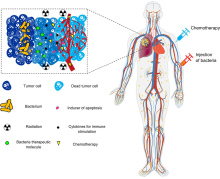
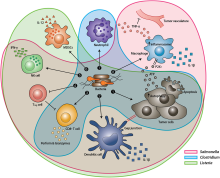
174:, cytokines, and chemokines, which further facilitate tumor regression. ① Bacterial toxins from S. Typhimurium, Listeria, and Clostridium can kill tumor cells directly by inducing apoptosis or autophagy. Toxins delivered via Salmonella can upregulate Connexin 43 (Cx43), leading to bacteria-induced gap junctions between the tumor and dendritic cells (DCs), which allow cross-presentation of tumor antigens to the DCs. ② Upon exposure to tumor antigens and interaction with bacterial components, DCs secrete robust amounts of the proinflammatory cytokine IL-1β, which subsequently activates CD8+ T cells. ③ The antitumor response of the activated CD8+ T cells is further enhanced by bacterial flagellin (a protein subunit of the bacterial flagellum) via TLR5 activation. The perforin and granzyme proteins secreted by activated CD8+ T cells efficiently kill tumor cells in primary and metastatic tumors. ④ Flagellin and TLR5 signaling also decreases the abundance of CD4+ CD25+ regulatory T (Treg) cells, which subsequently improves the antitumor response of the activated CD8+ T cells. ⑤ S. Typhimurium flagellin stimulates NK cells to produce interferon-γ (IFN-γ), an important cytokine for both innate and adaptive immunity. ⑥ Listeria-infected MDSCs shift into an immune-stimulating phenotype characterized by increased IL-12 production, which further enhances the CD8+ T and NK cell responses. ⑦ Both S. Typhimurium and Clostridium infection can stimulate significant neutrophil accumulation. Elevated secretion of TNF-α and TNF-related apoptosis-inducing ligand (TRAIL) by neutrophils enhances the immune response and kills tumor cells by inducing apoptosis. ⑧ The macrophage inflammasome is activated through contact with bacterial components (LPS and flagellin) and Salmonella-damaged cancer cells, leading to elevated secretion of IL-1β and TNF-α into the tumor microenvironment. NK cell: natural killer cell. Treg cell: regulatory T cell. MDSCs: myeloid-derived suppressor cells. P2X7 receptor: purinoceptor 7-extracellular ATP receptor. LPS: lipopolysaccharide
125:
109:
159:
27:
179:
167:
874:
317:
293:
225:
39:
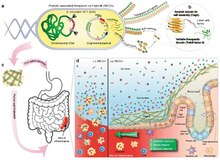
CsgA (yellow), the main proteinaceous component of the E. coli biofilm matrix, was genetically fused to a therapeutic domain—in this case, TFF3 (PDB ID: 19ET, bright green), which is a cytokine secreted by mucus-producing cells. The flexible linker (black) includes a 6xHis tag for detection purposes.
34:
Genetically engineered E. coli Nissle 1917 (EcN) with csg (curli) operon deletion (PBP8 strain) containing plasmids encoding a synthetic curli operon capable of producing chimeric CsgA proteins (yellow chevrons with appended bright green domains), which are secreted and self-assembled extracellularly
115:
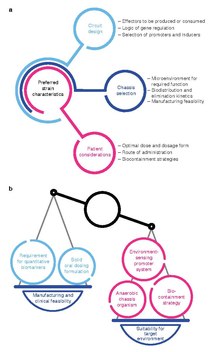
Several aspects require consideration during the design of an engineered bacterial therapeutic. The selection of a chassis organism can be guided by the desired site of activity and pharmacokinetic properties of the chassis, as well as manufacturing feasibility. The design of genetic circuits may
48:
Interaction of E. coli and the colonic mucosa. Inflammatory lesions in IBD result in loss of colonic crypt structure, damage to epithelial tissue, and compromised barrier integrity (left panel, (−) E. coli). The resulting invasion of luminal contents and recruitment of immune cells to the site
128:
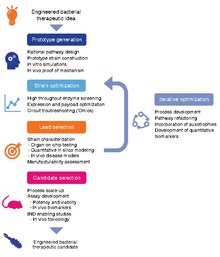
Schematic representation of a workflow for developing clinical candidate-quality engineered strains. The development workflow should incorporate technologies for optimizing strain potency, as well as predictive in vitro and in vivo assays, as well quantitative pharmacology models, to maximize
116:
also be influenced by the circuit's effectors, pragmatic concerns regarding inducer compounds, and the genetic stability of regulatory circuits. Critically, the design of an engineered bacterial drug may also be constrained by considerations for the needs of patients.
