:
628:
251:
210:
5318:
Bordería AV, Rozen-Gagnon K, Vignuzzi M (2016). "Fidelity Variants and RNA Quasispecies".
4075:
Moreno E, Gallego I, Gregori J, Lucía-Sanz A, Soria ME, Castro V, et al. (May 2017).
4009:"Mutational and fitness landscapes of an RNA virus revealed through population sequencing"
3510:
Hirose Y, Onuki M, Tenjimbayashi Y, Mori S, Ishii Y, Takeuchi T, et al. (June 2018).
2250:. Current Topics in Microbiology and Immunology. Vol. 299. Springer-Verlag. pp.
1428:
8:
5784:"Sequential bottlenecks drive viral evolution in early acute hepatitis C virus infection"
1299:"Evolution of digital organisms at high mutation rates leads to survival of the flattest"
1071:
861:
718:
602:
345:
309:
256:
186:
75:
24:
6875:
6781:
6722:
6464:
6347:
6155:
5901:
5846:
5699:
5545:
5490:
5435:
5278:
4351:
4141:
4124:
Ojosnegros S, Beerenwinkel N, Antal T, Nowak MA, Escarmís C, Domingo E (February 2010).
4024:
3866:
3751:"RNA Recombination Enhances Adaptability and Is Required for Virus Spread and Virulence"
3576:
3421:
2666:
2129:
1945:
Duarte EA, Novella IS, Ledesma S, Clarke DK, Moya A, Elena SF, et al. (July 1994).
1803:
1712:
1506:
1314:
1250:
6840:
6800:
6765:
6682:
6657:
6630:
6605:
6581:
6556:
6532:
6507:
6429:
6230:
6177:
5866:
5810:
5783:
5718:
5683:
5615:
5588:
5564:
5529:
5510:
5454:
5419:
5418:
Moreno E, Ojosnegros S, García-Arriaza J, Escarmís C, Domingo E, Perales C (May 2014).
5386:
5346:
5295:
5262:
5231:
5204:
5129:
5102:
5029:
5004:
4985:
4760:
4675:
4650:
4631:
4454:
4275:
4213:
4160:
4125:
4101:
4076:
4041:
4008:
3939:
3912:
3885:
3850:
3826:
3799:
3775:
3750:
3723:
3696:
3669:
3645:"QUENTIN: reconstruction of disease transmissions from viral quasispecies genomic data"
3644:
3536:
3511:
3487:
3462:
3438:
3405:
3381:
3354:
3327:
3302:
3283:
3178:
3153:
3120:
3093:
3064:
3037:
3010:
2983:
2959:
2934:
2910:
2883:
2777:
2750:
2626:
2601:
2582:
2501:
2408:
2365:
2319:
2088:
2063:
1924:
1833:
1732:
1676:
1651:
1603:
1433:
1383:
1356:
1334:
1236:
1114:
1077:
1004:
924:
857:
662:
365:(also termed reverse transcriptases, RTs). In addition, these enzymes are defective in
285:
264:
138:
95:
91:
55:
6894:
6859:
6483:
6448:
6115:
6090:
5920:
5885:
5395:
5371:"Evolutionary transition toward defective RNAs that are infectious by complementation"
5370:
5078:
5053:
4902:
4877:
4812:
4787:
4666:
4583:
4558:
4442:
1971:
1946:
672:(LCMV) (that cause growth hormone deficiency in mice) by non-pathogenic LCMV variants.
90:
when describing viruses, and a conceptual framework for a deeper understanding of the
6982:
6899:
6844:
6805:
6746:
6741:
6706:
6687:
6635:
6586:
6537:
6488:
6433:
6421:
6371:
6366:
6331:
6307:
6273:
6218:
6208:
6169:
6120:
6071:
6030:
5995:
5960:
5956:
5925:
5858:
5815:
5764:
5723:
5664:
5654:
5620:
5569:
5502:
5459:
5400:
5351:
5331:
5300:
5236:
5182:
5172:
5134:
5083:
5069:
5034:
4977:
4942:
4907:
4858:
4817:
4803:
4746:
4715:
4680:
4623:
4588:
4536:
4498:
4446:
4411:
4401:
4375:
4370:
4335:
4313:
4267:
4259:
4249:
4205:
4165:
4106:
4046:
3984:
3944:
3890:
3831:
3780:
3728:
3674:
3625:
3621:
3590:
3541:
3492:
3443:
3386:
3332:
3275:
3227:
3183:
3125:
3069:
3015:
2964:
2915:
2882:
Schmidt TT, Reyes G, Gries K, Ceylan CÜ, Sharma S, Meurer M, et al. (May 2017).
2864:
2854:
2823:
2782:
2731:
2726:
2709:
2690:
2685:
2650:
2631:
2574:
2570:
2536:
2532:
2491:
2449:
2439:
2400:
2369:
2357:
2311:
2307:
2273:
2263:
2217:
2177:
2173:
2141:
2093:
2044:
2040:
2007:
1976:
1916:
1912:
1874:
1864:
1825:
1772:
1768:
1724:
1681:
1595:
1555:
1530:
1525:
1490:
1468:
1464:
1388:
1326:
1274:
1269:
1224:
1202:
1194:
1119:
1101:
756:
747:
610:
568:
317:
297:
202:
134:
126:
122:
118:
63:
48:
44:
6572:
6181:
6106:
5514:
4989:
4893:
4574:
4458:
4279:
4217:
3981:
Virus as populations: composition, complexity, dynamics, and biological implications
3660:
3287:
2586:
2505:
2412:
2323:
2204:
Domingo E, Holland JJ, Ahlquist P (1988). Domingo E, Holland JJ, Ahlquist P (eds.).
1962:
1928:
1837:
1736:
1437:
1055:
357:
The molecular basis of high error rates is the limited template-copying fidelity of
6889:
6879:
6832:
6795:
6785:
6736:
6726:
6677:
6669:
6625:
6617:
6576:
6568:
6527:
6519:
6478:
6468:
6411:
6403:
6361:
6351:
6299:
6265:
6200:
6159:
6110:
6102:
6061:
6022:
5987:
5952:
5915:
5905:
5870:
5850:
5805:
5795:
5754:
5713:
5703:
5646:
5610:
5600:
5559:
5549:
5494:
5449:
5439:
5390:
5382:
5341:
5323:
5290:
5282:
5226:
5216:
5164:
5124:
5114:
5073:
5065:
5024:
5016:
4969:
4934:
4897:
4889:
4848:
4807:
4799:
4738:
4707:
4670:
4662:
4635:
4615:
4578:
4570:
4488:
4438:
4365:
4355:
4305:
4239:
4197:
4155:
4145:
4096:
4088:
4036:
4028:
3934:
3924:
3880:
3870:
3821:
3811:
3770:
3762:
3718:
3708:
3664:
3656:
3617:
3580:
3531:
3523:
3482:
3478:
3474:
3433:
3425:
3376:
3366:
3322:
3314:
3267:
3219:
3173:
3165:
3115:
3105:
3059:
3049:
3005:
2995:
2954:
2946:
2905:
2895:
2846:
2813:
2772:
2762:
2721:
2680:
2670:
2621:
2613:
2566:
2528:
2483:
2431:
2392:
2349:
2303:
2255:
2209:
2169:
2133:
2083:
2075:
2036:
1999:
1966:
1958:
1908:
1856:
1815:
1807:
1764:
1716:
1671:
1663:
1607:
1587:
1520:
1510:
1460:
1423:
1415:
1378:
1368:
1338:
1318:
1264:
1254:
1184:
1174:
1127:
1109:
1091:
645:
Intra-mutant spectrum interactions for interference, complementation or cooperation
472:
337:
114:
40:
992:
The long-term evolution of the virus may be influenced in that it may be a better
512:, interaction with host factors, etc.) or viral particles (for thermal stability,
5800:
5759:
5742:
5221:
5119:
5020:
3713:
3371:
3110:
3000:
2767:
1096:
849:
695:
694:
decrease fitness as a result of new mutations than the same variant located at a
598:
525:
509:
496:
395:
313:
281:
214:
145:
6673:
6303:
6204:
4876:
González-López C, Arias A, Pariente N, Gómez-Mariano G, Domingo E (April 2004).
4244:
4201:
3209:
3035:
2802:"Thinking Outside the Triangle: Replication Fidelity of the Largest RNA Viruses"
2079:
6864:
Proceedings of the National Academy of Sciences of the United States of America
6711:
Proceedings of the National Academy of Sciences of the United States of America
6453:
Proceedings of the National Academy of Sciences of the United States of America
6336:
Proceedings of the National Academy of Sciences of the United States of America
6330:
Loeb LA, Essigmann JM, Kazazi F, Zhang J, Rose KD, Mullins JI (February 1999).
6140:"Time-dependent estimates of molecular evolutionary rates: evidence and causes"
5890:
Proceedings of the National Academy of Sciences of the United States of America
5688:
Proceedings of the National Academy of Sciences of the United States of America
5424:
Proceedings of the National Academy of Sciences of the United States of America
4340:
Proceedings of the National Academy of Sciences of the United States of America
4130:
Proceedings of the National Academy of Sciences of the United States of America
3855:
Proceedings of the National Academy of Sciences of the United States of America
3804:
Proceedings of the National Academy of Sciences of the United States of America
3766:
3429:
2888:
Proceedings of the National Academy of Sciences of the United States of America
2655:
Proceedings of the National Academy of Sciences of the United States of America
1811:
1495:
Proceedings of the National Academy of Sciences of the United States of America
1229:
Proceedings of the National Academy of Sciences of the United States of America
1062:
804:
703:
560:
517:
476:
399:
182:
83:
71:
6941:
6917:
6606:"Human cancers express mutator phenotypes: origin, consequences and targeting"
5833:
Chao L (November 1990). "Fitness of RNA virus decreased by Muller's ratchet".
5369:
García-Arriaza J, Manrubia SC, Toja M, Domingo E, Escarmís C (November 2004).
4938:
4711:
4309:
3318:
3271:
2487:
869:
730:
605:. One category included mutants with an amino acid substitution that affected
455:(right box). Relevant adaptive mutations are highlighted with colored symbols.
443:
233:
6951:
6269:
5943:
Muller HJ (May 1964). "The relation of recombination to mutational advance".
5420:"Exploration of sequence space as the basis of viral RNA genome segmentation"
4742:
4415:
4334:
Chumakov KM, Powers LB, Noonan KE, Roninson IB, Levenbook IS (January 1991).
4263:
3231:
2259:
1599:
1559:
1515:
1198:
1105:
1082:
1035:
1034:– and hence reduce their mutation rate – showed these variants to have lower
901:
219:
178:
165:
error threshold relationship, indicating the maximum amount of information (ʋ
110:
36:
6884:
6523:
5910:
5886:"Rapid fitness losses in mammalian RNA virus clones due to Muller's ratchet"
5708:
5554:
5498:
5444:
4493:
4476:
4150:
3913:"Rethinking quasispecies theory: From fittest type to cooperative consortia"
3875:
3851:"Clonality and intracellular polyploidy in virus evolution and pathogenesis"
3816:
3585:
3560:
3054:
2900:
2137:
2003:
1259:
230:
increase of mutation rate on virus survival, each with several derivations.
133:
compatible with inheritable information. Crossing such a limit leads to RNA
78:
consisted of mutant distributions, as found experimentally with present-day
6903:
6809:
6790:
6750:
6731:
6691:
6639:
6590:
6541:
6492:
6473:
6425:
6356:
6277:
6173:
6075:
6026:
5991:
5964:
5819:
5768:
5727:
5668:
5624:
5573:
5506:
5463:
5404:
5355:
5304:
5240:
5186:
5138:
5087:
5038:
4981:
4946:
4911:
4853:
4836:
4719:
4684:
4627:
4592:
4557:
Ruiz-Jarabo CM, Arias A, Baranowski E, Escarmís C, Domingo E (April 2000).
4540:
4502:
4450:
4317:
4209:
4169:
4110:
4050:
3948:
3894:
3835:
3784:
3732:
3678:
3629:
3594:
3545:
3496:
3447:
3390:
3336:
3279:
3223:
3187:
3129:
3073:
3019:
2968:
2919:
2868:
2827:
2786:
2735:
2675:
2635:
2453:
2361:
2353:
2277:
2097:
1878:
1829:
1685:
1534:
1472:
1392:
1373:
1330:
1278:
1206:
1123:
1043:
905:
888:
Inhibitors used in combination should target different viral gene products.
793:
783:
504:
468:
273:
6407:
6375:
6222:
6195:
Domingo E (1989). "RNA virus evolution and the control of viral disease".
6124:
6066:
6049:
6034:
5999:
5929:
5862:
4862:
4821:
4379:
4360:
4271:
3929:
3169:
2694:
2578:
2540:
2315:
2213:
2181:
2145:
2048:
1980:
1776:
1728:
1667:
1131:
963:
employing lethal mutagenesis. For example, increased doses of the mutagen
835:
4092:
3527:
2950:
2884:"GLN3 inactivation cause imbalanced dNTP pools and increased mutagenesis"
2617:
2404:
1920:
1241:
983:
This visualization of "survival of the flattest" in evolutionary biology.
929:
881:
776:
690:
587:
513:
488:
375:
370:
102:
6416:
5650:
5605:
5327:
5286:
5168:
4875:
4619:
4032:
2850:
2435:
1860:
1419:
1189:
94:
of RNA viruses than is offered by classical studies based on simplified
6836:
6707:"RNA virus error catastrophe: direct molecular test by using ribavirin"
2396:
2244:"Transitions in understanding of RNA viruses: a historical perspective"
2158:
1820:
1720:
1591:
1031:
1027:
797:
658:
572:
379:
312:
without perturbations derived from modifications of the environment or
149:
6164:
6139:
5417:
5261:
Vignuzzi M, Stone JK, Arnold JJ, Cameron CE, Andino R (January 2006).
189:(a term used to refer to any replicating entity), as an ingredient of
16:
Population structure of viruses with a large number of variant genomes
5854:
5528:
Ciota AT, Ehrbar DJ, Van Slyke GA, Willsey GG, Kramer LD (May 2012).
5052:
Quer J, Hershey CL, Domingo E, Holland JJ, Novella IS (August 2001).
4786:
Borrego B, Novella IS, Giralt E, Andreu D, Domingo E (October 1993).
4295:
3463:"Marine RNA Virus Quasispecies Are Distributed throughout the Oceans"
1549:
1322:
1163:"Historical Perspective on the Discovery of the Quasispecies Concept"
1039:
964:
933:
681:
Suppression of drug-resistant viral mutants during antiviral therapy.
492:
464:
329:
260:
198:
87:
79:
6621:
1406:
Eigen M, McCaskill J, Schuster P (1988). "Molecular Quasi-Species".
398:, with complex dynamics of intra-host heterogeneity and variations.
6047:
5587:
Xue KS, Hooper KA, Ollodart AR, Dingens AS, Bloom JD (March 2016).
4973:
997:
827:
There is an increasing perception that Darwinian principles should
460:
194:
106:
5005:"My Cousin, My Enemy: quasispecies suppression of drug resistance"
4556:
2932:
2062:
Bradwell K, Combe M, Domingo-Calap P, Sanjuán R (September 2013).
1354:
6505:
5368:
3403:
2600:
Eckerle LD, Lu X, Sperry SM, Choi L, Denison MR (November 2007).
2061:
1947:"Subclonal components of consensus fitness in an RNA virus clone"
853:
6942:
Video: Using fitness landscapes to visualize evolution in action
6293:
4187:
4123:
2482:. Current Topics in Microbiology and Immunology. Vol. 392.
894:
Targeting of cellular functions needed for the virus life cycle.
6557:"Lethal mutagenesis: targeting the mutator phenotype in cancer"
5977:
4697:
944:
865:
845:
841:
341:
137:, a transition that is the basis of an antiviral design termed
32:
6973:
Knowledge articles published in peer-reviewed literature (J2W)
5884:
Duarte E, Clarke D, Moya A, Domingo E, Holland J (July 1992).
5530:"Cooperative interactions in the West Nile virus mutant swarm"
4074:
3607:
3559:
Gisder S, Möckel N, Eisenhardt D, Genersch E (December 2018).
3558:
2981:
2115:
1789:
6332:"Lethal mutagenesis of HIV with mutagenic nucleoside analogs"
1898:
1222:
28:
6088:
5317:
4333:
4233:
3509:
1297:
Wilke CO, Wang JL, Ofria C, Lenski RE, Adami C (July 2001).
706:
mutagenesis or when it was co-inoculated with wild type PV.
6766:"Quasispecies theory in the context of population genetics"
6393:
5527:
4605:
3642:
3258:
Domingo E, Perales C (May 2018). "Quasispecies and virus".
2293:
1161:
Domingo E, García-Crespo C, Perales C (29 September 2021).
1160:
1060:
This article was adapted from the following source under a
940:
185:
to explain self-organization and adaptability of primitive
5781:
5260:
4785:
4648:
2026:
1944:
678:
Suppression of FMDV by capsid and polymerase FMDV mutants.
6919:
Using fitness landscapes to visualize evolution in action
5101:
Codoñer FM, Darós JA, Solé RV, Elena SF (December 2006).
5051:
5002:
3154:"Emergency Services of Viral RNAs: Repair and Remodeling"
2840:
1851:
Schuster P (2016). "Quasispecies on Fitness Landscapes".
1076:
Esteban Domingo Solans, Celia Perales (17 October 2019).
592:
6958:
Knowledge articles published in peer-reviewed literature
6329:
5740:
5586:
4924:
4878:"Preextinction viral RNA can interfere with infectivity"
3748:
2556:
2518:
1993:
1223:
van Nimwegen E, Crutchfield JP, Huynen M (August 1999).
792:
Components of a mutant spectrum can exhibit a different
5883:
5640:
4835:
Teng MN, Oldstone MB, de la Torre JC (September 1996).
4834:
2881:
875:
836:
Vaccine exposure of multiple B cell and T cell epitopes
467:
variants, conferring upon the population some adaptive
70:
The term quasispecies was adopted from a theory of the
6138:
Ho SY, Duchêne S, Molak M, Shapiro B (December 2015).
5476:
5054:"Contingent neutrality in competing viral populations"
4291:
4289:
3091:
3031:
3029:
2203:
1405:
932:(T-705; 6-fluoro-3-hydroxy-2-pirazinecarboxamide) and
588:
Variants that drive responses to selective constraints
35:(related by mutations). Quasispecies result from high
6137:
6012:
5100:
5003:
Kirkegaard K, van Buuren NJ, Mateo R (October 2016).
3694:
3355:"Quasispecies theory and the behavior of RNA viruses"
1649:
974:
675:
Suppression of FMDV by a mutagenized FMDV population.
609:(since the antigenic determinant overlapped with the
4649:
Briones C, Domingo E, Molina-París C (August 2003).
4006:
3848:
3300:
3092:
Donohue RC, Pfaller CK, Cattaneo R (February 2019).
1296:
483:(in its replicative and non-replicative modes), and
39:
as mutants arise continually and change in relative
6704:
6199:. Vol. 33. Birkhäuser Basel. pp. 93–133.
5741:Gutiérrez S, Michalakis Y, Blanc S (October 2012).
5362:
4869:
4552:
4550:
4286:
4126:"Competition-colonization dynamics in an RNA virus"
3205:
3203:
3201:
3199:
3197:
3026:
2599:
2248:
Quasispecies: Concept and Implications for Virology
62:and highly connected (that is, flat) region in the
6944:—contains an example of "survival of the flattest"
5681:
5202:
4400:. Berlin, Heidelberg: Springer Berlin Heidelberg.
3906:
3904:
3695:Lowry K, Woodman A, Cook J, Evans DJ (June 2014).
2799:
2648:
1554:. Pretoria, South Africa: University of Pretoria.
1451:Nowak MA (April 1992). "What is a quasispecies?".
553:
540:
177:Quasispecies theory was developed in the 1970s by
6041:
5643:Quasispecies: From Theory to Experimental Systems
5636:
5634:
5320:Quasispecies: From Theory to Experimental Systems
5161:Quasispecies: From Theory to Experimental Systems
5158:
4959:
4391:
4389:
3910:
2843:Quasispecies: From Theory to Experimental Systems
2800:Smith EC, Sexton NR, Denison MR (November 2014).
2651:"A novel 3'-end repair mechanism in an RNA virus"
2649:Nagy PD, Carpenter CD, Simon AE (February 1997).
2480:Quasispecies: From Theory to Experimental Systems
2428:Quasispecies: From Theory to Experimental Systems
2289:
2287:
1998:. Wiley-VCH Verlag GmbH & Co. KGaA: 367–388.
1853:Quasispecies: From Theory to Experimental Systems
6949:
6857:
6499:
6389:
6387:
6385:
6289:
6287:
5198:
5196:
4547:
4329:
4327:
3194:
2473:
2471:
2469:
2467:
2465:
2463:
1645:
1643:
1641:
1639:
1637:
6651:
6649:
5256:
5254:
5252:
5250:
4996:
4732:
4518:
4516:
4514:
4512:
4229:
4227:
4183:
4181:
4179:
4117:
4007:Acevedo A, Brodsky L, Andino R (January 2014).
4002:
4000:
3974:
3972:
3970:
3968:
3966:
3964:
3962:
3960:
3958:
3901:
3348:
3346:
3253:
3251:
3249:
3247:
3245:
3243:
3241:
3087:
3085:
3083:
2477:
2237:
2235:
2233:
2199:
2197:
2195:
2193:
2191:
2111:
2109:
2107:
2020:
1894:
1892:
1890:
1888:
1754:
1750:
1748:
1746:
1635:
1633:
1631:
1629:
1627:
1625:
1623:
1621:
1619:
1617:
1218:
1216:
280:, or the high frequency of conversions between
6249:
6082:
6006:
5734:
5631:
5154:
5152:
5150:
5148:
4522:
4470:
4468:
4386:
4070:
4068:
4066:
4064:
4062:
4060:
3797:
3791:
3460:
3257:
2552:
2550:
2284:
1940:
1938:
1578:Eigen M, Schuster P (1979). "The Hypercycle".
1484:
1482:
451:(left box), or be adapted to a different host
6963:Knowledge articles published in PLOS Genetics
6851:
6698:
6655:
6382:
6325:
6323:
6284:
6188:
5971:
5675:
5311:
5193:
4474:
4431:International Journal of Antimicrobial Agents
4324:
3690:
3688:
3352:
2707:
2460:
2382:
2335:
2333:
1844:
1650:Domingo E, Sheldon J, Perales C (June 2012).
1577:
1357:"Experimental evolution of plant RNA viruses"
6705:Crotty S, Cameron CE, Andino R (June 2001).
6646:
5247:
5094:
4953:
4918:
4779:
4735:Sequence Space and Quasispecies Distribution
4642:
4509:
4422:
4224:
4176:
3997:
3955:
3849:Perales C, Moreno E, Domingo E (July 2015).
3842:
3744:
3742:
3343:
3301:Sanjuán R, Domingo-Calap P (December 2016).
3238:
3147:
3145:
3143:
3141:
3139:
3080:
2748:
2710:"Exonucleolytic proofreading by p53 protein"
2230:
2188:
2152:
2104:
1885:
1743:
1614:
1573:
1571:
1569:
1488:
1350:
1348:
1292:
1290:
1288:
1225:"Neutral evolution of mutational robustness"
1213:
1007:, analogous to the way sexual organisms use
259:, as still implied today in the contents of
6822:
6658:"Examining the theory of error catastrophe"
6440:
5936:
5877:
5470:
5411:
5145:
4599:
4465:
4057:
2547:
1935:
1479:
571:, or that can alter the capacity to induce
6757:
6597:
6449:"Error catastrophe and antiviral strategy"
6320:
5521:
5203:Pfeiffer JK, Kirkegaard K (October 2005).