604:
Kurtz, Caroline B.; Millet, Yves A.; Puurunen, Marja K.; Perreault, Mylène; Charbonneau, Mark R.; Isabella, Vincent M.; Kotula, Jonathan W.; Antipov, Eugene; Dagon, Yossi; Denney, William S.; Wagner, David A. (2019-01-16).
1002:
Sedighi, Mansour; Zahedi
Bialvaei, Abed; Hamblin, Michael R.; Ohadi, Elnaz; Asadi, Arezoo; Halajzadeh, Masoumeh; Lohrasbi, Vahid; Mohammadzadeh, Nima; Amiriani, Taghi; Krutova, Marcela; Amini, Abolfazl (2019-04-05).
49:
exacerbates the local inflammation. The application of E. coli (right panel, (+) E. coli) reinforces barrier function, promotes epithelial restitution, and dampens inflammatory signaling to ameliorate IBD activity.
206:
tumors. This property tends to increase their residence time in the tumor, giving them longer to exert their therapeutic effects, in contrast to other bacteria that would be quickly cleared by the immune system.
170:
After systemic administration, bacteria localize to the tumor microenvironment. The interactions between bacteria, cancer cells, and the surrounding microenvironment cause various alterations in
120:
Optimal strain design often requires a balance between strain suitability for function in the target microenvironment and concerns for feasibility of manufacturing and clinical development.
273:
61:
that consists of a living organism that is used to treat a disease. This usually takes the form of a cell (animal, bacterial, or fungal) or a virus that has been
565:
141:. Currently, there is a large focus on: 1) identifying microbes that naturally produce therapeutic effects (for example, probiotic bacteria), and 2)
17:
590:
124:
232: by Pichet Praveschotinunt, Anna M. Duraj-Thatte, Ilia Gelfat, Franziska Bahl, David B. Chou & Neel S. Joshi available under the
248:
195:
729:
44:
Engineered bacteria are produced in bulk before delivery to the GI tract. A site of colonic inflammation is highlighted in red.
607:"An engineered E. coli Nissle improves hyperammonemia and survival in mice and shows dose-dependent exposure in healthy humans"
782:
108:
513:
158:
1149:
756:"Why now is the time for programmable living medicines: insights from Jim Collins, Aoife Brennan, and Jason Kelly"
1154:
1139:
1129:
21:
755:
194:
environments are particularly attractive for this purpose, as they will tend to migrate to, invade (through the
171:
896:
Sieow, Brendan Fu-Long; Wun, Kwok Soon; Yong, Wei Peng; Hwang, In Young; Chang, Matthew Wook (December 2020).
1124:
142:
1144:
539:
26:
324: by Mark R. Charbonneau, Vincent M. Isabella, Ning Li & Caroline B. Kurtz available under the
300: by Mark R. Charbonneau, Vincent M. Isabella, Ning Li & Caroline B. Kurtz available under the
69:
properties that is injected into a patient. Perhaps the oldest use of a living medicine is the use of
178:
1159:
1134:
881: by Mai Thi-Quynh Duong, Yeshan Qin, Sung-Hwan You & Jung-Joon Min available under the
203:
199:
274:"Engineering Living Medicines for Chronic Diseases | SBE | Society for Biological Engineering"
187:
166:
566:"From 'living' cement to medicine-delivering biofilms, biologists remake the material world"
186:
There is tremendous interest in using bacteria as a therapy to treat tumors. In particular,
820:
675:
882:
873:
664:"Developing a new class of engineered live bacterial therapeutics to treat human diseases"
325:
316:
301:
292:
233:
224:
8:
62:
1005:"Therapeutic bacteria to combat cancer; current advances, challenges, and opportunities"
841:
824:
808:
679:
30:
Genetically engineered probiotics as living medicines to treat intestinal inflammation.
1096:
1061:
1037:
1004:
979:
946:
854:
706:
663:
662:
Charbonneau, Mark R.; Isabella, Vincent M.; Li, Ning; Kurtz, Caroline B. (2020-04-08).