4726:
4691:
3911:Villarreal LP, Witzany G (November 2013).
3685:
3158:Microbiology and Molecular Biology Reviews
2376:
2330:
1692:
1656:Microbiology and Molecular Biology Reviews
141:, and of relevance to antiviral medicine.
86:. The theory provided a new definition of
6893:
6883:
6858:Sanjuán R, Moya A, Elena SF (June 2004).
6799:
6789:
6740:
6730:
6681:
6629:
6580:
6548:
6531:
6482:
6472:
6415:
6365:
6355:
6163:
6114:
6065:
5919:
5909:
5826:
5809:
5799:
5775:
5758:
5717:
5707:
5614:
5604:
5563:
5553:
5453:
5443:
5394:
5345:
5294:
5230:
5220:
5128:
5118:
5077:
5045:
5028:
4901:
4852:
4828:
4811:
4674:
4582:
4492:
4369:
4359:
4243:
4159:
4149:
4100:
4040:
3938:
3928:
3884:
3874:
3825:
3815:
3774:
3739:
3722:
3712:
3668:
3584:
3535:
3486:
3461:Vlok M, Lang AS, Suttle CA (April 2019).
3437:
3380:
3370:
3326:
3177:
3136:
3119:
3109:
3063:
3053:
3009:
2999:
2958:
2909:
2899:
2817:
2776:
2766:
2725:
2684:
2674:
2625:
2425:
2087:
1970:
1819:
1675:
1566:
1547:
1524:
1514:
1427:
1399:
1382:
1372:
1345:
1285:
1268:
1258:
1240:
1188:
1178:
1156:
1154:
1152:
1150:
1148:
1146:
1144:
1113:
1095:
717:Therefore, complementation, cooperation,
6255:
6131:
5682:Pfeiffer JK, Kirkegaard K (April 2006).
5580:
3151:
1850:
987:
978:
729:
442:
438:
352:
232:
159:
6554:
6298:. Springer New York. pp. 123–147.
6194:
4428:
4395:
3978:
3798:Tibayrenc M, Ayala FJ (November 2012).
2241:
967:reduces the infectivity of poliovirus.
840:Vaccines should include repertoires of
822:
6950:
5942:
2819:10.1146/annurev-virology-031413-085507
1489:Drake JW, Holland JJ (November 1999).
1180:10.1146/annurev-virology-091919-105900
1141:
786:purposes can mutate to new phenotypes.
617:
593:Behavior of reconstructed quasispecies
578:
169:) and the maximum average error rate p
6763:
6446:
3917:World Journal of Biological Chemistry
2339:
1698:
1450:
919:
864:> several expressed proteins >
6656:Summers J, Litwin S (January 2006).
6603:
6296:Handbook of Antimicrobial Resistance
5832:
3307:Cellular and Molecular Life Sciences
2478:Domingo E, Schuster P, eds. (2016).
876:Antiviral agents used in combination
803:Viral pathogenesis is influenced by
4475:Martín V, Domingo E (August 2008).
2749:Smith EC, Denison MR (2013-12-05).
870:multiple synthetic peptide antigens
294:human immunodeficiency virus type 1
13:
5387:10.1128/JVI.78.21.11678-11685.2004
3353:Lauring AS, Andino R (July 2010).
1491:"Mutation rates among RNA viruses"
975:Possible evolutionary consequences
954:
14:
6999:
6968:Externally peer reviewed articles
6935:
4443:10.1016/j.ijantimicag.2018.10.005
1453:Trends in Ecology & Evolution
54:The theory predicts that a viral
6910:
6816:
6555:Fox EJ, Loeb LA (October 2010).
5070:10.1128/jvi.75.16.7315-7320.2001
4804:10.1128/JVI.67.10.6071-6079.1993
4398:Genetic Diversity of RNA Viruses
4236:Genetic Diversity of RNA Viruses
2727:10.1046/j.1432-1327.2001.02075.x
2714:European Journal of Biochemistry
1054:
766:
670:lymphocytic choriomengitis virus
303:
213:. One of its corollaries is the
6825:Journal of Mathematical Biology
6573:10.1016/j.semcancer.2010.10.005
6107:10.1128/JVI.66.6.3557-3565.1992
6054:The Journal of General Virology
4894:10.1128/jvi.78.7.3319-3324.2004
4737:. CRC Press. pp. 211–245.
4733:Eigen M, Biebricher CK (1988).
4575:10.1128/jvi.74.8.3543-3547.2000
4481:Molecular Biology and Evolution
3636:
3601:
3552:
3503:
3454:
3397:
3294:
2975:
2926:
2875:
2834:
2793:
2742:
2701:
2642:
2593:
2512:
2419:
2055:
1987:
1963:10.1128/JVI.68.7.4295-4301.1994
1783:
1030:to give them a higher-fidelity
554:Non-consensus-based descriptors
541:Limitations and indeterminacies
113:(approx one error per round of
31:with a large number of variant
4559:"Memory in viral quasispecies"
3622:10.1093/bioinformatics/17.1.16
3479:10.1128/mspheredirect.00157-19
3303:"Mechanisms of viral mutation"
3152:Agol VI, Gmyl AP (June 2018).
2708:Bakhanashvili M (April 2001).
1792:Journal of Theoretical Biology
1652:"Viral quasispecies evolution"
1541:
1444:
1429:11858/00-001M-0000-002C-84A7-C
1021:
994:evolutionarily stable strategy
725:
209:to understand the response to
1:
4667:10.1016/s0022-2836(03)00661-2
4523:Briones C, Domingo E (2008).
3661:10.1093/bioinformatics/btx402
1408:Journal of Physical Chemistry
1049:
1026:Experimental manipulation of
369:because they lack a 3’ to 5’
363:RNA-dependent DNA polymerases
359:RNA-dependent RNA polymerases
207:intra-population interactions
6978:Microbial population biology
6015:Journal of Molecular Biology
5980:Journal of Molecular Biology
5957:10.1016/0027-5107(64)90047-8
5801:10.1371/journal.ppat.1002243
5760:10.1016/j.coviro.2012.08.001
5222:10.1371/journal.ppat.0010011
5120:10.1371/journal.ppat.0020136
5021:10.1016/j.coviro.2016.09.011
4927:Journal of Molecular Biology
4700:Journal of Molecular Biology
4655:Journal of Molecular Biology
4298:Journal of Molecular Biology
3979:Domingo E (September 2015).
3714:10.1371/journal.ppat.1004191
3372:10.1371/journal.ppat.1001005
3111:10.1371/journal.ppat.1007605
3001:10.1371/journal.ppat.1007368
2768:10.1371/journal.ppat.1003760
2571:10.1016/0092-8674(89)90883-0
2533:10.1016/0378-1119(92)90216-c
2308:10.1016/0092-8674(89)90942-2
2174:10.1016/0378-1119(85)90017-4
2041:10.1016/0378-1119(76)90004-4
1913:10.1016/0092-8674(78)90223-4
1769:10.1016/0301-4622(82)87037-3
1465:10.1016/0169-5347(92)90145-2
1097:10.1371/JOURNAL.PGEN.1008271
661:(PV) by attenuated virus in
535:foot-and-mouth disease virus
7:
6674:10.1128/JVI.80.1.20-26.2006
6304:10.1007/978-1-4939-0694-9_1
6205:10.1007/978-3-0348-9146-2_5
5747:Current Opinion in Virology
5009:Current Opinion in Virology
4245:10.1007/978-3-642-77011-1_1
4202:10.1016/j.virol.2018.07.030
3260:European Biophysics Journal
2080:10.1534/genetics.113.154963
1548:Le Clercq LS (2014-06-18).
848:epitopes to evoke an ample
629:memory in the immune system
485:genome segment reassortment
328:, bacterial populations or
10:
7004:
6561:Seminars in Cancer Biology
6396:Future Medicinal Chemistry
4608:Nature Reviews. Immunology
3767:10.1016/j.chom.2017.08.006
3565:Environmental Microbiology
3430:10.1038/s41598-017-02652-w
3042:Frontiers in Plant Science
1812:10.1016/j.jtbi.2018.10.007
668:Suppression of pathogenic
338:conformation distributions
248:vesicular stomatitis virus
155:
6197:Progress in Drug Research
4939:10.1016/j.jmb.2007.03.074
4712:10.1016/j.jmb.2004.03.061
4310:10.1016/j.jmb.2006.05.027
3319:10.1007/s00018-016-2299-6
3272:10.1007/s00249-018-1282-6
2806:Annual Review of Virology
2488:10.1007/978-3-319-23898-2
2342:Seminars in Liver Disease
1167:Annual Review of Virology
292:have been performed with
109:) because they have high
6988:Infraspecific virus taxa
6770:BMC Evolutionary Biology
6764:Wilke CO (August 2005).
6447:Eigen M (October 2002).
6270:10.1016/j.pt.2009.11.009
5534:BMC Evolutionary Biology
4743:10.1201/9781351076449-12
2260:10.1007/3-540-26397-7_14
1516:10.1073/pnas.96.24.13910
657:Suppression of virulent
479:and it can also include
326:evolution of tumor cells
193:organizations that link
6885:10.1073/pnas.0400146101
5911:10.1073/pnas.89.13.6015
5709:10.1073/pnas.0600834103
5555:10.1186/1471-2148-12-58
5499:10.1126/science.1115030
5445:10.1073/pnas.1323136111
4151:10.1073/pnas.0909787107
3876:10.1073/pnas.1501715112
3817:10.1073/pnas.1212452109
3755:Cell Host & Microbe
3586:10.1111/1462-2920.14481
3055:10.3389/fpls.2018.00932
2901:10.1073/pnas.1618714114
2385:Die Naturwissenschaften
2138:10.1126/science.7041255
2004:10.1002/352760801x.ch17
1701:Die Naturwissenschaften
1260:10.1073/pnas.96.17.9716
862:inactivated whole virus
624:complex adaptive system
565:neutralizing antibodies
386:Studies with different
336:, drug resistance, and
334:chromosomal instability
278:Newcastle disease virus
6791:10.1186/1471-2148-5-44
6732:10.1073/pnas.111085598
6610:Nature Reviews. Cancer
6474:10.1073/pnas.212514799
6357:10.1073/pnas.96.4.1492
6258:Trends in Parasitology
6027:10.1006/jmbi.1998.2366
5992:10.1006/jmbi.1996.0639
4854:10.1006/viro.1996.0460
3224:10.1002/chin.200338243
2676:10.1073/pnas.94.4.1113
2354:10.1055/s-0031-1297925
1374:10.1038/sj.hdy.6801088
984:
829:assist in the planning
735:
531:experimental evolution
501:biological constraints
456:
238:
174:
60:evolutionarily neutral
6604:Loeb LA (June 2011).
6524:10.1093/annonc/mdy003
6408:10.4155/fmc-2018-0457
6067:10.1099/vir.0.80126-0
4494:10.1093/molbev/msn099
4361:10.1073/pnas.88.1.199
3930:10.4331/wjbc.v4.i4.79
3170:10.1128/mmbr.00067-17
2214:10.1201/9781351076432
1757:Biophysical Chemistry
1668:10.1128/MMBR.05023-11
1016:mutational robustness
988:Mutational robustness
982:
866:one expressed protein
819:helping to confront.