644:
584:
489:
454:
430:
397:
373:
340:
133:
Development of living medicines is an extremely active research area in the fields of
1164:
1101:
1083:
1062:"The role of bacteria in cancer therapy – enemies in the past, but allies at present"
1042:
1024:
984:
966:
927:
919:
858:
846:
711:
693:
636:
628:
494:
476:
435:
417:
378:
360:
191:
134:
93:
58:
648:
1091:
1073:
1032:
1016:
974:
958:
909:
836:
828:
701:
683:
618:
484:
466:
425:
409:
368:
352:
914:
897:
623:
606:
413:
945:
Duong, Mai Thi-Quynh; Qin, Yeshan; You, Sung-Hwan; Min, Jung-Joon (2019-12-11).
688:
1078:
962:
1118:
1087:
1028:
970:
923:
697:
632:
480:
421:
364:
321:
297:
229:
89:
85:
878:
832:
471:
453:
Kitada, Tasuku; DiAndreth, Breanna; Teague, Brian; Weiss, Ron (2018-02-09).
1105:
1046:
988:
931:
850:
715:
640:
498:
439:
382:
138:
81:
74:
807:
Gurbatri, Candice R.; Arpaia, Nicholas; Danino, Tal (25 November 2022).
455:"Programming gene and engineered-cell therapies with synthetic biology"
1020:
77:, though living medicines have advanced tremendously since that time.
97:
356:
783:"Living medicines: Ginkgo's machine to disrupt the pharma industry"
162:
Schematic of therapeutic bacteria strategies against hypoxic tumors
1001:
66:
730:"Gene Circuits Empower Next-Generation Cell and Gene Therapies"
947:"Bacteria-cancer interactions: bacteria-based cancer therapy"
396:
Fischbach, M. A.; Bluestone, J. A.; Lim, W. A. (2013-04-03).
70:
898:"Tweak to Treat: Reprograming Bacteria for Cancer Treatment"
603:
661:
452:
249:"'Living medicine' helps make toxic ammonia breakthrough"
1060:
Song, Shiyu; Vuai, Miza S.; Zhong, Mintao (2018-03-15).
341:"Emerging biomedical applications of synthetic biology"
809:"Engineering bacteria as interactive cancer therapies"
398:"Cell-Based Therapeutics: The Next Pillar of Medicine"
395:
339:
Weber, Wilfried; Fussenegger, Martin (January 2012).
806:
103:
895:
182:Bacteria involved in causing and treating cancers
18:Biological therapy for inflammatory bowel disease
1116:
734:GEN - Genetic Engineering and Biotechnology News
338:
129:translational potential for patient populations.
944:
564:ServiceFeb. 18, Robert F. (18 February 2020).
563:
1059:
514:"Why 2018 Was the Year of 'Living' Medicine"
589:: CS1 maint: numeric names: authors list (
145:organisms to produce therapeutic effects.
1095:
1077:
1036:
978:
913:
840:
705:
687:
622:
488:
470:
429:
372:
177:
165:
157:
123:
107:
25:
511:
1117:
246:
35:into therapeutic curli hybrid fibers.
951:Experimental & Molecular Medicine
780:
537:
80:Examples of living medicines include
13:
512:McCarty, Niko (18 December 2018).
14:
1176:
781:Costa, Kevin (20 February 2019).
153:
877: This article incorporates
872:
320: This article incorporates
315:
296: This article incorporates
291:
228: This article incorporates
223:
1053:
995:
938:
889:
865:
800:
774:
748:
722:
655:
597:
557:
531:
247:Sample, Ian (16 January 2019).
172:tumor-infiltrating immune cells
148:
104:Development of living medicines
96:, a subset of the latter being
22:Biologics for immunosuppression
611:Science Translational Medicine
505:
446:
402:Science Translational Medicine
389:
332:
308:
284:
266:
240:
216:
1:
540:"The Era of Living Medicines"
538:Kelly, Jason (12 June 2019).
210:
1066:Infectious Agents and Cancer
915:10.1016/j.trecan.2020.11.004
624:10.1126/scitranslmed.aau7975
414:10.1126/scitranslmed.3005568
7:
10:
1181:
689:10.1038/s41467-020-15508-1
15:
1079:10.1186/s13027-018-0180-y
963:10.1038/s12276-019-0297-0
1150:Pharmaceutical industry
833:10.1126/science.add9667
472:10.1126/science.aad1067
345:Nature Reviews Genetics
143:genetically programming
1155:Life sciences industry
1140:Biotechnology products
1130:Biological engineering
200:tumor microenvironment
183:
175:
163:
130:
121:
94:bacterial therapeutics
63:genetically engineered
50:
668:Nature Communications
188:tumor-homing bacteria
181:
169:
161:
127:
111:
82:cellular therapeutics
29:
1125:Bacteria and humans
825:2022Sci...378..858G
680:2020NatCo..11.1738C
1145:Biopharmaceuticals
465:(6376): eaad1067.
184:
176:
164:
131:
122:
90:phage therapeutics
86:immunotherapeutics
51:
1021:10.1002/cam4.2148
819:(6622): 858–864.
736:. 1 February 2020
617:(475): eaau7975.
196:leaky vasculature
135:synthetic biology
1172:
1110:
1109:
1099:
1081:
1057:
1051:
1050:
1040:
1015:(6): 3167–3181.
999:
993:
992:
982:
942:
936:
935:
917:
902:Trends in Cancer
893:
887:
876:
869:
863:
862:
844:
804:
798:
797:
795:
793:
778:
772:
771:
769:
767:
752:
746:
745:
743:
741:
726:
720:
719:
709:
691:
659:
653:
652:
626:
601:
595:
594:
588:
580:
578:
576:
561:
555:
554:
552:
550:
535:
529:
528:
526:
524:
509:
503:
502:
492:
474:
450:
444:
443:
433:
393:
387:
386:
376:
336:
330:
319:
312:
306:
295:
288:
282:
281:
270:
264:
263:
261:
259:
244:
238:
227:
220:
1180:
1179:
1175:
1174:
1173:
1171:
1170:
1169:
1160:Specialty drugs
1115:
1114:
1113:
1058:
1054:
1009:Cancer Medicine
1000:
996:
943:
939:
894:
890:
870:
866:
805:
801:
791:
789:
779:
775:
765:
763:
754:
753:
749:
739:
737:
728:
727:
723:
660:
656:
602:
598:
582:
581:
574:
572:
562:
558:
548:
546:
544:Ginkgo Bioworks
536:
532:
522:
520:
510:
506:
451:
447:
408:(179): 179ps7.
394:
390:
357:10.1038/nrg3094
337:
333:
313:
309:
289:
285:
272:
271:
267:
257:
255:
245:
241:
221:
217:
213:
190:that thrive in
156:
151:
106:
55:living medicine
24:
12:
11:
5:
1178:
1168:
1167:
1162:
1157:
1152:
1147:
1142:
1137:
1132:
1127:
1112:
1111:
1052:
994:
937:
908:(5): 447–464.
888:
864:
799:
773:
762:. 2 April 2019
747:
721:
654:
596:
556:
530:
504:
445:
388:
331:
307:
283:
265:
239:
214:
212:
209:
155:
154:Cancer therapy
152:
150:
147:
105:
102:
9:
6:
4:
3:
2:
1177:
1166:
1163:
1161:
1158:
1156:
1153:
1151:
1148:
1146:
1143:
1141:
1138:
1136:
1135:Biotechnology
1133:
1131:
1128:
1126:
1123:
1122:
1120:
1107:
1103:
1098:
1093:
1089:
1085:
1080:
1075:
1071:
1067:
1063:
1056:
1048:
1044:
1039:
1034:
1030:
1026:
1022:
1018:
1014:
1010:
1006:
998:
990:
986:
981:
976:
972:
968:
964:
960:
956:
952:
948:
941:
933:
929:
925:
921:
916:
911:
907:
903:
899:
892:
886:
884:
880:
875:
868:
860:
856:
852:
848:
843:
838:
834:
830:
826:
822:
818:
814:
810:
803:
788:
784:
777:
761:
757:
751:
735:
731:
725:
717:
713:
708:
703:
699:
695:
690:
685:
681:
677:
673:
669:
665:
658:
650:
646:
642:
638:
634:
630:
625:
620:
616:
612:
608:
600:
592:
586:
571:
567:
560:
545:
541:
534:
519:
515:
508:
500:
496:
491:
486:
482:
478:
473:
468:
464:
460:
456:
449:
441:
437:
432:
427:
423:
419:
415:
411:
407:
403:
399:
392:
384:
380:
375:
370:
366:
362:
358:
354:
350:
346:
342:
335:
329:
327:
323:
318:
311:
305:
303:
299:
294:
287:
279:
278:www.aiche.org
275:
269:
254:
250:
243:
237:
235:
231:
226:
219:
215:
208:
205:
201:
197:
193:
189:
180:
173:
168:
160:
146:
144:
140:
136:
126:
119:
114:
110:
101:
99:
95:
91:
87:
83:
78:
76:
72:
68:
64:
60:
57:is a type of
56:
47:
43:
38:
33:
28:
23:
19:
1069:
1065:
1055:
1012:
1008:
997:
957:(12): 1–15.