733:
446:
439:Phenotypic reservoirs
353:Dynamic heterogeneity
236:
222:of the population in
211:selective constraints
163:
105:(including important
4093:10.1128/jvi.02505-16
3528:10.1128/jvi.00017-18
2951:10.1128/jvi.01335-16
2618:10.1128/jvi.01296-07
1078:"Viral quasispecies"
823:Antiviral strategies
607:receptor recognition
561:antiviral inhibitors
449:bottleneck transfers
415:uasispecies, PAQ or
322:RNA-RNA interactions
252:replicative capacity
243:RNA bacteriophage Qβ
25:population structure
6876:2004PNAS..101.8396S
6782:2005BMCEE...5...44W
6723:2001PNAS...98.6895C
6662:Journal of Virology
6465:2002PNAS...9913374E
6348:1999PNAS...96.1492L
6156:2015MolEc..24.6007H
6095:Journal of Virology
5902:1992PNAS...89.6015D
5847:1990Natur.348..454C
5700:2006PNAS..103.5520P
5651:10.1007/82_2015_461
5606:10.7554/elife.13974
5546:2012BMCEE..12...58C
5491:2006Sci...311..236A
5436:2014PNAS..111.6678M
5375:Journal of Virology
5328:10.1007/82_2015_483
5287:10.1038/nature04388
5279:2006Natur.439..344V
5169:10.1007/82_2015_463
5058:Journal of Virology
4882:Journal of Virology
4792:Journal of Virology
4620:10.1038/nri.2016.13
4563:Journal of Virology
4396:Holland JJ (1992).
4352:1991PNAS...88..199C
4142:2010PNAS..107.2108O
4081:Journal of Virology
4033:10.1038/nature12861
4025:2014Natur.505..686A
3867:2015PNAS..112.8887P
3577:2018EnvMi..20.4612G
3516:Journal of Virology
3422:2017NatSR...7.2368V
2939:Journal of Virology
2894:(22): E4442–E4451.
2851:10.1007/82_2015_456
2667:1997PNAS...94.1113N
2606:Journal of Virology
2436:10.1007/82_2015_471
2242:Holland JJ (2006).
2130:1982Sci...215.1577H
1951:Journal of Virology
1861:10.1007/82_2015_469
1804:2019JThBi.460..170F
1713:1971NW.....58..465E
1580:Naturwissenschaften
1507:1999PNAS...9613910D
1420:10.1021/j100335a010
1315:2001Natur.412..331W
1251:1999PNAS...96.9716V
663:poliovirus vaccines
618:Quasispecies memory
603:monoclonal antibody
579:Collective response
435:ference, QUENTIN).
346:population dynamics
310:genetic equilibrium
257:nucleotide sequence
125:, and may act as a
96:consensus sequences
74:in which primitive
6837:10.1007/BF00160372
6512:Annals of Oncology
3983:. Academic Press.
3410:Scientific Reports
2397:10.1007/bf00450633
1721:10.1007/bf00623322
1592:10.1007/bf00420631
1005:selection pressure
985:
925:Lethal mutagenesis
920:Lethal mutagenesis
736:
599:sweeping selection
457:
388:virus-host systems
318:fitness landscapes
286:Coxsackie A9 virus
284:and dependence in
239:
175:
150:adaptive processes
139:lethal mutagenesis
121:, competition and
92:adaptive potential
21:viral quasispecies
6402:(13): 1645–1657.
6165:10.1111/mec.13450
6144:Molecular Ecology
6060:(Pt 8): 2289–97.
5945:Mutation Research
3990:978-0-12-800837-9
3571:(12): 4612–4628.
3522:(12): e00017-18.
3313:(23): 4433–4448.
2497:978-3-319-23897-5
2124:(4540): 1577–85.
1414:(24): 6881–6891.
1014:However, whether
805:microevolutionary
748:hepatitis C virus
611:integrin receptor
597:In some cases of
569:cytotoxic T cells
298:hepatitis C virus
127:unit of selection
119:genetic variation
64:fitness landscape
45:viral replication
6995:
6929:
6928:
6927:
6926:
6914:
6908:
6907:
6897:
6887:
6870:(22): 8396–401.
6855:
6849:
6848:
6820:
6814:
6813:
6803:
6793:
6761:
6755:
6754:
6744:
6734:
6717:(12): 6895–900.
6702:
6696:
6695:
6685:
6653:
6644:
6643:
6633:
6601:
6595:
6594:
6584:
6552:
6546:
6545:
6535:
6503:
6497:
6496:
6486:
6476:
6444:
6438:
6437:
6419:
6391:
6380:
6379:
6369:
6359:
6327:
6318:
6317:
6291:
6282:
6281:
6253:
6247:
6246:
6240:
6236:
6234:
6226:
6192:
6186:
6185:
6167:
6135:
6129:
6128:
6118:
6086:
6080:
6079:
6069:
6045:
6039:
6038:
6010:
6004:
6003:
5975:
5969:
5968:
5940:
5934:
5933:
5923:
5913:
5881:
5875:
5874:
5855:10.1038/348454a0
5830:
5824:
5823:
5813:
5803:
5779:
5773:
5772:
5762:
5738:
5732:
5731:
5721:
5711:
5679:
5673:
5672:
5638:
5629:
5628:
5618:
5608:
5584:
5578:
5577:
5567:
5557:
5525:
5519:
5518:
5474:
5468:
5467:
5457:
5447:
5415:
5409:
5408:
5398:
5381:(21): 11678–85.
5366:
5360:
5359:
5349:
5315:
5309:
5308:
5298:
5258:
5245:
5244:
5234:
5224:
5200:
5191:
5190:
5156:
5143:
5142:
5132:
5122:
5098:
5092:
5091:
5081:
5049:
5043:
5042:
5032:
5000:
4994:
4993:
4957:
4951:
4950:
4922:
4916:
4915:
4905:
4873:
4867:
4866:
4856:
4832:
4826:
4825:
4815:
4783:
4777:
4776:
4770:
4766:
4764:
4756:
4730:
4724:
4723:
4695:
4689:
4688:
4678:
4646:
4640:
4639:
4603:
4597:
4596:
4586:
4554:
4545:
4544:
4520:
4507:
4506:
4496:
4472:
4463:
4462:
4426:
4420:
4419:
4393:
4384:
4383:
4373:
4363:
4331:
4322:
4321:
4293:
4284:
4283:
4247:
4231:
4222:
4221:
4185:
4174:
4173:
4163:
4153:
4121:
4115:
4114:
4104:
4072:
4055:
4054:
4044:
4019:(7485): 686–90.
4004:
3995:
3994:
3976:
3953:
3952:
3942:
3932:
3908:
3899:
3898:
3888:
3878:
3846:
3840:
3839:
3829:
3819:
3810:(48): E3305-13.
3795:
3789:
3788:
3778:
3746:
3737:
3736:
3726:
3716:
3692:
3683:
3682:
3672:
3640:
3634:
3633:
3605:
3599:
3598:
3588:
3556:
3550:
3549:
3539:
3507:
3501:
3500:
3490:
3473:(2): e00157-19.
3458:
3452:
3451:
3441:
3401:
3395:
3394:
3384:
3374:
3350:
3341:
3340:
3330:
3298:
3292:
3291:
3255:
3236:
3235:
3207:
3192:
3191:
3181:
3149:
3134:
3133:
3123:
3113:
3089:
3078:
3077:
3067:
3057:
3033:
3024:
3023:
3013:
3003:
2994:(10): e1007368.
2979:
2973:
2972:
2962:
2930:
2924:
2923:
2913:
2903:
2879:
2873:
2872:
2838:
2832:
2831:
2821:
2797:
2791:
2790:
2780:
2770:
2761:(12): e1003760.
2746:
2740:
2739:
2729:
2705:
2699:
2698:
2688:
2678:
2646:
2640:
2639:
2629:
2612:(22): 12135–44.
2597:
2591:
2590:
2554:
2545:
2544:
2516:
2510:
2509:
2475:
2458:
2457:
2423:
2417:
2416:
2380:
2374:
2373:
2337:
2328:
2327:
2291:
2282:
2281:
2239:
2228:
2227:
2201:
2186:
2185:
2156:
2150:
2149:
2113:
2102:
2101:
2091:
2059:
2053:
2052:
2024:
2018:
2017:
1991:
1985:
1984:
1974:
1942:
1933:
1932:
1896:
1883:
1882:
1848:
1842:
1841:
1823:
1787:
1781:
1780:
1752:
1741:
1740:
1696:
1690:
1689:
1679:
1647:
1612:
1611:
1575:
1564:
1563:
1545:
1539:
1538:
1528:
1518:
1486:
1477:
1476:
1448:
1442:
1441:
1431:
1403:
1397:
1396:
1386:
1376:
1352:
1343:
1342:
1323:10.1038/35085569
1294:
1283:
1282:
1272:
1262:
1244:
1242:adap-org/9903006
1220:
1211:
1210:
1192:
1182:
1158:
1136:
1135:
1117:
1099:
1090:(10): e1008271.
1072:reviewer reports
1065:
1058:
900:Combined use of
744:Müller’s ratchet
135:virus extinction
7003:
7002:
6998:
6997:
6996:
6994:
6993:
6992:
6948:
6947:
6938:
6933:
6932:
6924:
6922:
6916:
6915:
6911:
6856:
6852:
6821:
6817:
6762:
6758:
6703:
6699:
6654:
6647:
6622:10.1038/nrc3063
6602:
6598:
6553:
6549:
6504:
6500:
6459:(21): 13374–6.
6445:
6441:
6392:
6383:
6328:
6321:
6314:
6292:
6285:
6254:
6250:
6238:
6237:
6228:
6227:
6215:
6193:
6189:
6150:(24): 6007–12.
6136:
6132:
6087:
6083:
6046:
6042:
6011:
6007:
5976:
5972:
5941:
5937:
5882:
5878:
5841:(6300): 454–5.
5831:
5827:
5794:(9): e1002243.
5780:
5776:
5739:
5735:
5680:
5676:
5661:
5639:
5632:
5585:
5581:
5526:
5522:
5485:(5758): 236–8.
5475:
5471:
5430:(18): 6678–83.
5416:
5412:
5367:
5363:
5338:
5316:
5312:
5273:(7074): 344–8.
5259:
5248:
5201:
5194:
5179:
5157:
5146:
5099:
5095:
5064:(16): 7315–20.
5050:
5046:
5001:
4997:
4962:Nature Genetics
4958:
4954:
4933:(4): 985–1000.
4923:
4919:
4874:
4870:
4833:
4829:
4784:
4780:
4768:
4767:
4758:
4757:
4753:
4731:
4727:
4696:
4692:
4647:
4643:
4604:
4600:
4555:
4548:
4521:
4510:
4473:
4466:
4427:
4423:
4408:
4394:
4387:
4332:
4325:
4294:
4287:
4256:
4232:
4225:
4186:
4177:
4122:
4118:
4073:
4058:
4005:
3998:
3991:
3977:
3956:
3909:
3902:
3861:(29): 8887–92.
3847:
3843:
3796:
3792:
3747:
3740:
3707:(6): e1004191.
3693:
3686:
3641:
3637:
3606:
3602:
3557:
3553:
3508:
3504:
3459:
3455:
3402:
3398:
3365:(7): e1001005.
3351:
3344:
3299:
3295:
3256:
3239:
3208:
3195:
3164:(2): e00067-1.
3150:
3137:
3104:(2): e1007605.
3090:
3081:
3034:
3027:
2980:
2976:
2931:
2927:
2880:
2876:
2861:
2839:
2835:
2798:
2794:
2747:
2743:
2706:
2702:
2647:
2643:
2598:
2594:
2555:
2548:
2517:
2513:
2498:
2476:
2461:
2446:
2424:
2420:
2381:
2377:
2338:
2331:
2292:
2285:
2270:
2240:
2231:
2224:
2202:
2189:
2157:
2153:
2114:
2105:
2060:
2056:
2025:
2021:
2014:
1992:
1988:
1957:(7): 4295–301.
1943:
1936:
1897:
1886:
1871:
1849:
1845:
1788:
1784:
1753:
1744:
1707:(10): 465–523.
1697:
1693:
1648:
1615:
1576:
1567:
1546:
1542:
1501:(24): 13910–3.
1487:
1480:
1449:
1445:
1404:
1400:
1353:
1346:
1309:(6844): 331–3.
1295:
1286:
1235:(17): 9716–20.
1221:
1214:
1159:
1142:
1075:
1061:
1059:
1052:
1024:
990:
977:
957:
955:Error threshold
922:
878:
850:immune response
838:
825:
769:
728:
696:fitness plateau
687:complementation
647:
620:
595:
590:
581:
556:
543:
526:virus isolation
510:gene expression
497:virus evolution
477:point mutations
441:
355:
314:population size
306:
282:drug resistance
246:documented for
215:error threshold
172:
168:
158:
146:deep sequencing
17:
12:
11:
5:
7001:
6991:
6990:
6985:
6980:
6975:
6970:
6965:
6960:
6946:
6945:
6937:
6936:External links
6934:
6931:
6930:
6909:
6850:
6815:
6756:
6697:
6645:
6596:
6547:
6518:(3): 563–572.
6498:
6439:
6381:
6319:
6312:
6283:
6248:
6239:|journal=
6213:
6187:
6130:
6101:(6): 3557–65.
6081:
6040:
6021:(2): 495–505.
6005:
5970:
5935:
5896:(13): 6015–9.
5876:
5825:
5788:PLOS Pathogens
5774:
5733:
5694:(14): 5520–5.
5674:
5659:
5630:
5579:
5520:
5469:
5410:
5361:
5336:
5310:
5246:
5209:PLOS Pathogens
5192:
5177:
5144:
5107:PLOS Pathogens
5093:
5044:
4995:
4974:10.1038/ng1583
4952:
4917:
4888:(7): 3319–24.
4868:
4827:
4798:(10): 6071–9.
4778:
4769:|journal=
4751:
4725:
4690:
4641:
4598:
4546:
4508:
4487:(8): 1544–54.
4464:
4421:
4406:
4385:
4346:(1): 199–203.
4323:
4285:
4254:
4223:
4175:
4136:(5): 2108–12.
4116:
4056:
3996:
3989:
3954:
3900:
3841:
3790:
3738:
3701:PLOS Pathogens
3684:
3655:(1): 163–170.
3649:Bioinformatics
3635:
3610:Bioinformatics
3600:
3551:
3502:
3453:
3396:
3359:PLOS Pathogens
3342:
3293:
3266:(4): 443–457.
3237:
3193:
3135:
3098:PLOS Pathogens
3079:
3025:
2988:PLOS Pathogens
2974:
2925:
2874:
2859:
2833:
2792:
2755:PLOS Pathogens
2741:
2720:(7): 2047–54.
2700:
2641:
2592:
2546:
2511:
2496:
2459:
2444:
2418:
2391:(11): 541–65.
2375:
2329:
2283:
2268:
2229:
2222:
2187:
2151:
2103:
2054:
2019:
2012:
1986:
1934:
1884:
1869:
1843:
1782:
1742:
1691:
1662:(2): 159–216.
1613:
1565:
1540:
1478:
1443:
1398:
1344:
1284:
1212:
1139:
1138:
1051:
1048:
1023:
1020:
989:
986:
976:
973:
956:
953:
921:
918:
913:
912:
909:
898:
895:
892:
889:
877:
874:
837:
834:
824:
821:
816:
815:
811:
808:
801:
790:
787:
780:
768:
765:
727:
724:
704:5-fluorouracil
683:
682:
679:
676:
673:
666:
655:
646:
643:
619:
616:
594:
591:
589:
586:
580:
577:
555:
552:
542:
539:
440:
437:
354:
351:
305:
302:
276:morphology of
224:sequence space
183:Peter Schuster
170:
166:
157:
154:
111:mutation rates
72:origin of life
37:mutation rates
15:
9:
6:
4:
3:
2:
7000:
6989:
6986:
6984:
6981:
6979:
6976:
6974:
6971:
6969:
6966:
6964:
6961:
6959:
6956:
6955:
6953:
6943:
6940:
6939:
6921:
6920:
6913:
6905:
6901:
6896:
6891:
6886:
6881:
6877:
6873:
6869:
6865:
6861:
6854:
6846:
6842:
6838:
6834:
6830:
6826:
6819:
6811:
6807:
6802:
6797:
6792:
6787:
6783:
6779:
6775:
6771:
6767:
6760:
6752:
6748:
6743:
6738:
6733:
6728:
6724:
6720:
6716:
6712:
6708:
6701:
6693:
6689:
6684:
6679:
6675:
6671:
6667:
6663:
6659:
6652:
6650:
6641:
6637:
6632:
6627:
6623:
6619:
6615:
6611:
6607:
6600:
6592:
6588:
6583:
6578:
6574:
6570:
6566:
6562:
6558:
6551:
6543:
6539:
6534:
6529:
6525:
6521:
6517:
6513:
6509:
6502:
6494:
6490:
6485:
6480:
6475:
6470:
6466:
6462:
6458:
6454:
6450:
6443:
6435:
6431:
6427:
6423:
6418:
6413:
6409:
6405:
6401:
6397:
6390:
6388:
6386:
6377:
6373:
6368:
6363:
6358:
6353:
6349:
6345:
6342:(4): 1492–7.
6341:
6337:
6333:
6326:
6324:
6315:
6313:9781493906932
6309:
6305:
6301:
6297:
6290:
6288:
6279:
6275:
6271:
6267:
6263:
6259:
6252:
6244:
6232:
6224:
6220:
6216:
6214:9783034899253
6210:
6206:
6202:
6198:
6191:
6183:
6179:
6175:
6171:
6166:
6161:
6157:
6153:
6149:
6145:
6141:
6134:
6126:
6122:
6117:
6112:
6108:
6104:
6100:
6096:
6092:
6085:
6077:
6073:
6068:
6063:
6059:
6055:
6051:
6044:
6036:
6032:
6028:
6024:
6020:
6016:
6009:
6001:
5997:
5993:
5989:
5986:(2): 255–67.
5985:
5981:
5974:
5966:
5962:
5958:
5954:
5950:
5946:
5939:
5931:
5927:
5922:
5917:
5912:
5907:
5903:
5899:
5895:
5891:
5887:
5880:
5872:
5868:
5864:
5860:
5856:
5852:
5848:
5844:
5840:
5836:
5829:
5821:
5817:
5812:
5807:
5802:
5797:
5793:
5789:
5785:
5778:
5770:
5766:
5761:
5756:
5753:(5): 546–55.
5752:
5748:
5744:
5737:
5729:
5725:
5720:
5715:
5710:
5705:
5701:
5697:
5693:
5689:
5685:
5678:
5670:
5666:
5662:
5660:9783319238975
5656:
5652:
5648:
5644:
5637:
5635:
5626:
5622:
5617:
5612:
5607:
5602:
5598:
5594:
5590:
5583:
5575:
5571:
5566:
5561:
5556:
5551:
5547:
5543:
5539:
5535:
5531:
5524:
5516:
5512:
5508:
5504:
5500:
5496:
5492:
5488:
5484:
5480:
5473:
5465:
5461:
5456:
5451:
5446:
5441:
5437:
5433:
5429:
5425:
5421:
5414:
5406:
5402:
5397:
5392:
5388:
5384:
5380:
5376:
5372:
5365:
5357:
5353:
5348:
5343:
5339:
5337:9783319238982
5333:
5329:
5325:
5321:
5314:
5306:
5302:
5297:
5292:
5288:
5284:
5280:
5276:
5272:
5268:
5264:
5257:
5255:
5253:
5251:
5242:
5238:
5233:
5228:
5223:
5218:
5214:
5210:
5206:
5199:
5197:
5188:
5184:
5180:
5178:9783319238975
5174:
5170:
5166:
5162:
5155:
5153:
5151:
5149:
5140:
5136:
5131:
5126:
5121:
5116:
5112:
5108:
5104:
5097:
5089:
5085:
5080:
5075:
5071:
5067:
5063:
5059:
5055:
5048:
5040:
5036:
5031:
5026:
5022:
5018:
5014:
5010:
5006:
4999:
4991:
4987:
4983:
4979:
4975:
4971:
4967:
4963:
4956:
4948:
4944:
4940:
4936:
4932:
4928:
4921:
4913:
4909:
4904:
4899:
4895:
4891:
4887:
4883:
4879:
4872:
4864:
4860:
4855:
4850:
4846:
4842:
4838:
4831:
4823:
4819:
4814:
4809:
4805:
4801:
4797:
4793:
4789:
4782:
4774:
4762:
4754:
4752:9781351076449
4748:
4744:
4740:
4736:
4729:
4721:
4717:
4713:
4709:
4706:(2): 405–12.
4705:
4701:
4694:
4686:
4682:
4677:
4672:
4668:
4664:
4661:(1): 213–29.
4660:
4656:
4652:
4645:
4637:
4633:
4629:
4625:
4621:
4617:
4613:
4609:
4602:
4594:
4590:
4585:
4580:
4576:
4572:
4569:(8): 3543–7.
4568:
4564:
4560:
4553:
4551:
4542:
4538:
4535:(2): 93–109.
4534:
4530:
4526:
4519:
4517:
4515:
4513:
4504:
4500:
4495:
4490:
4486:
4482:
4478:
4471:
4469:
4460:
4456:
4452:
4448:
4444:
4440:
4437:(1): 105562.
4436:
4432:
4425:
4417:
4413:
4409:
4407:9783642770111
4403:
4399:
4392:
4390:
4381:
4377:
4372:
4367:
4362:
4357:
4353:
4349:
4345:
4341:
4337:
4330:
4328:
4319:
4315:
4311:
4307:
4304:(3): 558–72.
4303:
4299:
4292:
4290:
4281:
4277:
4273:
4269:
4265:
4261:
4257:
4255:9783642770111
4251:
4246:
4241:
4237:
4230:
4228:
4219:
4215:
4211:
4207:
4203:
4199:
4195:
4191:
4184:
4182:
4180:
4171:
4167:
4162:
4157:
4152:
4147:
4143:
4139:
4135:
4131:
4127:
4120:
4112:
4108:
4103:
4098:
4094:
4090:
4086:
4082:
4078:
4071:
4069:
4067:
4065:
4063:
4061:
4052:
4048:
4043:
4038:
4034:
4030:
4026:
4022:
4018:
4014:
4010:
4003:
4001:
3992:
3986:
3982:
3975:
3973:
3971:
3969:
3967:
3965:
3963:
3961:
3959:
3950:
3946:
3941:
3936:
3931:
3926:
3922:
3918:
3914:
3907:
3905:
3896:
3892:
3887:
3882:
3877:
3872:
3868:
3864:
3860:
3856:
3852:
3845:
3837:
3833:
3828:
3823:
3818:
3813:
3809:
3805:
3801:
3794:
3786:
3782:
3777:
3772:
3768:
3764:
3760:
3756:
3752:
3745:
3743:
3734:
3730:
3725:
3720:
3715:
3710:
3706:
3702:
3698:
3691:
3689:
3680:
3676:
3671:
3666:
3662:
3658:
3654:
3650:
3646:
3639:
3631:
3627:
3623:
3619:
3615:
3611:
3604:
3596:
3592:
3587:
3582:
3578:
3574:
3570:
3566:
3562:
3555:
3547:
3543:
3538:
3533:
3529:
3525:
3521:
3517:
3513:
3506:
3498:
3494:
3489:
3484:
3480:
3476:
3472:
3468:
3464:
3457:
3449:
3445:
3440:
3435:
3431:
3427:
3423:
3419:
3415:
3411:
3407:
3400:
3392:
3388:
3383:
3378:
3373:
3368:
3364:
3360:
3356:
3349:
3347:
3338:
3334:
3329:
3324:
3320:
3316:
3312:
3308:
3304:
3297:
3289:
3285:
3281:
3277:
3273:
3269:
3265:
3261:
3254:
3252:
3250:
3248:
3246:
3244:
3242:
3233:
3229:
3225:
3221:
3217:
3213:
3206:
3204:
3202:
3200:
3198:
3189:
3185:
3180:
3175:
3171:
3167:
3163:
3159:
3155:
3148:
3146:
3144:
3142:
3140:
3131:
3127:
3122:
3117:
3112:
3107:
3103:
3099:
3095:
3088:
3086:
3084:
3075:
3071:
3066:
3061:
3056:
3051:
3047:
3043:
3039:
3032:
3030:
3021:
3017:
3012:
3007:
3002:
2997:
2993:
2989:
2985:
2978:
2970:
2966:
2961:
2956:
2952:
2948:
2944:
2940:
2936:
2929:
2921:
2917:
2912:
2907:
2902:
2897:
2893:
2889:
2885:
2878:
2870:
2866:
2862:
2860:9783319238975
2856:
2852:
2848:
2844:
2837:
2829:
2825:
2820:
2815:
2812:(1): 111–32.