954:
950:
940:
905:
901:
891:
871:
867:
816:
812:
802:
790:. Retrieved
786:
776:
764:. Retrieved
759:
750:
738:. Retrieved
733:
724:
671:
667:
657:
614:
610:
599:
573:. Retrieved
569:
559:
547:. Retrieved
543:
533:
521:. Retrieved
517:
507:
462:
458:
448:
405:
401:
391:
351:(1): 21–35.
348:
344:
334:
314:
310:
290:
286:
277:
268:
256:. Retrieved
253:The Guardian
252:
242:
222:
218:
185:
149:Applications
139:microbiology
132:
117:
112:
79:
75:bloodletting
54:
52:
45:
41:
36:
31:
674:(1): 1738.
84:(including
67:therapeutic
65:to possess
1119:Categories
787:SynBioBeta
760:SynBioBeta
211:References
98:probiotics
16:See also:
1088:1750-9378
1029:2045-7634
971:2092-6413
924:2405-8033
883:CC BY 4.0
859:253839557
698:2041-1723
633:1946-6234
585:cite news
481:0036-8075
422:1946-6234
365:1471-0056
326:CC BY 4.0
302:CC BY 4.0
234:CC BY 4.0
1165:Pharmacy
1106:29568324
1072:(1): 9.
1047:30950210
989:31827064
932:33303401
885:license.
851:36423303
842:10584033
716:32269218
649:58031579
641:30651324
499:29439214
440:23552369
383:22124480
328:license.
304:license.
236:license.
204:colonize
59:biologic
1097:5856380
1038:6558487
980:6906302
821:Bibcode
813:Science
792:5 April
766:5 April
740:5 April
707:7142098
676:Bibcode
575:5 April
549:5 April
523:5 April
490:7643872
459:Science
431:3772767
374:7097403
258:5 April
198:in the
192:hypoxic
71:leeches
1104:
1094:
1086:
1045:
1035:
1027:
987:
977:
969:
930:
922:
857:
849:
839:
714:
704:
696:
647:
639:
631:
518:Medium
497:
487:
479:
438:
428:
420:
381:
371:
363:
202:) and
92:, and
855:S2CID
645:S2CID
1102:PMID
1084:ISSN
1043:PMID
1025:ISSN
985:PMID
967:ISSN
928:PMID
920:ISSN
879:text
847:PMID
794:2020
768:2020
742:2020
712:PMID
694:ISSN
637:PMID
629:ISSN
591:link
577:2020
570:AAAS
551:2020
525:2020
495:PMID
477:ISSN
436:PMID
418:ISSN
379:PMID
361:ISSN
322:text
298:text
260:2020
230:text
137:and
73:for
20:and
1092:PMC
1074:doi
1033:PMC
1017:doi
975:PMC
959:doi
910:doi
837:PMC
829:doi
817:378
702:PMC
684:doi
619:doi
485:PMC
467:doi
463:359
426:PMC
410:doi
369:PMC
353:doi
88:),
1121::
1100:.
1090:.
1082:.
1070:13
1068:.
1064:.
1041:.
1031:.
1023:.
1011:.
1007:.
983:.
973:.
965:.
955:51
953:.
949:.
926:.
918:.
904:.
900:.
853:.
845:.
835:.
827:.
815:.
811:.
785:.
758:.
732:.
710:.
700:.
692:.
682:.
672:11
670:.
666:.
643:.
635:.
627:.
615:11
613:.
609:.
587:}}
583:{{
568:.
542:.
516:.
493:.
483:.
475:.
461:.
457:.
434:.
424:.
416:.
404:.
400:.
377:.
367:.
359:.
349:13
347:.
343:.
276:.
251:.
100:.
53:A
1108:.
1076::
1049:.
1019::
1013:8
991:.
961::
934:.
912::
906:7
861:.
831::
823::
796:.
770:.
744:.
718:.
686::
678::
651:.
621::
593:)
579:.
553:.
527:.
501:.
469::
442:.
412::
406:5
385:.
355::
280:.
262:.
118:b
113:a
46:d
42:c
37:b
32:a
Text is available under the Creative Commons Attribution-ShareAlike License. Additional terms may apply.