2811:
2807:
2803:
2796:
2788:
2784:
2779:
2774:
2769:
2764:
2760:
2756:
2752:
2745:
2737:
2733:
2728:
2723:
2719:
2715:
2711:
2704:
2696:
2692:
2687:
2682:
2677:
2672:
2668:
2664:
2661:(4): 1113–8.
2660:
2656:
2652:
2645:
2637:
2633:
2628:
2623:
2619:
2615:
2611:
2607:
2603:
2596:
2588:
2584:
2580:
2576:
2572:
2568:
2565:(1): 219–28.
2564:
2560:
2553:
2551:
2542:
2538:
2534:
2530:
2526:
2522:
2515:
2507:
2503:
2499:
2493:
2489:
2485:
2481:
2474:
2472:
2470:
2468:
2466:
2464:
2455:
2451:
2447:
2445:9783319238975
2441:
2437:
2433:
2429:
2422:
2414:
2410:
2406:
2402:
2398:
2394:
2390:
2386:
2379:
2371:
2367:
2363:
2359:
2355:
2351:
2348:(4): 356–74.
2347:
2343:
2336:
2334:
2325:
2321:
2317:
2313:
2309:
2305:
2302:(5): 901–10.
2301:
2297:
2290:
2288:
2279:
2275:
2271:
2265:
2261:
2257:
2253:
2249:
2245:
2238:
2236:
2234:
2225:
2223:9781351076432
2219:
2215:
2211:
2207:
2200:
2198:
2196:
2194:
2192:
2183:
2179:
2175:
2171:
2167:
2163:
2155:
2147:
2143:
2139:
2135:
2131:
2127:
2123:
2119:
2112:
2110:
2108:
2099:
2095:
2090:
2085:
2081:
2077:
2074:(1): 243–51.
2073:
2069:
2065:
2058:
2050:
2046:
2042:
2038:
2034:
2030:
2023:
2015:
2013:9783527608010
2009:
2005:
2001:
1997:
1996:Pathogenomics
1990:
1982:
1978:
1973:
1968:
1964:
1960:
1956:
1952:
1948:
1941:
1939:
1930:
1926:
1922:
1918:
1914:
1910:
1907:(4): 735–44.
1906:
1902:
1895:
1893:
1891:
1889:
1880:
1876:
1872:
1870:9783319238975
1866:
1862:
1858:
1854:
1847:
1839:
1835:
1831:
1827:
1822:
1817:
1813:
1809:
1805:
1801:
1797:
1793:
1786:
1778:
1774:
1770:
1766:
1763:(4): 329–45.
1762:
1758:
1751:
1749:
1747:
1738:
1734:
1730:
1726:
1722:
1718:
1714:
1710:
1706:
1702:
1695:
1687:
1683:
1678:
1673:
1669:
1665:
1661:
1657:
1653:
1646:
1644:
1642:
1640:
1638:
1636:
1634:
1632:
1630:
1628:
1626:
1624:
1622:
1620:
1618:
1609:
1605:
1601:
1597:
1593:
1589:
1585:
1581:
1574:
1572:
1570:
1561:
1557:
1553:
1552:
1544:
1536:
1532:
1527:
1522:
1517:
1512:
1508:
1504:
1500:
1496:
1492:
1485:
1483:
1474:
1470:
1466:
1462:
1459:(4): 118–21.
1458:
1454:
1447:
1439:
1435:
1430:
1425:
1421:
1417:
1413:
1409:
1402:
1394:
1390:
1385:
1380:
1375:
1370:
1367:(5): 478–83.
1366:
1362:
1358:
1351:
1349:
1340:
1336:
1332:
1328:
1324:
1320:
1316:
1312:
1308:
1304:
1300:
1293:
1291:
1289:
1280:
1276:
1271:
1266:
1261:
1256:
1252:
1248:
1243:
1238:
1234:
1230:
1226:
1219:
1217:
1208:
1204:
1200:
1196:
1191:
1186:
1181:
1176:
1172:
1168:
1164:
1157:
1155:
1153:
1151:
1149:
1147:
1145:
1140:
1137:
1133:
1129:
1125:
1121:
1116:
1111:
1107:
1103:
1098:
1093:
1089:
1085:
1084:
1083:PLOS Genetics
1079:
1073:
1069:
1064:
1057:
1047:
1045:
1041:
1037:
1036:pathogenicity
1033:
1029:
1019:
1017:
1012:
1010:
1009:recombination
1006:
1001:
999:
995:
981:
972:
968:
966:
960:
952:
950:
946:
942:
937:
935:
931:
926:
917:
910:
907:
903:
902:immunotherapy
899:
896:
893:
890:
887:
886:
885:
883:
873:
871:
867:
863:
859:
855:
851:
847:
843:
833:
830:
820:
812:
809:
806:
802:
799:
795:
791:
788:
785:
781:
778:
774:
773:
772:
767:Viral disease
764:
760:
758:
753:
749:
745:
740:
732:
723:
720:
715:
711:
707:
705:
699:
697:
692:
688:
680:
677:
674:
671:
667:
664:
660:
656:
653:
652:
651:
642:
639:
635:
630:
625:
615:
612:
608:
604:
600:
585:
576:
574:
570:
566:
562:
551:
547:
538:
536:
532:
527:
522:
519:
515:
511:
506:
502:
498:
494:
490:
486:
482:
481:recombination
478:
474:
470:
466:
462:
454:
450:
445:
436:
434:
430:
427:etwork-based
426:
422:
418:
414:
410:
406:
401:
400:Bioinformatic
397:
394:
389:
384:
381:
377:
376:coronaviruses
372:
368:
364:
360:
350:
347:
343:
339:
335:
331:
327:
323:
319:
315:
311:
304:Current scope
301:
299:
295:
291:
287:
283:
279:
275:
269:
266:
262:
258:
253:
249:
244:
235:
231:
227:
225:
221:
216:
212:
208:
204:
200:
196:
192:
188:
184:
180:
179:Manfred Eigen
162:
153:
151:
147:
142:
140:
136:
132:
128:
124:
120:
116:
112:
108:
104:
99:
97:
93:
89:
85:
82:within their
81:
77:
73:
68:
65:
61:
58:at a low but
57:
52:
50:
46:
42:
38:
34:
30:
26:
22:
6923:, retrieved
6918:
6912:
6867:
6863:
6853:
6831:(1): 33–44.
6828:
6824:
6818:
6773:
6769:
6759:
6714:
6710:
6700:
6665:
6661:
6616:(6): 450–7.
6613:
6609:
6599:
6567:(5): 353–9.
6564:
6560:
6550:
6515:
6511:
6501:
6456:
6452:
6442:
6417:10261/216260
6399:
6395:
6339:
6335:
6295:
6264:(2): 83–92.
6261:
6257:
6251:
6196:
6190:
6147:
6143:
6133:
6098:
6094:
6084:
6057:
6053:
6043:
6018:
6014:
6008:
5983:
5979:
5973:
5948:
5944:
5938:
5893:
5889:
5879:
5838:
5834:
5828:
5791:
5787:
5777:
5750:
5746:
5736:
5691:
5687:
5677:
5642:
5596:
5592:
5582:
5537:
5533:
5523:
5482:
5478:
5472:
5427:
5423:
5413:
5378:
5374:
5364:
5319:
5313:
5270:
5266:
5212:
5208:
5160:
5113:(12): e136.
5110:
5106:
5096:
5061:
5057:
5047:
5012:
5008:
4998:
4968:(7): 701–9.
4965:
4961:
4955:
4930:
4926:
4920:
4885:
4881:
4871:
4847:(1): 113–9.
4844:
4840:
4830:
4795:
4791:
4781:
4734:
4728:
4703:
4699:
4693:
4658:
4654:
4644:
4614:(2): 124–8.
4611:
4607:
4601:
4566:
4562:
4532:
4529:AIDS Reviews
4528:
4484:
4480:
4434:
4430:
4424:
4397:
4343:
4339:
4301:
4297:
4235:
4193:
4189:
4133:
4129:
4119:
4084:
4080:
4016:
4012:
3980:
3923:(4): 79–90.
3920:
3916:
3858:
3854:
3844:
3807:
3803:
3793:
3758:
3754:
3704:
3700:
3652:
3648:
3638:
3616:(1): 16–22.
3613:
3609:
3603:
3568:
3564:
3554:
3519:
3515:
3505:
3470:
3466:
3456:
3413:
3409:
3399:
3362:
3358:
3310:
3306:
3296:
3263:
3259:
3215:
3211:
3161:
3157:
3101:
3097:
3045:
3041:
2991:
2987:
2977:
2942:
2938:
2928:
2891:
2887:
2877:
2842:
2836:
2809:
2805:
2795:
2758:
2754:
2744:
2717:
2713:
2703:
2658:
2654:
2644:
2609:
2605:
2595:
2562:
2558:
2527:(2): 281–8.
2524:
2520:
2514:
2479:
2427:
2421:
2388:
2384:
2378:
2345:
2341:
2299:
2295:
2247:
2206:RNA Genetics
2205:
2165:
2161:
2154:
2121:
2117:
2071:
2067:
2057:
2035:(1): 27–32.
2032:
2028:
2022:
1995:
1989:
1954:
1950:
1904:
1900:
1852:
1846:
1795:
1791:
1785:
1760:
1756:
1704:
1700:
1694:
1659:
1655:
1583:
1579:
1550:
1543:
1498:
1494:
1456:
1452:
1446:
1411:
1407:
1401:
1364:
1360:
1306:
1302:
1232:
1228:
1190:10261/265663
1173:(1): 51–72.
1170:
1166:
1087:
1081:
1053:
1044:adaptability
1025:
1013:
1002:
991:
969:
961:
958:
938:
923:
914:
906:chemotherapy
879:
839:
826:
817:
794:cell tropism
784:pest control
770:
761:
752:BHK-21 cells
741:
737:
719:interference
716:
712:
708:
700:
684:
648:
633:
621:
596:
582:
557:
548:
544:
523:
505:host defense
469:pluripotency
463:but also of
458:
452:
432:
431:ransmission
428:
424:
420:
416:
412:
408:
404:
392:
385:
367:proofreading
361:(RdRps) and
356:
307:
296:(HIV-1) and
289:
270:
240:
228:
176:
143:
115:replication)
100:
69:
56:quasispecies
53:
20:
18:
6668:(1): 20–6.
5015:: 106–111.
4196:: 100–109.
3416:(1): 2368.
1821:2072/378023
1798:: 170–183.
1586:(1): 7–41.
1022:Cooperation
930:favipiravir
882:monotherapy
777:vaccination
757:attenuation
726:Bottlenecks
691:cooperation
489:enterovirus
419:asispecies
411:nalysis of
371:exonuclease
250:(VSV). The
191:hypercyclic
131:upper limit
103:RNA viruses
80:RNA viruses
6952:Categories
6925:2019-10-22
5951:(1): 2–9.
5599:: e13974.
5215:(2): e11.
3761:(3): 420.
3212:ChemInform
2269:3540263950
2168:(1): 1–8.
1050:References
1032:polymerase
1028:poliovirus
880:Antiviral
858:attenuated
798:host range
659:poliovirus
638:bottleneck
573:interferon
514:entry into
465:phenotypic
423:volution,
396:infections
380:retroviral
330:stem cells
261:data banks
199:phenotypic
51:proceeds.
6845:122142345
6776:(1): 44.
6434:201673871
6241:ignored (
6231:cite book
5540:(1): 58.
4771:ignored (
4761:cite book
4416:851813241
4264:851813241
3232:0931-7597
2370:260313185
1600:0028-1042
1560:958495192
1199:2327-056X
1132:Q86320171
1106:1553-7390
1066:license (
1063:CC BY 4.0
1040:wild-type
965:ribavirin
934:ribavirin
518:exit from
493:virulence
461:genotypic
407:artition
265:tolerance
195:genotypic
187:replicons
123:selection
107:pathogens
88:wild type
76:replicons
49:selection
41:frequency
6983:Virology
6904:15159545
6810:16107214
6751:11371613
6692:16352527
6640:21593786
6591:20934515
6542:29324969
6493:12370416
6426:31469331
6278:20036799
6182:14433111
6174:26769402
6076:15269370
5965:14195748
5820:21912520
5769:22921636
5728:16567621
5669:26162566
5625:26978794
5574:22541042
5515:21605827
5507:16410525
5464:24757055
5405:15479809
5356:26499340
5305:16327776
5241:16220146
5187:26210988
5139:17196038
5088:11462003
5039:27764731
4990:16501575
4982:15965477
4947:17481660
4912:15016853
4841:Virology
4720:15136042
4685:12875847
4628:26831526
4593:10729128
4541:18615120
4503:18436553
4459:52980171
4451:30315919
4318:16797586
4280:46530529
4218:52006443
4210:30107298
4190:Virology
4170:20080701
4111:28275194
4051:24284629
3949:24340131
3895:26195777
3836:22949662
3785:28910639
3733:24945141
3679:29304222
3630:11222259
3595:30452113
3546:29593040
3497:30944212
3448:28539654
3391:20661479
3337:27392606
3288:13239618
3280:29397419
3188:29540453
3130:30768648
3074:30013589
3020:30335851
2969:27795410
2920:28416670
2869:26373410
2828:26958717
2787:24348241
2736:11277927
2636:17804504
2587:23203156
2506:19514308
2454:26342705
2413:42131267
2362:22189976
2324:35903181
2278:16568907
2098:23852383
2068:Genetics
1929:19109157
1879:26597856
1838:52947045
1830:30300648
1737:38296619
1686:22688811
1535:10570172
1473:21235976
1438:96727272
1393:18253158
1361:Heredity
1331:11460163
1279:10449760
1207:34586874
1128:Wikidata
1124:31622336
998:genotype
6872:Bibcode
6801:1208876
6778:Bibcode
6719:Bibcode
6683:1317512
6631:4007007
6582:3256989
6533:5888943
6461:Bibcode
6376:9990051
6344:Bibcode
6223:2687948
6152:Bibcode
6125:1316467
6035:9878424
6000:8951375
5930:1321432
5898:Bibcode
5871:4235839
5863:2247152
5843:Bibcode
5811:3164670
5719:1414638
5696:Bibcode
5616:4805539
5565:3358237
5542:Bibcode
5487:Bibcode
5479:Science
5455:4020086
5432:Bibcode
5347:7121553
5296:1569948
5275:Bibcode
5232:1250929
5130:1757203
5030:5298929
4863:8806545
4822:7690417
4676:7173031
4636:8000538
4380:1846038
4348:Bibcode
4272:1600747
4161:2836666
4138:Bibcode
4102:5411618
4042:4111796
4021:Bibcode
3940:3856310
3886:4517279
3863:Bibcode
3827:3511763
3776:5807061
3724:4055744
3670:6355096
3573:Bibcode
3537:5974501
3488:6449609
3467:mSphere
3439:5443807
3418:Bibcode
3382:2908548
3328:5075021
3179:5968460
3121:6395005
3065:6036239
3048:: 932.
3011:6207329
2960:5165223
2911:5465912
2778:3857799
2695:9037015
2663:Bibcode
2627:2169014
2579:2790959
2541:1336756
2316:2550139
2252:371–401
2182:3912262
2146:7041255
2126:Bibcode
2118:Science
2089:3761305
2049:1052321
1981:8207804
1800:Bibcode
1777:7159681
1729:4942363
1709:Bibcode
1677:3372249
1608:1812273
1503:Bibcode
1384:7094686
1339:1482925
1311:Bibcode
1247:Bibcode
1115:6797082
854:epitope
634:in vivo
473:neutral
453:in vivo
393:in vivo
290:in vivo
203:fitness
156:History
33:genomes
29:viruses
6902:
6895:420405
6892:
6843:
6808:
6798:
6749:
6739:
6690:
6680:
6638:
6628:
6589:
6579:
6540:
6530:
6491:
6484:129678
6481:
6432:
6424:
6374:
6364:
6310:
6276:
6221:
6211:
6180:
6172:
6123:
6116:241137
6113:
6074:
6033:
5998:
5963:
5928:
5921:402129
5918:
5869:
5861:
5835:Nature
5818:
5808:
5767:
5726:
5716:
5667:
5657:
5623:
5613:
5572:
5562:
5513:
5505:
5462:
5452:
5403:
5396:523252
5393:
5354:
5344:
5334:
5303:
5293:
5267:Nature
5239:
5229:
5185:
5175:
5137:
5127:
5086:
5079:114966
5076:
5037:
5027:
4988:
4980:
4945:
4910:
4903:371084
4900:
4861:
4820:
4813:238028
4810:
4749:
4718:
4683:
4673:
4634:
4626:
4591:
4584:111862
4581:
4539:
4501:
4457:
4449:
4414:
4404:
4378:
4368:
4316:
4278:
4270:
4262:
4252:
4216:
4208:
4168:
4158:
4109:
4099:
4087:(10).
4049:
4039:
4013:Nature
3987:
3947:
3937:
3893:
3883:
3834:
3824:
3783:
3773:
3731:
3721:
3677:
3667:
3628:
3593:
3544:
3534:
3495:
3485:
3446:
3436:
3389:
3379:
3335:
3325:
3286:
3278:
3230:
3218:(38).
3186:
3176:
3128:
3118:
3072:
3062:
3018:
3008:
2967:
2957:
2918:
2908:
2867:
2857:
2826:
2785:
2775:
2734:
2693:
2683:
2634:
2624:
2585:
2577:
2539:
2504:
2494:
2452:
2442:
2411:
2405:593400
2403:
2368:
2360:
2322:
2314:
2276:
2266:
2220:
2180:
2144:
2096:
2086:
2047:
2010:
1979:
1972:236352
1969:
1927:
1921:657273
1919:
1877:
1867:
1836:
1828:
1775:
1735:
1727:
1684:
1674:
1606:
1598:
1558:
1533:
1523:
1471:
1436:
1391:
1381:
1337:
1329:
1303:Nature
1277:
1267:
1205:
1197:
1130:
1122:
1112:
1104:
945:APOBEC
846:T cell
842:B cell
342:prions
274:plaque
6841:S2CID
6742:34449
6430:S2CID
6367:15492
6178:S2CID
5867:S2CID
5593:eLife
5511:S2CID
4986:S2CID
4632:S2CID
4455:S2CID
4371:50777
4276:S2CID
4214:S2CID
3284:S2CID
2945:(1).
2686:19753
2583:S2CID
2502:S2CID
2409:S2CID
2366:S2CID
2320:S2CID
1925:S2CID
1834:S2CID
1733:S2CID
1604:S2CID
1526:24164
1434:S2CID
1335:S2CID
1270:22276
1237:arXiv
1038:than
868:>
860:>
220:drift
23:is a
6900:PMID
6806:PMID
6747:PMID
6688:PMID
6636:PMID
6587:PMID
6538:PMID
6489:PMID
6422:PMID
6372:PMID
6308:ISBN
6274:PMID
6243:help
6219:PMID
6209:ISBN
6170:PMID
6121:PMID
6072:PMID
6031:PMID
5996:PMID
5961:PMID
5926:PMID
5859:PMID
5816:PMID
5765:PMID
5724:PMID
5665:PMID
5655:ISBN
5621:PMID
5570:PMID
5503:PMID
5460:PMID
5401:PMID
5352:PMID
5332:ISBN
5301:PMID
5237:PMID
5183:PMID
5173:ISBN
5135:PMID
5084:PMID
5035:PMID
4978:PMID
4943:PMID
4908:PMID
4859:PMID
4818:PMID
4773:help
4747:ISBN
4716:PMID
4681:PMID
4624:PMID
4589:PMID
4537:PMID
4499:PMID
4447:PMID
4412:OCLC
4402:ISBN
4376:PMID
4314:PMID
4268:PMID
4260:OCLC
4250:ISBN
4206:PMID
4166:PMID
4107:PMID
4047:PMID
3985:ISBN
3945:PMID
3891:PMID
3832:PMID
3781:PMID
3729:PMID
3675:PMID
3626:PMID
3591:PMID
3542:PMID
3493:PMID
3444:PMID
3387:PMID
3333:PMID
3276:PMID
3228:ISSN
3184:PMID
3126:PMID
3070:PMID
3016:PMID
2965:PMID
2916:PMID
2865:PMID
2855:ISBN
2824:PMID
2783:PMID
2732:PMID
2691:PMID
2632:PMID
2575:PMID
2559:Cell
2537:PMID
2521:Gene
2492:ISBN
2450:PMID
2440:ISBN
2401:PMID
2358:PMID
2312:PMID
2296:Cell
2274:PMID
2264:ISBN
2218:ISBN
2178:PMID
2162:Gene
2142:PMID
2094:PMID
2045:PMID
2029:Gene
2008:ISBN
1977:PMID
1917:PMID
1901:Cell
1875:PMID
1865:ISBN
1826:PMID
1773:PMID
1725:PMID
1682:PMID
1596:ISSN
1556:OCLC
1531:PMID
1469:PMID
1389:PMID
1327:PMID
1275:PMID
1203:PMID
1195:ISSN
1120:PMID
1102:ISSN
1074:):
1068:2000
941:ADAR
904:and
844:and
197:and
181:and
84:host
47:and
6890:PMC
6880:doi
6868:101
6833:doi
6796:PMC
6786:doi
6737:PMC
6727:doi
6678:PMC
6670:doi
6626:PMC
6618:doi
6577:PMC
6569:doi
6528:PMC
6520:doi
6479:PMC
6469:doi
6412:hdl
6404:doi
6362:PMC
6352:doi
6300:doi
6266:doi
6201:doi
6160:doi
6111:PMC
6103:doi
6062:doi
6023:doi
6019:285
5988:doi
5984:264
5953:doi
5949:106
5916:PMC
5906:doi
5851:doi
5839:348
5806:PMC
5796:doi
5755:doi
5714:PMC
5704:doi
5692:103
5647:doi
5611:PMC
5601:doi
5560:PMC
5550:doi
5495:doi
5483:311
5450:PMC
5440:doi
5428:111
5391:PMC
5383:doi
5342:PMC
5324:doi
5291:PMC
5283:doi
5271:439
5227:PMC
5217:doi
5165:doi
5125:PMC
5115:doi
5074:PMC
5066:doi
5025:PMC
5017:doi
4970:doi
4935:doi
4931:369
4898:PMC
4890:doi
4849:doi
4845:223
4808:PMC
4800:doi
4739:doi
4708:doi
4704:339
4671:PMC
4663:doi
4659:331
4616:doi
4579:PMC
4571:doi
4489:doi
4439:doi
4366:PMC
4356:doi
4306:doi
4302:360
4240:doi
4198:doi
4194:523
4156:PMC
4146:doi
4134:107
4097:PMC
4089:doi
4037:PMC
4029:doi
4017:505
3935:PMC
3925:doi
3881:PMC
3871:doi
3859:112
3822:PMC
3812:doi
3808:109
3771:PMC
3763:doi
3719:PMC
3709:doi
3665:PMC
3657:doi
3618:doi
3581:doi
3532:PMC
3524:doi
3483:PMC
3475:doi
3434:PMC
3426:doi
3377:PMC
3367:doi
3323:PMC
3315:doi
3268:doi
3220:doi
3174:PMC
3166:doi
3116:PMC
3106:doi
3060:PMC
3050:doi
3006:PMC
2996:doi
2955:PMC
2947:doi
2906:PMC
2896:doi
2892:114
2847:doi
2814:doi
2773:PMC
2763:doi
2722:doi
2718:268
2681:PMC
2671:doi
2622:PMC
2614:doi
2567:doi
2529:doi
2525:122
2484:doi
2432:doi
2393:doi
2350:doi
2304:doi
2256:doi
2210:doi
2170:doi
2134:doi
2122:215
2084:PMC
2076:doi
2072:195
2037:doi
2000:doi
1967:PMC
1959:doi
1909:doi
1857:doi
1816:hdl
1808:doi
1796:460
1765:doi
1717:doi
1672:PMC
1664:doi
1588:doi
1521:PMC
1511:doi
1461:doi
1424:hdl
1416:doi
1379:PMC
1369:doi
1365:100
1319:doi
1307:412
1265:PMC
1255:doi
1185:hdl
1175:doi
1110:PMC
1092:doi
1070:) (
949:RIP
796:or
689:or
567:or
516:or
340:in
171:max
167:max
43:as
27:of
6954::
6898:.
6888:.
6878:.
6866:.
6862:.
6839:.
6829:32
6827:.
6804:.
6794:.
6784:.
6772:.
6768:.
6745:.
6735:.
6725:.
6715:98
6713:.
6709:.
6686:.
6676:.
6666:80
6664:.
6660:.
6648:^
6634:.
6624:.
6614:11
6612:.
6608:.
6585:.
6575:.
6565:20
6563:.
6559:.
6536:.
6526:.
6516:29
6514:.
6510:.
6487:.
6477:.
6467:.
6457:99
6455:.
6451:.
6428:.
6420:.
6410:.
6400:11
6398:.
6384:^
6370:.
6360:.
6350:.
6340:96
6338:.
6334:.
6322:^
6306:.
6286:^
6272:.
6262:26
6260:.
6235::
6233:}}
6229:{{
6217:.
6207:.
6176:.
6168:.
6158:.
6148:24
6146:.
6142:.
6119:.
6109:.
6099:66
6097:.
6093:.
6070:.
6058:85
6056:.
6052:.
6029:.
6017:.
5994:.
5982:.
5959:.
5947:.
5924:.
5914:.
5904:.
5894:89
5892:.
5888:.
5865:.
5857:.
5849:.
5837:.
5814:.
5804:.
5790:.
5786:.
5763:.
5749:.
5745:.
5722:.
5712:.
5702:.
5690:.
5686:.
5663:.
5653:.
5633:^
5619:.
5609:.
5595:.
5591:.
5568:.
5558:.
5548:.
5538:12
5536:.
5532:.
5509:.
5501:.
5493:.
5481:.
5458:.
5448:.
5438:.
5426:.
5422:.
5399:.
5389:.
5379:78
5377:.
5373:.
5350:.
5340:.
5330:.
5299:.
5289:.
5281:.
5269:.
5265:.
5249:^
5235:.
5225:.
5211:.
5207:.
5195:^
5181:.
5171:.
5147:^
5133:.
5123:.
5109:.
5105:.
5082:.
5072:.
5062:75
5060:.
5056:.
5033:.
5023:.
5013:20
5011:.
5007:.
4984:.
4976:.
4966:37
4964:.
4941:.
4929:.
4906:.
4896:.
4886:78
4884:.
4880:.
4857:.
4843:.
4839:.
4816:.
4806:.
4796:67
4794:.
4790:.
4765::
4763:}}
4759:{{
4745:.
4714:.
4702:.
4679:.
4669:.
4657:.
4653:.
4630:.
4622:.
4612:16
4610:.
4587:.
4577:.
4567:74
4565:.
4561:.
4549:^
4533:10
4531:.
4527:.
4511:^
4497:.
4485:25
4483:.
4479:.
4467:^
4453:.
4445:.
4435:56
4433:.
4410:.
4388:^
4374:.
4364:.
4354:.
4344:88
4342:.
4338:.
4326:^
4312:.
4300:.
4288:^
4274:.
4266:.
4258:.
4248:.
4226:^
4212:.
4204:.
4192:.
4178:^
4164:.
4154:.
4144:.
4132:.
4128:.
4105:.
4095:.
4085:91
4083:.
4079:.
4059:^
4045:.
4035:.
4027:.
4015:.
4011:.
3999:^
3957:^
3943:.
3933:.
3919:.
3915:.
3903:^
3889:.
3879:.
3869:.
3857:.
3853:.
3830:.
3820:.
3806:.
3802:.
3779:.
3769:.
3759:22
3757:.
3753:.
3741:^
3727:.
3717:.
3705:10
3703:.
3699:.
3687:^
3673:.
3663:.
3653:34
3651:.
3647:.
3624:.
3614:17
3612:.
3589:.
3579:.
3569:20
3567:.
3563:.
3540:.
3530:.
3520:92
3518:.
3514:.
3491:.
3481:.
3469:.
3465:.
3442:.
3432:.
3424:.
3412:.
3408:.
3385:.
3375:.
3361:.
3357:.
3345:^
3331:.
3321:.
3311:73
3309:.
3305:.
3282:.
3274:.
3264:47
3262:.
3240:^
3226:.
3216:34
3214:.
3196:^
3182:.
3172:.
3162:82
3160:.
3156:.
3138:^
3124:.
3114:.
3102:15
3100:.
3096:.
3082:^
3068:.
3058:.
3044:.
3040:.
3028:^
3014:.
3004:.
2992:14
2990:.
2986:.
2963:.
2953:.
2943:91
2941:.
2937:.
2914:.
2904:.
2890:.
2886:.
2863:.
2853:.
2822:.
2808:.
2804:.
2781:.
2771:.
2757:.
2753:.
2730:.
2716:.
2712:.
2689:.
2679:.
2669:.
2659:94
2657:.
2653:.
2630:.
2620:.
2610:81
2608:.
2604:.
2581:.
2573:.
2563:59
2561:.
2549:^
2535:.
2523:.
2500:.
2490:.
2462:^
2448:.
2438:.
2407:.
2399:.
2389:64
2387:.
2364:.
2356:.
2346:31
2344:.
2332:^
2318:.
2310:.
2300:58
2298:.
2286:^
2272:.
2262:.
2254:.
2246:.
2232:^
2216:.
2208:.
2190:^
2176:.
2166:40
2164:.
2140:.
2132:.
2120:.
2106:^
2092:.
2082:.
2070:.
2066:.
2043:.
2031:.
2006:.
1975:.
1965:.
1955:68
1953:.
1949:.
1937:^
1923:.
1915:.
1905:13
1903:.
1887:^
1873:.
1863:.
1832:.
1824:.
1814:.
1806:.
1794:.
1771:.
1761:16
1759:.
1745:^
1731:.
1723:.
1715:.
1705:58
1703:.
1680:.
1670:.
1660:76
1658:.
1654:.
1616:^
1602:.
1594:.
1584:65
1582:.
1568:^
1529:.
1519:.
1509:.
1499:96
1497:.
1493:.
1481:^
1467:.
1455:.
1432:.
1422:.
1412:92
1410:.
1387:.
1377:.
1363:.
1359:.
1347:^
1333:.
1325:.
1317:.
1305:.
1301:.
1287:^
1273:.
1263:.
1253:.
1245:.
1233:96
1231:.
1227:.
1215:^
1201:.
1193:.
1183:.
1169:.
1165:.
1143:^
1126:.
1118:.
1108:.
1100:.
1088:15
1086:.
1080:.
947:,
943:,
759:.
563:,
433:In
417:QU
332:,
300:.
226:.
152:.
98:.
19:A
6906:.
6882::
6874::
6847:.
6835::
6812:.
6788::
6780::
6774:5
6753:.
6729::
6721::
6694:.
6672::
6642:.
6620::
6593:.
6571::
6544:.
6522::
6495:.
6471::
6463::
6436:.
6414::
6406::
6378:.
6354::
6346::
6316:.
6302::
6280:.
6268::
6245:)
6225:.
6203::
6184:.
6162::
6154::
6127:.
6105::
6078:.
6064::
6037:.
6025::
6002:.
5990::
5967:.
5955::
5932:.
5908::
5900::
5873:.
5853::
5845::
5822:.
5798::
5792:7
5771:.
5757::
5751:2
5730:.
5706::
5698::
5671:.
5649::
5627:.
5603::
5597:5
5576:.
5552::
5544::
5517:.
5497::
5489::
5466:.
5442::
5434::
5407:.
5385::
5358:.
5326::
5307:.
5285::
5277::
5243:.
5219::
5213:1
5189:.
5167::
5141:.
5117::
5111:2
5090:.
5068::
5041:.
5019::
4992:.
4972::
4949:.
4937::
4914:.
4892::
4865:.
4851::
4824:.
4802::
4775:)
4755:.
4741::
4722:.
4710::
4687:.
4665::
4638:.
4618::
4595:.
4573::
4543:.
4505:.
4491::
4461:.
4441::
4418:.
4382:.
4358::
4350::
4320:.
4308::
4282:.
4242::
4220:.
4200::
4172:.
4148::
4140::
4113:.
4091::
4053:.
4031::
4023::
3993:.
3951:.
3927::
3921:4
3897:.
3873::
3865::
3838:.
3814::
3787:.
3765::
3735:.
3711::
3681:.
3659::
3632:.
3620::
3597:.
3583::
3575::
3548:.
3526::
3499:.
3477::
3471:4
3450:.
3428::
3420::
3414:7
3393:.
3369::
3363:6
3339:.
3317::
3290:.
3270::
3234:.
3222::
3190:.
3168::
3132:.
3108::
3076:.
3052::
3046:9
3022:.
2998::
2971:.
2949::
2922:.
2898::
2871:.
2849::
2830:.
2816::
2810:1
2789:.
2765::
2759:9
2738:.
2724::
2697:.
2673::
2665::
2638:.
2616::
2589:.
2569::
2543:.
2531::
2508:.
2486::
2456:.
2434::
2415:.
2395::
2372:.
2352::
2326:.
2306::
2280:.
2258::
2226:.
2212::
2184:.
2172::
2148:.
2136::
2128::
2100:.
2078::
2051:.
2039::
2033:1
2016:.
2002::
1983:.
1961::
1931:.
1911::
1881:.
1859::
1840:.
1818::
1810::
1802::
1779:.
1767::
1739:.
1719::
1711::
1688:.
1666::
1610:.
1590::
1562:.
1537:.
1513::
1505::
1475:.
1463::
1457:7
1440:.
1426::
1418::
1395:.
1371::
1341:.
1321::
1313::
1281:.
1257::
1249::
1239::
1209:.
1187::
1177::
1171:8
1134:.
1094::
908:.
665:.
429:T
425:N
421:E
413:Q
409:A
405:P
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.